Convolvulus arvensis L., commonly known as field bindweed, is a perennial plant in the family Convolvulaceae. The native origin of this plant is Eurasia, however, in recent centuries, its distribution has expanded to most temperate-subtropical regions. In East Asia, it was introduced into Japan in the 1900s and spread to all areas in the 1940s, while its introduction into Korea was recorded in 1980 along the western coast of the Korean peninsula. Nowadays C. arvensis is invasive in all areas of South Korea [1].
In the course of our routine forays to collect phytopathogenic fungi, a previously unknown powdery mildew was found on this plant in Seoul in the summer of 2021 (Fig. 1A). Initially, the infected plants displayed white powdery colonies due to the net-like mycelial mats with an abundance of conidiophores and conidia on both sides of the leaves and stems (Fig. 1B). As the disease progressed, the powdery patches grew larger and denser, causing leaf senescence and early defoliation. No chasmothecia were found until the death of the infected leaves in December 2021.
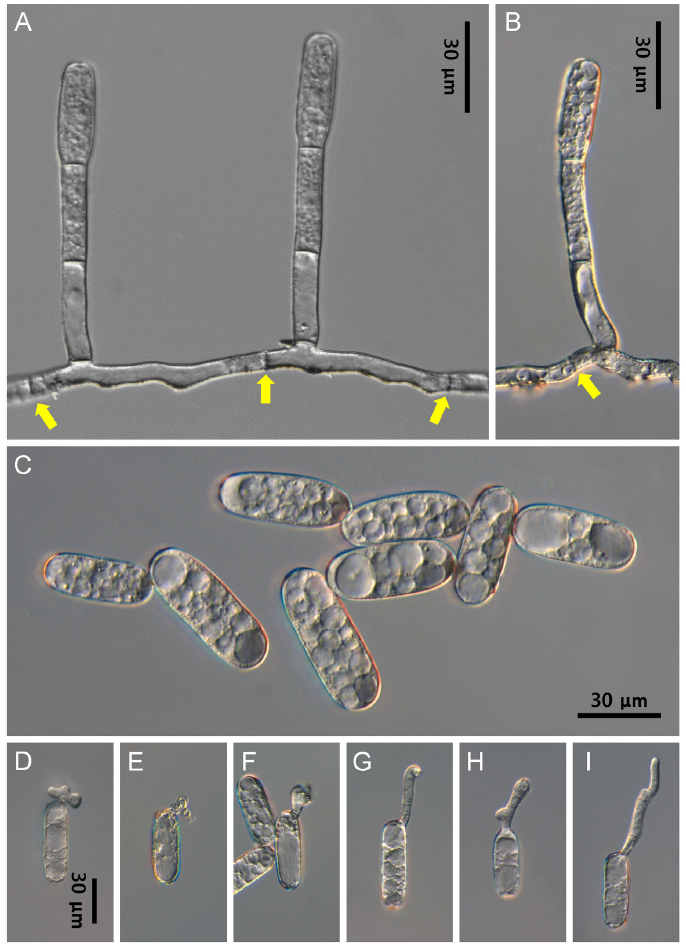
Detailed morphological characteristics of the powdery mildew were observed using an Olympus BX50 microscope (Olympus, Tokyo, Japan). Photomicrographs were taken with a Zeiss AX10 microscope equipped with an AxioCam MRc5 camera (Carl Zeiss, Oberkochen, Germany). The hyphae were superficial, straight to wavy, branched, and 4-7 μm wide. Hyphal appressoria were multi-lobed, single or in pairs, and 3-7 μm wide. Conidiophores are arising from superficial hyphae, solitary, positioned between two hyphal septa usually not central, upright, 54-102×7-9 μm, and composed of 3-4 cells with straight foot cells (Fig. 2A and B). Conidia were formed singly on conidiophores, cylindrical to ellipsoid-cylindrical, 34-54×14-18 μm, vacuolate, and devoid of conspicuous fibrosin bodies (Fig. 2C). Germ tubes were produced on the perihilar position of the conidia and were variable in shape (Figs. 2D-I). These characteristics were consistent with the description of Erysiphe convolvuli DC. [2,3].
