1Apple Research Center, National Institute of Horticultural & Herbal Science, Gunwi 43100, Korea
2Department of Plant Medicine, Kyungpook National University, Daegu 41566, Korea
*Correspondence to heeyoung@knu.ac.kr
Korean Journal of Mycology (Kor J Mycol) 2025 June, Volume 53, Issue 2, pages 79-89.
https://doi.org/10.4489/kjm.2025.53.2.3
Received on April 30, 2025, Revised on May 29, 2025, Accepted on May 30, 2025, Published on June 30, 2025.
Copyright © The Korean Society of Mycology.
This is an Open Access article which is freely available under the Creative Commons Attribution-NonCommercial 4.0 International License (CC BY-NC) (https://creativecommons.org/licenses/by-nc/4.0/).
A fungal strain designated ARI-24-A9 was isolated from ambrosia beetles collected from apple orchards in Korea. The strain was characterized using culture, morphological, and molecular phylogenetic approaches to determine its taxonomic identity. When cultured on potato dextrose agar (PDA) and malt extract agar enriched with 1% Difco yeast extract (YEME) media at 25℃ for 7 days, the colony diameter ranged from 72.3–74.0 mm on PDA and 82.1–85.3 mm on YEME. The colonies exhibited a golden-yellow center with a characteristic growth pattern. Growth condition tests revealed that the optimal temperature and pH for colony development were 30℃ and pH 4–5, respectively. Most hyphae were hyaline with an average width of 2.9 μm, whereas thicker, pale brown hyphae with a width of up to 7.4 μm were observed in aerial mycelium-forming regions. These culture and morphological characteristics were consistent with those of Dryadomyces sulphureus. Molecular analyses using four genetic markers (internal transcribed spacer, ITS; large subunit ribosomal RNA gene, LSU; small subunit ribosomal RNA gene, SSU; β-tubulin, β-TUB) revealed that ARI-24-A9 shared over 99% sequence similarity with D. sulphureus and formed a highly supported clade with this species in both single-locus (ITS) and multilocus (LSU, SSU, β-TUB) phylogenetic trees. These findings confirmed that ARI-24-A9 is conspecific to D. sulphureus. This is the first known instance of D. sulphureus recovered from an ambrosia beetle in Korea.
Ambrosia beetles, Dryadomyces sulphureus, Korean apple orchard, Symbiotic fungi
Among the various symbiotic relationships between insects and fungi, fungal farming has been observed in ants, macrotermites (Macrotermes), and ambrosia beetles (Scolytinae and Platypodinae) [1,2]. Ambrosia beetles typically infest stressed or dead trees, excavating tunnels and inoculating symbiotic fungi. Both larval and adult stages of ambrosia beetles rely on these fungi as their principal diet.
Fungi associated with ambrosia beetles have been reported in at least seven families, with notable genera including Ambrosiella, Fusarium, and Raffaelea. Most of these fungi belong to the phylum Ascomycota, except for Flavodon ambrosius, a basidiomycete (Basidiomycota) symbiont that was identified as a new species isolated from ambrosia beetles by Simmons et al. [3]. Ambrosia beetles cultivate Raffaelea and Ambrosiella species within their galleries and rely on this symbiosis for survival and development [4]. Although most Raffaelea species are saprophytes, R. lauricola, R. quercivora, and R. quercus-mongolicae are pathogenic and cause significant damage to forests and fruit crops [5–7].
Recent phylogenetic studies have led to taxonomic revisions of several species previously classified under Ambrosiella or Raffaelea, reinstating them into the genus Dryadomyces [8,9]. To address the incomplete taxonomy of the order Ophiostomatales, De Beer et al. [10] conducted a phylogenomic re-evaluation, recognizing five species within the genus Dryadomyces: D. amasae, D. montetyi, D. quercivorus, D. quercus-mongolicae, and D. sulphureus. All five species are associated with ambrosia beetles [10].
In this study, a fungal strain was isolated from ambrosia beetles collected from Korean apple orchards and designated ARI-24-A9. To identify this strain, its culture, morphological, and molecular phylogenetic characteristics were analyzed. This study reports the mycological characteristics of ARI-24-A9.
