Lactuca serriola L. (syn. L. scariola L.), commonly known as prickly lettuce, is native to Europe and is considered to be a major gene source for the breeding of lettuce (L. sativa). L. serriola invasion in Korea was first detected in the late 1970s [1]. Since then, the plant has become widely naturalized. It has disrupted native plant communities and has been designated as one of 12 “Harmful nonindigenous plants” by the Korean Ministry of Environment [2]. In May 2020, downy mildew infections on L. serriola were observed on the roadside (35°52'07" N, 126°50'25" E) in Gimje-si, Korea.
Symptoms of infected L. serriola appeared as yellow brownish or light brownish spots on the leaves (Fig. 1A and 1B). The lesions were water-soaked, polyangular, and clearly delimited by the leaf veins (Fig. 1C). White dense oomycete growth was observed on the corresponding abaxial leaf surface (Fig. 1D). A dried specimen was deposited in the Kunsan National University Herbarium (KSNUH656), Korea.
To identify the causal agent, morphological and molecular sequence analyses were performed. For detailed microscopic examination, conidiophores and conidia formed underneath the infected leaves were examined and photographed using an Imager M2 AX10 microscope (Carl Zeiss, Jena, Germany) equipped with an AxioCam 512 camera (Carl Zeiss). Conidiophores were tree-like, hyaline, straight to slightly curved, with sizes of (211.1-)290.4 to 503.1(-632.0)×(7.3-)8.7 to 11.6(-14.4) μm (n=50). The conidiophores were monopodially branched in four to six orders (Fig. 1E and 1F). Trunks were hyaline with sizes of (77.7-)119.5 to 292.9(-409.6)×(7.0-)8.6 to 11.6(-14.3) μm (n=50). Ultimate branchlets were in pairs or triplets and were straight to slightly curved, 7.8 to 26.7 μm long and 8.2 to 10.5 μm wide at the base (n=50), with truncate or infrequently swollen tips (Fig. 1G and 1H). Conidia were hyaline, ovoidal or lemon-shaped, and measured (17.2-)17.5 to 21.3(-24.6)×(15.2-)15.1 to 17.5(-19.8) μm, with a length to width ratio of (1.09-)1.14 to 1.24(-1.29) (n=50) (Fig. 1I and 1J). The morphological characteristics unequivocally identified the pathogen as belonging to the genus Bremia, with close resemblance to B. lactucae [3]. Two different species of Bremia have been accepted as the causal agents of downy mildew on Lactuca species: B. lactucae and B. elliptica. These two species are easily distinguishable by multigene phylogeny but are indistinguishable morphologically. Instead, the latter species is a specialist parasitic only to L. indica but has not previously been reported in L. serriola and L. sativa.
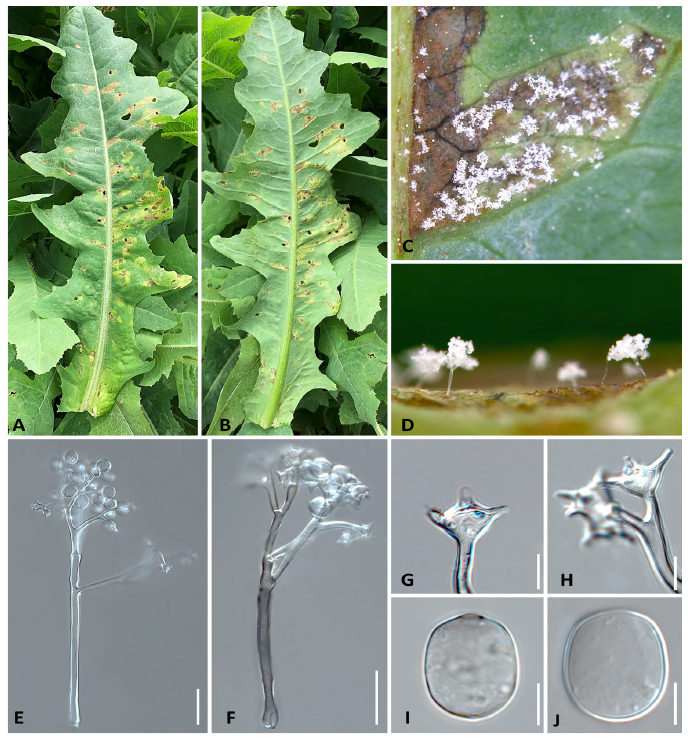

Fig. 1. Downy mildew disease caused by Bremia lactucae on Lactuca serriola in Korea. A and B, Vein-limited spots above (A) and below (B) an infected leaf. C, Close-up view of veinlimited downy mildew growth developing on the lower surface. D, Conidiophores emerging from stomata under a stereomicroscope. E and F, Conidiophore under a differential interference contrast microscope. G and H, Ultimate branchlets. I and J, Conidium. Scale bars: 50 μm for E and F, 10 μm for G-J.
