1Korean Agricultural Culture Collection, Agricultural Microbiology Division, National Institute of Agricultural Sciences, RDA, Wanju 55365, Korea
2Industrial Insect Division, National Institute of Agricultural Sciences, RDA, Wanju 55365, Korea
*Correspondence to funguy@korea.kr
Korean Journal of Mycology (Kor J Mycol) 2025 June, Volume 53, Issue 2, pages 91-100.
https://doi.org/10.4489/kjm.2025.53.2.4
Received on April 10, 2025, Revised on June 10, 2025, Accepted on June 10, 2025, Published on June 30, 2025.
Copyright © The Korean Society of Mycology.
This is an Open Access article which is freely available under the Creative Commons Attribution-NonCommercial 4.0 International License (CC BY-NC) (https://creativecommons.org/licenses/by-nc/4.0/).
The genus Cordyceps, a group of entomopathogenic fungi, comprises approximately 750 identified species, mainly distributed in Asia, Europe, and North America. During our investigation in 2024, fungal strains were isolated from larvae of Lepidoptera infected with entomopathogenic fungi on a Perilla farm in Korea. Fungal strains from the infected larvae were isolated and identified as Cordyceps koratensis based on their morphological traits and multi-locus phylogenetic analysis of the internal transcribed spacer (ITS), large subunit ribosomal RNA (LSU), translation elongation factor 1-alpha (TEF), and the largest (RPB1) and second-largest (RPB2) subunits of RNA polymerase II loci. To our knowledge, this is the first report of C. koratensis infection in Korea. These findings may support the development of biocontrol agents within the framework of eco-friendly pest management for C. koratensis.
Cordyceps koratensis, Entomopathogenic fungi, Identification
Entomopathogenic fungi (EPF) are microbiological insecticides with a broad host range [1,2]. Some EPF, such as Beauveria bassiana (Balsamo) Vuill (Hypocreales: Cordycipitaceae), Cordyceps fumosorosea (Wize) Kepler, B. Shrestha, Spatafora (Hypocreales: Cordycipitaceae), and Cordyceps javanica (Frieder. & Bally) Kepler, B. Shrestha & Spatafora, have been commercially produced as biocontrol agents [3,4]. Infection begins with the attachment of conidia or blastospores to the host cuticle, followed by the growth and penetration of the fungus through the host integument [5]. Fungal growth blocks the host’s digestive and circulatory systems and produces toxins, leading to host death; eventually, aerial conidia are formed on the cadaver and disperse to infect new hosts [6,7]. However, the virulence of fungal strains may vary considerably both between and within species [8,9].
Cordyceps is a well-known genus of entomopathogenic fungi, with at least 700 known species. Cordyceps species reproduce through sexual (ascospores) or asexual (conidia) spores or both [10]. The name Cordyceps derives from the Greek term “kordyle”, which means “club”, and the Latin etymon “ceps”, which means “head” [11]. Cordyceps species invade insects and arthropods, and evade the host immune system by harmonizing the life cycle of their hosts with the intention of survival and multiplication. Many Cordyceps species play a significant role in the cycling of matter in ecological systems, have high environmental and economic value for biocontrol and bioactive compounds, and serve as a model system for research on fungal-insect pathology [12,13]. Known hosts of Cordyceps span seven orders of Arthropoda: Araneae, Coleoptera, Dermaptera, Hemiptera, Hymenoptera, Lepidoptera, and Orthoptera. Among them, Coleoptera and Lepidoptera are the two most significant host orders beyond the estimated 200 Cordyceps species [14–17].
A molecular phylogenetic investigation of Clavicipitaceae, with an emphasis on Cordyceps, was conducted by Sung et al. [18] and revealed that both Clavicipitaceae and Cordyceps were not monophyletic. Two additional families, Cordycipitaceae and Ophiocordycipitaceae, were recognized, and species previously classified as Cordyceps were supported as members of all three families. Clavicipitaceae and Ophiocordycipitaceae collectively formed a monophyletic group, whereas Cordycipitaceae, defined by the phylogenetic position of the type species of Cordyceps, C. militaris, shared a more recent common ancestor with Hypocreaceae [15]. The classification of Cordyceps species has been conventionally based on morphology [18]. However, significant changes in the taxonomy of Cordyceps have occurred since research on EPF entered the molecular era. Currently, multilocus phylogenetic analyses have gained importance for delimiting Cordyceps species [17,19–22].