Ambrosia beetles were collected using traps in an orchard at the Apple Research Center in Gunwi-gun, Daegu-si, Korea (36°29′68.9″N, 128°46′56.1″E). The bodies of the collected beetles were surface-sterilized with 70% ethanol for 1 min and thoroughly dried. The beetles were then placed on potato dextrose agar (PDA; Difco, Detroit, MI, USA) plates and incubated at 25℃ for 3 days. The resulting mycelial growth was subcultured on fresh PDA plates and incubated for an additional 8 days at the same temperature. After confirming the absence of contamination in pure cultures, a single fungal strain was successfully isolated and designated ARI-24-A9. The pure culture was preserved in 20% glycerol and stored at −80℃.
Strain ARI-24-A9 was cultured on 90 mm plates containing PDA and malt extract agar enriched with 1% Difco yeast extract (YEME) at 25℃ in the dark to observe its growth rate (temperature and pH dependence), culture characteristics, and morphological features. Key colony traits such as diameter and pigmentation were also recorded. Growth condition testing was conducted on PDA, with temperature settings of 5, 10, 15, 20, 25, 30, and 35℃, and pH conditions ranging from 4 to 9. Additionally, the morphology of the hyphae, conidia, and conidiophores was examined using a light microscope (CX-43; Olympus, Tokyo, Japan), and their dimensions were measured and documented.
For the molecular and phylogenetic analyses of the ARI-24-A9 strain, genomic DNA was extracted using a HiGene Genomic DNA Preparation Kit (Biofact, Daejeon, Korea). Partial sequences of the internal transcribed spacer (ITS) region, large subunit ribosomal RNA gene (LSU), small subunit ribosomal RNA gene (SSU), and β-tubulin (β-TUB) genes were amplified using PCR.
The ITS region was amplified using ITS1F/ITS4 primers [11,12], LSU with LROR/LR7 primers [13], SSU with NS1/NS4 primers [12], and β-TUB with Bt2a/Bt2b primers [14]. The PCR products were verified by electrophoresis on a 1% agarose gel and staining with ethidium bromide. They were then purified using EXOSAP-IT reagent (Thermo Fisher Scientific, Waltham, MA, USA). Sequencing was performed by Solgent (Daejeon, Korea), and the sequences were analyzed using SeqMan Lasergene software (DNAStar, Madison, WI, USA). The nucleotide sequences obtained for each genetic marker were registered in GenBank under accession numbers PV465310 (ITS), PV465507 (LSU), PV465508 (SSU), and PV476179 (β-TUB).
A phylogenetic tree was constructed to determine the taxonomic position of ARI-24-A9. Nucleotide sequences of species from the genus Dryadomyces used for phylogenetic analysis were retrieved from the National Center for Biotechnology Information (NCBI) database. Sequence alignment was performed using Clustal X 2.0, integrated into MEGA 7 software [15].
Phylogenetic trees were constructed based on the ITS region sequence and concatenated sequences of the LSU, SSU, and β-TUB genes. For the analysis, Kimura’s two-parameter model [16] was applied along with the nearest neighbor interchange method, excluding gaps. Trees were generated using the maximum likelihood (ML) method [17] and bootstrap analysis with 1,000 replicates was conducted to assess its reliability.
When cultured on YEME medium at 25℃ for 7 days, the colony diameter ranged from 82.1 to 85.3 mm. On the front side, the colony exhibited a golden-yellow center, which gradually faded to white toward the margins. Aerial hyphae were observed in scattered areas and were often accompanied by droplets containing brownish pigments. On the reverse side, the colony had a deep yellow center that became progressively paler toward the edges and appeared nearly white. The culture emitted a distinct sweet odor, reminiscent of fruit (Fig. 1A, 1B). Under the same conditions (25℃, 7 days) on PDA medium, the colony diameter ranged from 72.3 to 74.0 mm, and the cultural characteristics were nearly identical to those observed on YEME medium.
In growth tests conducted on PDA at different temperatures and pH levels, colony diameters were measured on days 5 and 7 of incubation (Fig. 2A). Under these temperature conditions, growth was minimal at 5℃ and 10℃. From 15℃ onward, colony growth increased gradually with temperature, reaching the highest growth at 30℃ with a colony diameter of 81.0 mm, indicating that 30℃ was the optimal growth temperature for this strain. In contrast, at 35℃, growth was significantly reduced, with a colony diameter of only 40.0 mm, and the colony morphology was too weak to distinguish any clear culture features (Fig. 2B).
In the pH-dependent growth test, the largest colony diameter was observed at pH 4, reaching 79.0 mm. As the pH increased, the growth rate gradually decreased, indicating that the optimal pH range for this strain was 4–5 (Fig. 2C).
Hyphae were generally hyaline with thin cell walls, showing frequent local swelling and the formation of a densely interwoven structure. The average hyphal width was 2.9 μm (n = 30) (Fig. 1C).
