Gastrodia elata is a saprophytic perennial plant [1] in the Achlorophyllous Orchidaceae family [2], which has a mycoheterotrophic lifestyle with reduced leaves and bracts in the scape [3]. The G. elata life cycle depends on two types of fungi: Mycena spp. support seed germination for vegetative propagation, and Armillaria mellea support vegetative growth to develop tubers [4]. G. elata rhizomes (tubers) accumulate valuable secondary metabolic compounds such as gastrodin (GA, p-(hydroxymethyl)
phenyl-β-D-glucopyranoside), vanillyl alcohol, and p-hydroxybenzyl alcohol [5]. It is used as a traditional medicinal herb to treat many human diseases, such as convulsion, vertigo, ischemia, hemiplegia, epilepsy, and tremors [5,6]. G. elata is widely cultivated in northeast Asia, including Korea, China, Japan, and India [7].
Tuber rot disease of G. elata has reduced total yields in Korea by approximately 70% in recent years [8]. Multiple fungal pathogens were reported to infect G. elata in Korea, including Clonostachys rosea, Fusarium solani, and Trichoderma hamatum [9-11], and Fusarium oxysporum caused G. elata root rot during the growing and post-harvest seasons in China [12]. Fusarium pathogens cause various diseases in plants, produce a number of mycotoxins, and are recognized as human pathogens. Fusarium has a significant economic impact on global agricultural and horticultural industries [13]. Fusarium belongs to the Ascomycota phylum and contains a large number of species, which primarily exist in the soil and are related to various plants. These species cause rot diseases on plant bulbs, cobs, corms, roots, seedlings, stalks, and tubers [14]. F. oxysporum causes disease in many important plant species, including banana, bean, cotton, chickpea, melon, tomato, and Arabidopsis [15].
In October 2018, we observed root rot disease (incidence 75% and severity 38%, n=266) on G. elata tubers from four commercial fields in Gimcheon, Korea (36°7′N 128°7′E), and obtained symptomatic tissues, which exhibited soft dark-grey lesions on the tuber surface (Fig. 1A). Pathogens were isolated as follows. The symptomatic tuber was surface-sterilized with 2% sodium hypochlorite (NaOCl) for 2 min, then rinsed twice with sterile distilled water for 1 min [16]. The tubers were dried with sterilized filter paper, and small pieces (5-7 mm diameter) of the lesions were incubated on potato dextrose agar (PDA, Difco, Detroit, MI, USA) in 90-mm Petri dishes for 7 days at 25℃ in the dark. Pure cultures were generated by isolating single spores and culturing on PDA for 10 days. The early-stage colonies were white and became purplish-white with age. The pure culture was used for Koch’s postulates, molecular analysis, and deposited in the Forest Bio Resources Bank, the National Institute of Forest Science (Accession no. NIFoS-FBRB181015-Fo01).
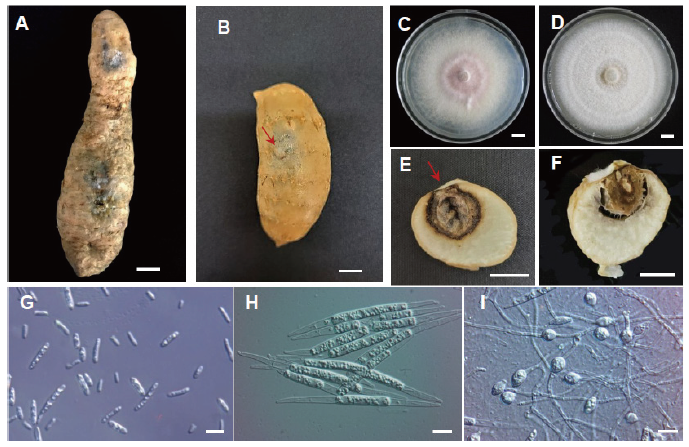
Fig. 1. Root rot caused by Fusarium oxysporum on Gastrodia elata tubers. (A) Symptoms of infected G. elata tubers collected from fields. (B) G. elata tubers inoculated with F. oxysporum developed rot symptoms after 10 days. (C) A 10-day-old colony of F. oxysporum growing on potato dextrose agar medium. (D) A 7-day-old colony of F. solani growing on potato dextrose agar medium. (E) Cross-section of the inoculated tuber in (B). (F) Cross-section of the inoculated tuber with F. solani. (G) microconidia and (H) Macroconidia. (I) Chlamydospores. Scale bar=1 cm (A-F) or 10 μm (G-I). Inoculation point was indicated with red arrow.