To confirm morphological identification, genomic DNA was extracted from diseased leaves of KSNUH656 using the DNeasy Plant Mini Kit (Qiagen, Germantown, MD, USA). Three DNA gene regions were sequenced: the internal transcribed spacer 1 (ITS1) rDNA region using primers DC6 and ITS2, partial large subunit (LSU) ribosomal DNA (rDNA) using LROR and LR6-O [3], and cytochrome c oxidase II (cox2) mitochondrial DNA (mtDNA) using cox2-F and cox2-RC4 [4]. Amplicons were sequenced using a DNA sequencing service (Macrogen, Seoul, Korea) with the primers used for amplification. The resulting sequences were edited using the Lasergene software package (DNASTAR, Madison, WI, USA) and deposited in GenBank (accession no. MZ389974 for ITS, MZ389907 for LSU, and MZ435743 for cox2). The BLASTn search of these sequences revealed that the Korean sample was identical to the sequences of B. lactucae parasitic to L. sativa (KT249021 for ITS, MF693907 for LSU, and MF687313 for cox2).
Alignments of ITS, LSU rDNA, and cox2 mtDNA sequences were generated using MAFFT7 [5] by choosing the Q-INS-1 algorithm and concatenated using SequenceMatrix [6]. A minimum evolution tree was constructed using MEGA 7.0 [7] with the Tamura–Nei model. In the phylogenetic tree for the concatenated dataset (Fig. 2), the Korean sample fell into the crown clade, representing B. lactucae sensu stricto, parasitic to L. serriola and L. sativa, with a high bootstrap support of 91%, but distant from B. elliptica parasitic to L. indica. Based on morphological characteristics and molecular sequencing data, the downy mildew pathogen was identified as B. lactucae.
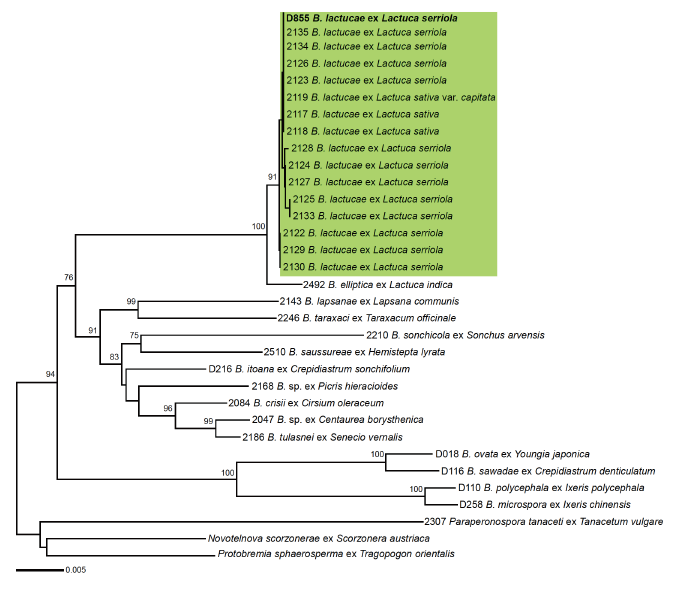
Fig. 2. Minimum evolution tree based on a concatenated alignment of the internal transcribed spacer 1, large subunit of ribosomal rDNA, and cox2 mtDNA sequences. Bootstrap support values higher than 70% are displayed above the branches. Green box presents a clade of Bremia lactucae specimens originating from Lactuca serriola and L. sativa, where the Korean downy mildew specimen from L. serriola is shown in bold. Branch lengths are proportional to the estimated number of nucleotide substitutions. The scale bar indicates the number of nucleotide substitutions per site.
Downy mildews (Peronosporaceae, Oomycota) are a monophyletic group that consists of obligate biotrophic species that are parasitic on a wide range of mono-and dicotyledonous plants [8]. Downy mildew is a notorious disease for many genera and species of the family Asteraceae. Among these, B. lactucae is one of the most well-known species that causes devastating damage in the cultivation of lettuce (L. sativa) [9]. Recent phylogenetic studies with multigene sequences have provided a good resolution for discriminating Bremia species [3,4,10-12] and have led to the description of three novel species [13,14].
Previously, B. lactucae infection of prickly lettuce was described mainly in Europe, and that in Asia was restricted to Pakistan [15]. To our knowledge, this is the first report of downy mildew caused by B. lactucae in prickly lettuce in East Asia. As B. lactucae populations that originate from prickly lettuce may play a major role in the epidemiology of downy mildew on edible lettuce (L. sativa) [16], the present finding of downy mildew on prickly lettuce indicates a potential threat to lettuce cultivation in Korea.