In 2024, an EPF Lepidoptera larva was discovered in a Perilla field in Korea. The infected specimens were completely covered with mycelia, suggesting the presence of a potentially novel EPF species that was previously unreported in Korea. This study aimed to identify the fungal species using morphological characteristics and to provide the first report of this EPF species in Korea.
Lepidopteran larvae infected with an entomopathogenic fungus were collected from a perilla (Perilla frutescens L. Britton) farm in Gwangju, Korea (N 35.185835° E 126.783697°). Isolation was performed using a tissue transplantation technique. The samples were washed with sterile distilled water and sectioned into three pieces. The inner tissues of larvae were cut (5 × 5 mm2), surface-sterilized by dipping in 1% sodium hypochlorite (NaOCl) for 1 min, and rinsed several times with sterile distilled water before being transferred onto the surface of potato dextrose agar (PDA). The mycelium growing out of the moth larvae tissue was sub-cultured on PDA and incubated at 25℃ for 14 days. All single-spore cultures were transferred to PDA slants and preserved at 4℃. One of the isolated strains was deposited at the Korean Agricultural Culture Collection (KACC410975).
Fungal mycelia were scraped from 14-day-old cultures grown on PDA plates incubated at 25℃. Approximately 50 mg of fresh mycelia was used for DNA extraction using the DNeasy® Plant Mini Kit (Qiagen, Hilden, Germany), following the manufacturer’s instructions.
The primer pairs ITS5/ITS4, LROR/LR5, EF1-983F/EF1-2218R, RPB1AF/RPB1C, and RPB2-5f2/ RPB2-7cr were used to amplify the internal transcribed spacer (ITS), large subunit ribosomal RNA (LSU), translation elongation factor 1-alpha (TEF), and the largest (RPB1) and second-largest (RPB2) subunits of RNA polymerase II, respectively (Table 1). Each polymerase chain reaction (PCR) volume (25 μL) consisted of 12.5 μL MyTaq HS Mix, 1 μL (10 pmol) of each primer, 8.5 μL nuclease-free water, and 2 μL DNA template (100 ng/μL). PCR reactions were performed in an MJ Research PTC-200 Thermal Cycler (MJ Research, Ramsey, MN, USA) with an initial denaturation step at 94°C for 5 min, followed by 30 cycles: denaturation at 94°C for 30 s; annealing at 52℃ (ITS, LSU), 56℃ (RPB1, RPB2), and 54℃ (TEF) for 30 s; extension at 72°C for 1 min; and a final extension at 72°C for 10 min. PCR products were checked by gel electrophoresis before being sent to Macrogen (Seoul, Korea) for sequencing using the amplified primer pairs.
Table 1. Primer pairs used for PCR amplification and sequencing in this study
테이블
ITS, internal transcribed spacer; LSU, large subunit ribosomal RNA; TEF, translation elongation factor 1-alpha, RPB1, largest subunits of RNA polymerase II; RPB2, second-largest subunits of RNA polymerase II.
The sequence datasets from each locus were individually aligned using the multiple sequence alignment program MAFFT version 11 (https://mafft.cbrc.jp/alignment/server/) with the G-INS-1 option. Poor terminal alignments were trimmed and further refined by visual inspection using MEGA 11 [23], followed by concatenation. Subsequently, a maximum likelihood (ML) phylogenetic tree was constructed using concatenated ITS, LSU, TEF, RPB1, and RPB2 sequences. IQ-TREE was used for phylogenetic analysis, and the model employed was the best-fit model “TIM2 + F + I + G4” with 1,000 ultrafast bootstrap replicates. The sequence datasets included sequences from KACC 410975, 34 reference species in the genus Cordyceps, and the outgroup (Simplicillium lanosoniveum CBS 704.86). Information on the reference sequences is provided in Table 2. All sequences obtained in this study were deposited in RDAGenBank (http://genebank.rda.go.kr).
Table 2. List of species and GenBank accession numbers of sequences used in this study
테이블
ITS, internal transcribed spacer; LSU, large subunit ribosomal RNA; TEF, translation elongation factor 1-alpha, RPB1, largest subunits of RNA polymerase II; RPB2, second-largest subunits of RNA polymerase II.
The accession numbers marked with T refer to sequences from ex-type strain. The sequences generated in this study are in bold.