We obtained 150-200 mg of mycelia from a 10-day-old culture that was grown on PDA at 25℃ in the dark. The mycelia were ground using a pestle with liquid nitrogen (N2), and genomic DNA was extracted using the AccuPrep Genomic DNA Extraction Kit (K-3032, Bioneer, Daejeon, Korea) according to the manufacturer’s instructions. The internal transcribed spacer (ITS) region of ribosomal DNA was amplified using ITS1/ITS4 primers [17], and translation elongation factor 1-alpha (TEF-1α) was analyzed using EF-1/EF-2 primers [18]. The PCR reactions were performed in a 20 μL mixture containing 5 μL of template genomic DNA, 1 μL each of ITS1/ITS4 or EF-1/EF-2 primer, 0.5 μL of RealTaq DNA polymerase (RTT01, RBC, New Taipei, Taiwan), 0.5 μL of each dNTP, 2 μL of 10× PCR reaction buffer, and 15 μL of distilled water. The PCR amplification was performed in a T100 Thermal Cycler (Bio-Rad, Foster, California, USA) under the following conditions: one cycle of pre-denaturation at 95℃ for 3 min, followed by 35 cycles of denaturation at 95℃ for 33 s, annealing at 58℃ for 1 min, and elongation at 72℃ for 1 min, with one final extension cycle at 72℃ for 5 min. The products were checked on a 1% agarose gel at 100 V for 35 min. The PCR product was sequenced using the ABI 3730xl system (Macrogen, Seoul, Korea). The ITS1/ITS4 primers generated a 472-bp amplicon (GenBank accession no. MW617294), while the EF-1/EF-2 primers generated a 712-bp amplicon (GenBank accession no. OM145982). NCBI BLASTn analysis identified a 100% match with F. oxysporum ITS (KY910846) and TEF-1α (MH341211).
For Koch’s postulates analyses, we incubated the culture on PDA for 10 days at 25℃ in the dark to obtain conidia. One-year-old healthy G. elata tubers harvested from fields were surface-sterilized. Experimentally inoculated tubers were injected with 10 µL of conidial suspension (8×105 conidia/ml) in potato dextrose broth (PDB, Difco, Detroit, MI, USA) (n=10), whereas control disease-free tubers were injected with 10 µL of sterile distilled water (n=10). All tubers were incubated at 25℃ with 80% relative humidity in the dark for 10 days. The rot symptoms on experimentally inoculated tubers were similar to that observed in the field (Fig. 1B), whereas control tubers did not exhibit symptoms. The dark-brown lesion on inoculated tubers developed with variable severity to cover an area of 28-60% of tuber diameter (Fig. 1E). The fungus was then isolated from symptomatic tissues, subjected to analysis, and determined to be 100% identical to F. oxysporum. These combined results satisfy Koch’s postulates. In comparison, F. solani had abundant cottony mycelium with creamy white colored growth (Fig. 1D), and tissues infected with F. solani were softer than those infected with F. oxysporum (Fig. 1F) [10].
The fungal isolate FO2SA was obtained from an infected G. elata tuber. Morphological characterization and rDNA sequencing analysis identified the isolate as F. oxysporum. The fungal colonies were cultured on PDA at 25℃ for 10 days. The colonies were initially white and became bright violet with age (Fig. 1C). Conidiophores were shaped as branched monophialides. Microconidia lacked septa and were oval-ellipsoidal and straight to curved (Fig. 1G). Macroconidia were slightly curved and slender with 1-5 septa pointed at both ends (Fig. 1H). By contrast, other F. oxysporum had 3-5 septa pointed at both ends, and the macroconidia of F. solani isolated from G. elata had 2-5 septa [10]. In the FO2SA isolate, chlamydospores were single and globose-elliopsoidal, whereas other chlamydospores were pairs (Fig. 1I). The morphological characteristics of F. oxysporum infecting G. elata tubers are presented in Table 1.