In addition, some hyphae within the clusters forming aerial mycelium exhibited thicker cell walls and broader widths (7.4 µm), appeared brown to pale brown, and showed branching at locally swollen regions (Fig. 1D, 1E). These hyphae were presumed to play a supporting role in the formation of aerial structures. Conidiophores and conidia were not observed on PDA or YEME.
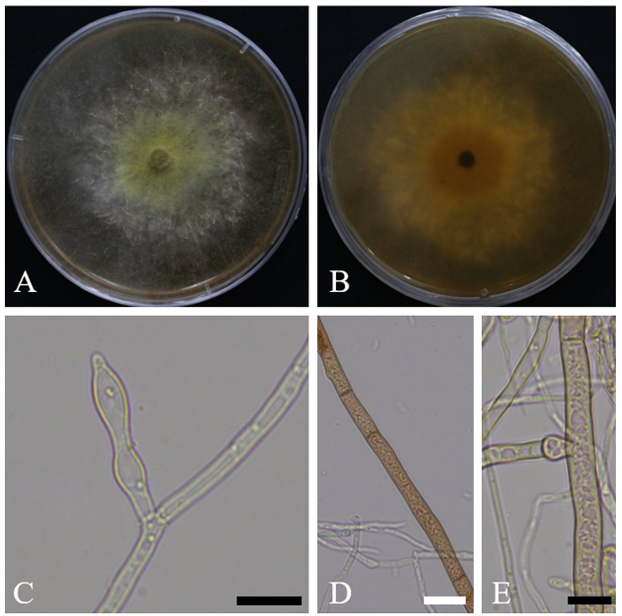
Fig. 1. Cultural and morphological characteristics of ARI-24-A9 (Dryadomyces sulphureus). A, B: colony on malt extract agar enriched with 1% Difco yeast extract (YEME) after 7 days of growth at 25°C; C: swollen regions within hyphae; D: Aerial hyphae; E: Branching at locally swollen regions along aerial hyphae. Scale bars: C–E = 10 μm, D = 20 μm.
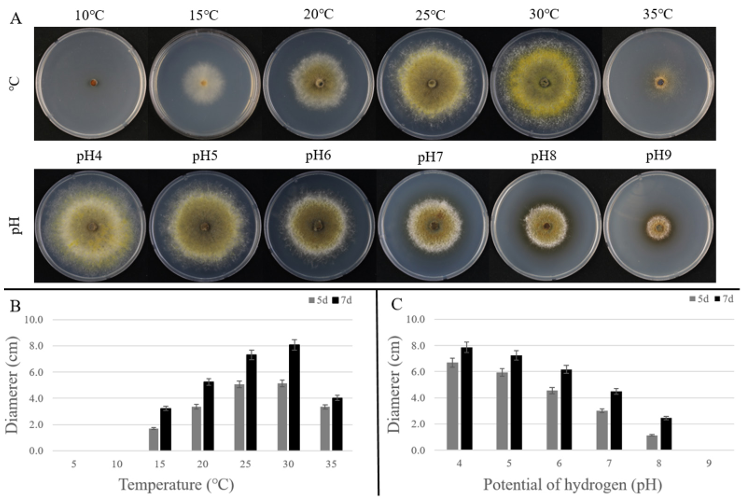
Fig. 2. Growth of ARI-24-A9 on potato dextrose agar (PDA) under different temperature and pH conditions. A: Colony morphology after 7 days of incubation at temperatures (10–35°C) and pH levels (pH 4–9); B: Colony diameters measured at 5 and 7 days under different temperature conditions; C: Colony diameters measured at 5 and 7 days under different pH conditions.
The culture and morphological characteristics of strain ARI-24-A9 were consistent with those of Dryadomyces sulphureus. Although D. sulphureus has been reported to be most closely related to D. amasae, strain ARI-24-A9 showed clear morphological differences from D. amasae. Aerial and immersed hyphae were observed in both species, and the morphology of the hyphae was similar. However, a distinct difference in coloration was observed. Although D. amasae hyphae changed from hyaline to dark, ARI24-A9 hyphae transitioned from hyaline and appeared pale brown. Differences in culture characteristics were also observed. ARI-24-A9 formed a golden-yellow colony, whereas D. amasae showed a yellowishgreen coloration (Table 1).
Table 1. A comparative analysis of the mycological characteristics of ARI-24-A9 and its closely related species in the genus Dryadomyces.
테이
a Fungal strain studied in this paper; b Source of description [4]; c Sources of description [8].