To confirm the morphology-based identification, two specimens (KUS-F32396 and F32411, Korea University Herbarium, Seoul, Korea) were used for molecular-phylogenetic analysis. Genomic DNA was extracted from the mycelium using MagListoTM 5M plant genomic DNA extraction kits (BIONEER, Daejeon, Korea) according to the manufacturer’s instructions. The internal transcribed spacer (ITS1 and ITS2) and large subunit (LSU) gene of rDNA were amplified and sequenced with an application of ITS1-F/PM6 and PM3/TW14 primers, respectively [4]. Newly obtained sequences in this study were evaluated using a BLASTN search and deposited in GenBank (Accession Nos. OM033508 and OM033509 for ITS; OM033511 and OM033512 for LSU). The results for ITS and the LSU were 100% identical with sequences from MN203981, MT377709, LC328325 of Erysiphe convolvuli. Phylogenetic analysis was conducted in PAUP* 4.0.b using an alignment consisting of 18 sequences, of which 16 were retrieved from GenBank [5]. Golovinomyces vincae (AB769444) was used as an outgroup taxon. The robustness of a maximum parsimony tree was evaluated using bootstrap (BS) analysis with the application of 1000 replicates. Sequences from our specimens were grouped with sequences of Erysiphe convolvuli in a distinct clade, which is supported by the highest BS value (Fig. 3).
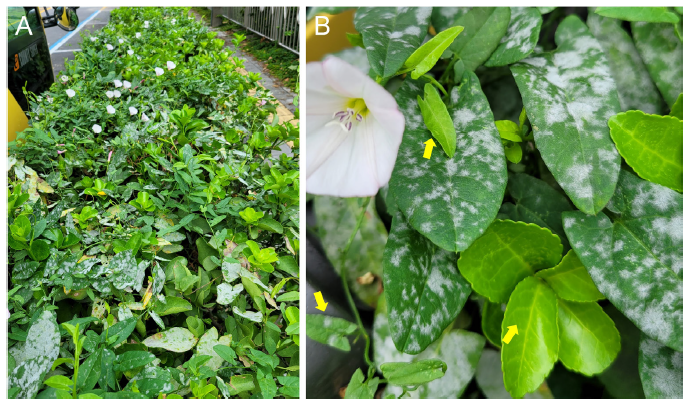

Fig. 1. Powdery mildew caused by Erysiphe convolvuli on Convolvulus arvensis, which is climbing over hedge-grown Euonymus japonicus shrubs on the roadside (A). Close-up view of E. convolvuli infection on C. arvensis growing over E. japonicus (right arrow). Note that young C. arvensis leaves are newly infected (left arrow) and uninfected (upper arrow) (B).
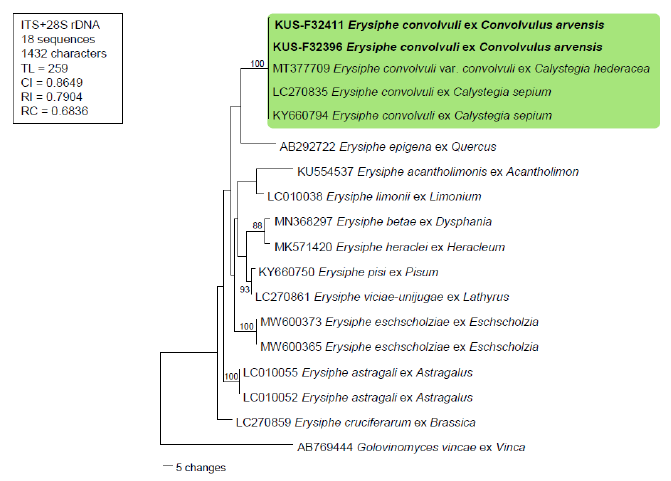
Fig. 3. Maximum parsimony tree based on the combined internal transcribed spacer (ITS)+ large subunit (LSU) sequences of Erysiphe convolvuli and other Erysiphe spp. retrieved from GenBank. The isolates obtained in this study are shown in bold. Bootstrap values (>80%) are indicated on related branches.
Erysiphe convolvuli has been found on C. arvensis plants within its indigenous area, including northern Africa (Canary Islands, Egypt, Libya, and Morocco), western Asia (Afghanistan, western part of China, India, Iran, Iraq, Israel, Lebanon, Pakistan, Saudi Arabia, Azerbaijan, and Turkey), and nearly all of Europe [6]. This powdery mildew species has also been recorded on C. arvensis plants in its invasive area, including China (Beijing) and the United States [7]. Interestingly, there have been no reports of this powdery mildew in Japan [8], suggesting the absence of E. convolvuli in Japan, where five introduced species of Convolvulus plants have been found (http://www.ylist.info). Likewise, E. convolvuli was recently found on Calystegia hederacea in Korea [3]. This is the first report of powdery mildew caused by E. convolvuli on C. arvensis in Korea. This finding could help understand recent invasions and the spread of this powdery mildew.
Acknowledgements
This work was supported by Korea Institute of Planning and Evaluation for Technology in Food, Agriculture and Forestry (IPET) through the Crop Viruses and Pests Response Industry Technology Development Program, funded by the Ministry of Agriculture, Food and Rural Affairs (MAFRA) (Project No. 320043-05). H.D. Shin was supported by a grant (K2123391) from Korea University, Korea.