Abstract
Freshwater fungi are a poly-phylogenetic group of taxonomically diverse organisms. In this study, we isolated diverse fungal strains from various environmental samples obtained from freshwaters in Korea. These strains were identified by performing molecular phylogenetic analyses of rDNA and/or other sequences (beta-tubulin, RNA polymerase II, and translation elongation factor 1). In addition, we examined their morphological characteristics
microscopically and cultural characteristics using different media. We identified eleven unrecorded
Figures & Tables
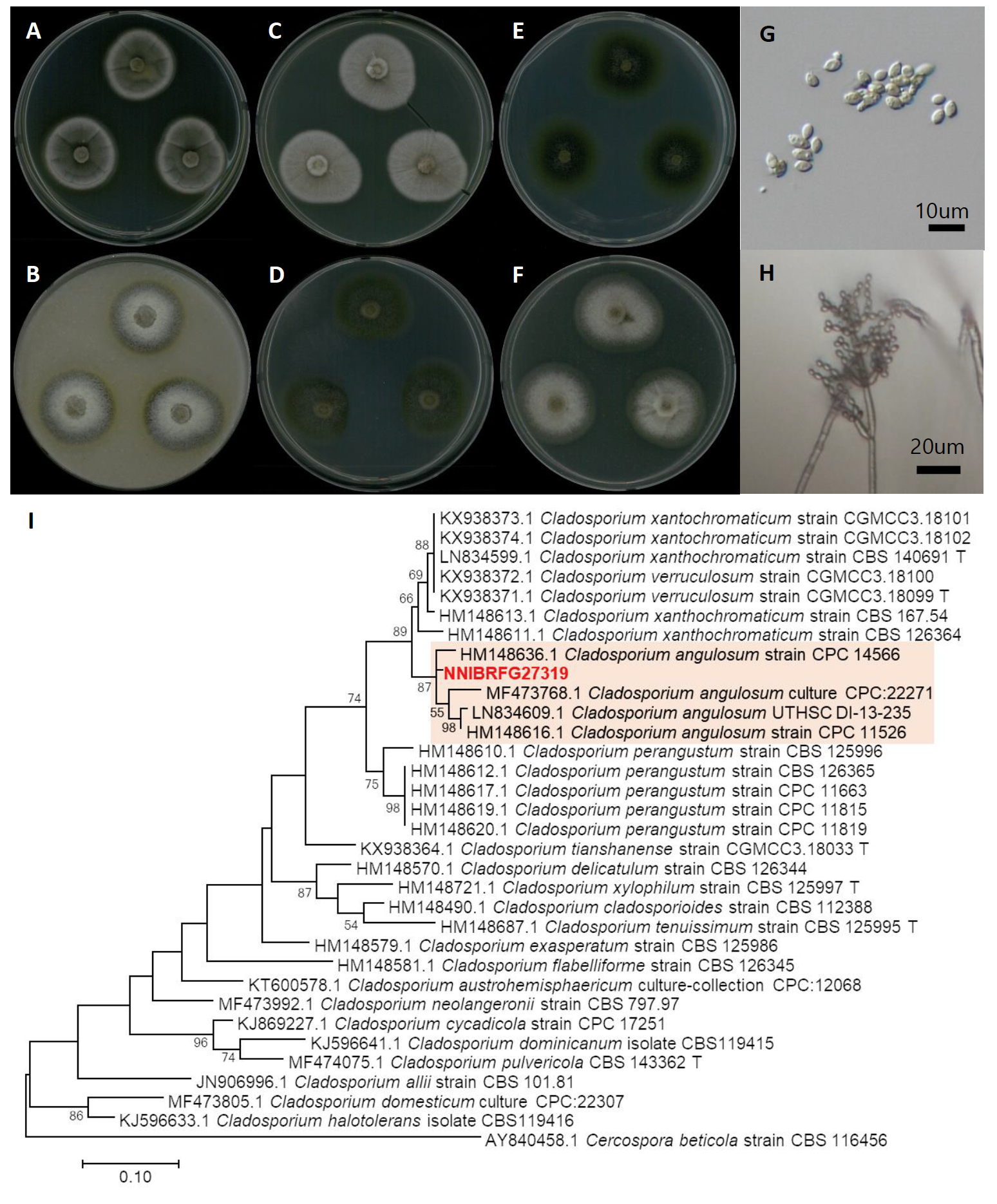
Fig. 1.Characters of NNIBRFG27319. Mycelial growth on (A) potato dextrose agar (PDA), (B) oatmeal agar (OA), (C) potato carrot agar (PCA), (D) malt extract agar (MEA), (E) corn meal dextrose agar (CMDA), (F)V 8-juice agar (V8A) for 7 days at 25℃. Microscopic observation showed (G) conidia and (H) conidiphore (Scale bars: 10 μm). Phylogenetic tree of NNIBRFG27319 and related species, based on maximum-likelihood analysis of (I) the actin gene sequences. Numbers at the nodes indicate the bootstrap values (>50%) from 1,000 replications. The bar indicates the number of substitutions per position. New isolates from the present study are shown in bold. “T” means type material.