YEME, malt extract agar enriched with 1% Difco yeast extract; PDA, potato dextrose agar; N/A, not available.
To analyze the molecular and phylogenetic relationships of ARI-24-A9, four molecular markers (ITS region, LSU, SSU, and β-TUB genes) were utilized. The obtained sequence lengths were 609, 1,305, 1,020, and 325 bp, respectively.
A BLAST (Basic Local Alignment Search Tool) search was conducted to compare these sequences with those of other strains in the NCBI database. For the ITS region, ARI-24-A9 showed the highest similarity (99.8%) with D. sulphureus CBS 380.68. The LSU gene sequence of ARI-24-A9 exhibited 99.8% similarity with D. sulphureus CBS 380.68, 97.3% similarity with D. quercivorus CBS 122982, and 96.8% similarity with D. quercus-mongolicae KACC 44405. For the SSU gene sequence, ARI-24-A9 exhibited 100.0% similarity with D. sulphureus CBS 380.68, and 99.5% similarity with D. quercivorus PC10.921, D. amasae C2750, and D. quercivorus C2526. Additionally, it displayed 99.4% similarity with D. quercusmongolicae C3013. Finally, the β-TUB gene sequence of ARI-24-A9 showed 99.6% similarity with D. sulphureus CBS 132735 and 99.1% similarity with D. montetyi P105.
This analysis confirmed that ARI-24-A9 showed the highest similarity to D. sulphureus across all four genetic markers (ITS, LSU, SSU, and β-TUB). Notably, SSU exhibited 100% identity, whereas the other genes showed > 99% similarity.
To evaluate the phylogenetic relationships of ARI-24-A9, a phylogenetic tree was constructed based on the ITS region sequence alone (Fig. 3), and multilocus sequence analysis (MLSA) was performed using
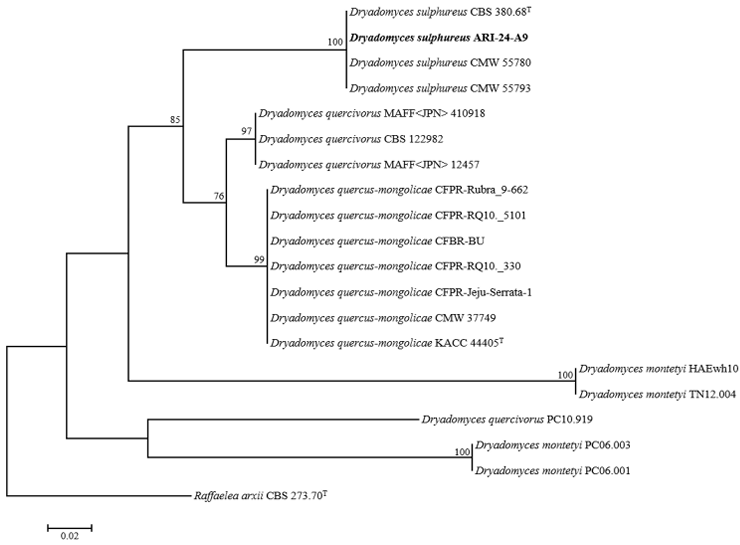
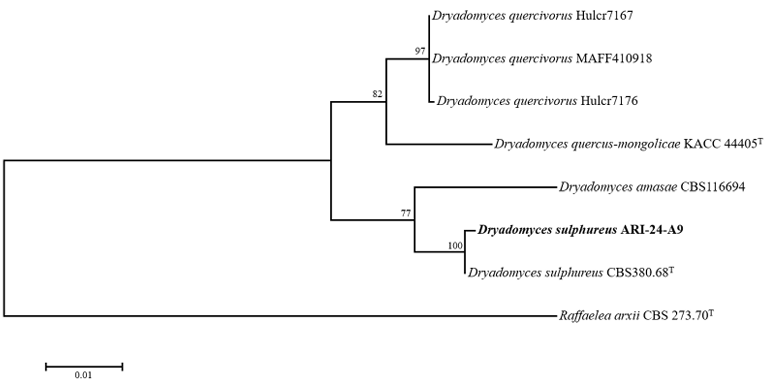
concatenated sequences of the LSU, SSU, and β-TUB genes (Fig. 4). The phylogenetic tree constructed using the ML method confirmed that ARI-24-A9 shared the same phylogenetic position as D. sulphureus and was clearly distinguished from other species. Based on comprehensive molecular and phylogenetic analyses, ARI-24-A9 was conclusively identified as identical to D. sulphureus at the species level.