Table 1. Comparison of morphological characteristics of the Gastrodia elata tuber isolate with Fusarium oxysporum .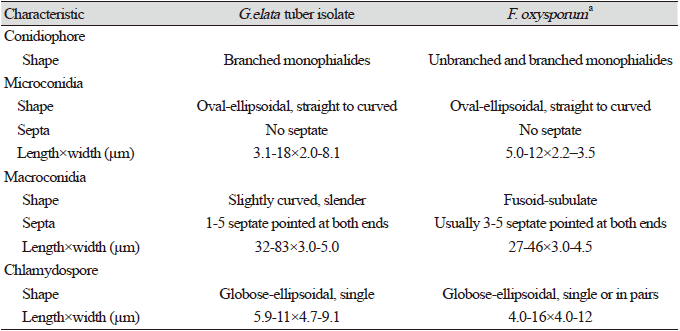
|
Phylogenetic analysis of the combined multilocus (ITS and TEF-1α) was performed using the maximum likelihood method and MEGA X (MEGA Software) [20]. The sequence distance was calculated based on the Tamura-Nei model, and bootstrap analysis was conducted with 1,000 replications. The FO2SA isolate was closely related to three other strains (XC02, ZBM, NRRL 32158) (Fig. 2).
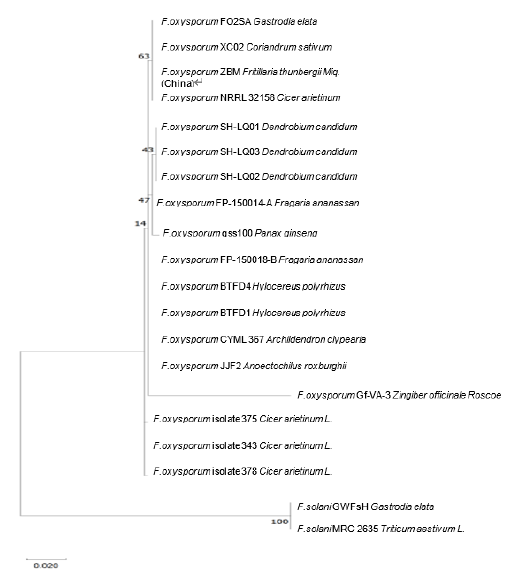

Fig. 2. Phylogenetic analysis using MUSCLE alignment based on a concatenated sequence (internal transcribed spacer [ITS] and translation elongation factor 1-alpha [TEF-1α]) of Fusarium oxysporum isolated from Gastrodia elata with 12 members, and two strains of F. solani (GWFsH and MRC2635) are used as the outgroup. The evolutionary history was inferred using the maximum likelihood method and the Tamura-Nei model. GenBank accession numbers (ITS TEF-1α) of the strains quoted are as follows: FO2SA (MW617294, OM145982), XC02 (MT579855, MT597425), ZBM (MH290452, MZ292517), NRRL32158 (MK074845, FJ538245), SH-LQ01 (JQ809654, JQ809657), SH-LQ03 (JQ809656, JQ809659), SH-LQ02 (JQ809655, JQ809658), FP-150014-A (MK085079, MK086961), gss100 (MH290452, MH341210), FP-150018-B (MK085070, MK086959), BTFD4 (MN727095, MN752124), BTFD1 (MN727096, MN752123), CYML367 (MN826824, MN839682), JJF2 (MN626452, MN629915), Gf-VA-3 (MT337417, MW816632), isolate 375 (MK928423, MN788463), 343 (MK920204, MN788462), 378 (MK928424, MN788464), F. solani GWFsH (KX588131, KX951419), and MRC2635 (MH582403, MH582423), respectively. The scale bar indicates the number of nucleotide substitutions. The alignment and tree were constructed in MEGA X [20]. Values at nodes are based on 1,000 bootstrap replicates.
This study identified F. oxysporum as the causative element of root rot in G. elata tubers in Korea. Our result provides critical information for management strategies to control pathogen fungi in commercial fields of G. elata in Korea.