The genus Dryadomyces was first described by Gebhardt et al. [8] with D. amasae as the type species. The genus name derives from “Dryads”, the tree nymphs in Greek mythology, reflecting the habitat of these fungi in woody plants. Species within Dryadomyces are associated with ambrosia beetles and currently include five species, which were previously classified under Ambrosiella or Raffaelea [8,9]. Harrington et al. [18] noted that, although D. amasae exhibited morphological differences in conidia compared to other Raffaelea species, the classification of ambrosia-associated fungi within Ophiostomatales remains unresolved. Therefore, they proposed that all such fungi should be retained within Raffaelea until further studies provide more clarity. Subsequently, Harrington et al. [19] maintained D. amasae within Raffaelea and asserted that morphological differences alone were insufficient to justify a separate genus. As a result, species of Dryadomyces were classified under Raffaelea. However, De Beer and Wingfield [9] identified a distinct lineage comprising Raffaelea amasae, R. montetyi, R. sulphurea, R. quercus-mongolicae, and R. quercivora, designating it the R. sulphurea complex. Phylogenetic analyses suggested that this complex might belong to Leptographium sensu lato; however, the data at that time were insufficient to determine whether it was part of Leptographium sensu lato or represented a separate lineage. Further studies by Dreaden et al. [20] assessed the monophyly of Raffaelea and concluded that the R. sulphurea complex should be excluded from Raffaelea. Their analyses revealed that this complex clustered closely with Esteya vermicola within Leptographium sensu lato. However, the authors emphasized the need for broader phylogenetic studies incorporating additional Leptographium and Raffaelea species to resolve its classification. This led to continued debate regarding whether these species should be integrated into Leptographium sensu lato or reclassified under a reinstated Dryadomyces with D. amasae as the type species. De Beer et al. [10] conducted morphological and molecular analyses to confirm that Raffaelea amasae, R. montetyi, R. sulphurea, R. quercus-mongolicae, and R. quercivora form a distinct monophyletic lineage separate from Leptographium and Raffaelea. As a result, the authors determined that reinstating Dryadomyces was the most appropriate taxonomic decision. The genus definition was revised to encompass these five species, providing robust evidence for the independent classification of Dryadomyces as a distinct lineage.
In the present study, the fungal strain ARI-24-A9 was isolated from an ambrosia beetle collected from an apple orchard in Korea. BLAST analyses of the ITS region, LSU, SSU, and β-TUB gene sequences confirmed that the strain belongs to the genus Dryadomyces, and is most closely related to D. sulphurea. A single-gene phylogenetic tree based on ITS sequences was constructed to determine the phylogenetic position of ARI-24-A9 among the five species recently reclassified under Dryadomyces. However, the ITS sequence of D. amasae was not available in the NCBI database, which prevented a direct comparison. Nonetheless, phylogenetic analyses using LSU, SSU, and β-TUB sequences revealed a clear distinction between ARI-24-A9 and D. amasae. Furthermore, ARI-24-A9 formed a distinct clade with D. sulphurea, as supported by high bootstrap values, and was distinct from the other species. These findings are consistent with those of Dreaden et al. [20], who reported that the R. sulphurea complex is phylogenetically distinct from Raffaelea based on LSU, SSU, and β-TUB sequences data.
Although various culture conditions were used to induce the formation of conidiophores and conidia in the ARI-24-A9 strain, no spore production was observed. Nevertheless, the culture characteristics and hyphal morphology of this strain were consistent with those of D. sulphurea and also showed partial similarity to D. amasae. However, distinct differences were observed in the colony pigmentation between ARI-24-A9 and D. amasae, and phylogenetic analyses placed them in separate clades. These findings strongly suggest that ARI-24-A9 is conspecific to D. sulphurea based on both morphological and molecular phylogenetic evidence.
The morphological features of D. sulphurea were first described in 1967 when the species was introduced by Batra. However, the study did not include detailed morphological data. Photomicrographs of the conidial structures were not provided, and the characteristics are represented only through line drawings. This limitation raises the question of whether conidial formation can be induced under artificial culture conditions.
In conclusion, this study presents the first report of a symbiotic fungus belonging to the genus Dryadomyces isolated from an ambrosia beetle in Korean apple orchards. This finding represents a significant expansion of the geographical distribution of D. sulphurea. Despite extensive efforts to induce conidial formation under various culture conditions, no spores have been observed, suggesting that the reproductive structures and mechanisms of this species remain poorly understood. Further research is necessary to clarify not only its ecological role but also the environmental factors required for conidial development and its interactions with host beetles.
The authors declare that they have no potential conflicts of interest.
This study was supported by the “Cooperative Research Program for Agriculture Science and Technology Development (Project No. PJ017183)” funded by the Rural Development Administration, Republic of Korea.
1. Mueller UG, Gerardo NM, Aanen DK, Six DL, Schultz TR. The evolution of agriculture in insects. Annu Rev Ecol Evol Syst 2005;36:563-95. [DOI]
2. Biedermann PHW, Vega FE. Ecology and evolution of insect-fungus mutualisms. Annu Rev Entomol 2020;65:431-55. [DOI]
3. Simmons DR, Li Y, Bateman CC, Hulcr J. Flavodon ambrosius sp. nov., a basidiomycetous mycosymbiont of Ambrosiodmus ambrosia beetles. Mycotaxon 2016;131:277-85. [DOI]
4. Batra LR. Ambrosia fungi: A taxonomic revision, and nutritional studies of some species. Mycologia 1967;59:976-1017. [DOI]
5. Kim KH, Choi YJ, Seo ST, Shin HD. Raffaelea quercus-mongolicae sp. nov. associated with Platypus koryoensis on oak in Korea. Mycotaxon 2009;10:189-97. [DOI]
6. Kubono T, Ito S. Raffaelea quercivora sp. nov. associated with mass mortality of Japanese oak, and the ambrosia beetle (Platypus quercivorus). Mycoscience 2002;43:255-60. [DOI]
7. Ploetz RC, Hulcr J, Wingfield MJ, de Beer ZW. Destructive tree diseases associated with ambrosia and bark beetles: Black swan events in tree pathology? Plant Dis 2013;97: 856-72. [DOI]
8. Gebhardt H, Weiss M, Oberwinkler F. Dryadomyces amasae: A nutritional fungus associated with ambrosia beetles of the genus Amasa (Coleoptera: Curculionidae, Scolytinae). Mycol Res 2005;109:687-96. [DOI]
9. De Beer ZW, Wingfield MJ. Emerging lineages in the Ophiostomatales. In: Seifert KA, De Beer ZW, Wingfield MJ, editors. The Ophiostomatoid fungi: Expanding frontiers, CBS Biodiversity Series, vol. 12. Utrecht, The Netherlands: CBS-KNAW Fungal Biodiversity Centre; 2013. p. 21-46.
10. De Beer ZW, Procter M, Wingfield MJ, Marincowitz S, Duong TA. Generic boundaries in the Ophiostomatales reconsidered and revised. Stud Mycol 2022;101:57-120. [DOI]
11. Gardes M, Bruns TD. ITS primers with enhanced specificity for basidiomycetes – application to the identification of mycorrhizae and rusts. Mol Ecol 1993;2:113-8. [DOI]
12. White TJ, Bruns T, Lee S, Taylor J. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ, editors. PCR protocols: A guide to methods and applications. New York: Academic Press; 1990. p. 315-22. [DOI]
13. Vilgalys R, Hester M. Rapid genetic identification and mapping of enzymatically amplified ribosomal DNA from several Cryptococcus species. J Bacteriol 1990;172:4238-46. [DOI]
14. Glass NL, Donaldson GC. Development of primer sets designed for use with the PCR to amplify conserved genes from filamentous ascomycetes. Appl Environ Microbiol 1995;61:1323-30. [DOI]
15. Kumar S, Stecher G, Tamura K. MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 2016;33:1870-4. [DOI]
16. Kimura M. A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 1980;16:111-20. [DOI]
17. Fitch WM. Toward defining the course of evolution: Minimum change for a specific tree topology. Syst Biol 1971;20:406-16. [DOI]
18. Harrington T, Fraedrich S, Aghayeva D. Raffaelea lauricola, a new ambrosia beetle symbiont and pathogen on the Lauraceae. Mycotaxon 2008;104:399-404. [DOI]
19. Harrington TC, Aghayeva DN, Fraedrich SW. New combinations in Raffaelea, Ambrosiella, and Hyalorhinocladiella, and four new species from the redbay ambrosia beetle, Xyleborus glabratus. Mycotaxon 2010;111:337-61. [DOI]
20. Dreaden TJ, Davis JM, De Beer ZW, Ploetz RC, Soltis PS, Wingfield MJ, Smith JA. Phylogeny of ambrosia beetle symbionts in the genus Raffaelea. Fungal Biol 2014;118:970-8. [DOI]