For morphological observation, the strain was grown on PDA and incubated at 25℃ in the dark for 14 days. At the end of the incubation period, initial investigations included recording the colony diameters and specific characteristics. Colonies grown on PDA were examined microscopically. Slides were prepared by mounting with lactic acid and were observed under a Zeiss AXIO Imager A1 microscope with differential interference contrast (DIC) illumination equipped with a digital AxioCam ICc3 camera (Carl Zeiss, Oberkochen, Germany). The recorded microscopic characteristics included size, shape, conidial pigmentation, and conidiogenic properties.
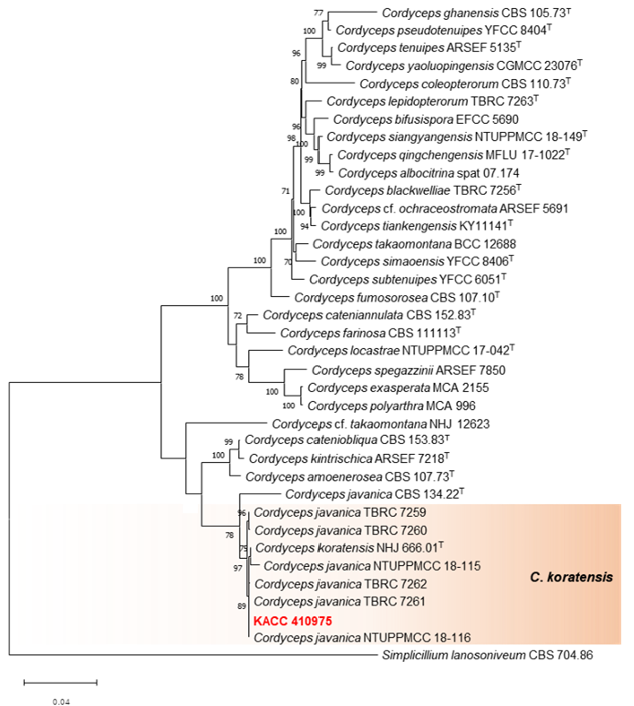
The phylogenetic position of strain KACC 410975 was analyzed using the concatenated sequence data of the ITS, LSU, TEF, RPB1, and RPB2 loci (Fig. 1). The concatenated dataset alignment contained 4034 characters. The concatenated alignment consisted of 555 characters from ITS, 860 from LSU, 1006 from TEF, 711 from RPB1, and 902 from RPB2. During the BLASTn analyses of the sequences, the KACC 410975 sequence of ITS, LSU, and TEF showed 100% similarity with C. koratensis NHJ 666.01T.
Phylogenetic analysis indicated that the isolate KACC 410975 was well clustered with the ex-type strain NHJ 666.01 of C. koratensis, supported by an ML bootstrap value of 97%, and was a sister of C. javanica.
The phylogenetic tree indicated that strain KACC 410975, an ex-type strain of C. koratensis (NHJ 666.01), and some strains (NTUPPMCC 18-115, NTUPPMCC 18-116, TBRC 7259, TBRC 7260, TBRC 7261, and TBRC 7262) previously regarded as C. javanica [26] were clustered in the same clade, supported by an ML bootstrap value of 97%. They are genetically distinct from C. javanica (ex-type strain CBS 134.22).
The morphological characteristics of KACC 410975 resembled those of C. koratensis (syn. Parahevansia koratensis, Hevansia koratensis, Akanthomyces koratensis) described by Hywel-Jones [24], as follows: phialides in a monolayer, single on basal lateral cells of synnemata, crowded, obovoid to ellipsoid with distinct necks.
Based on molecular phylogenetic and morphological analyses, the causative agent in lepidopteran larvae was identified as C. koratensis. This represents the first confirmed report of this species in Korea.
Cordyceps koratensis (Hywel-Jones) H.A. Ariyaw, M. Stadler, and Luangsa-ard (Figure 2) [MycoBank no.: 851851]
Colony morphology: Colonies on PDA reached 35–42 mm in diameter in 14 days at 25℃. Exhibited an entire margin and produced white mycelium with grey sporulation, becoming grey white, with a white to pale yellow reverse.
Micromorphology: Conidiophores arose from prostrate or aerial hyphae, consisting of solitary phialides along the hyphae or verticillate branches with two–four phialides in the whorls. Phialides were flask-shaped, measuring (6–)8–11(–13) × (2–)2.5–3(–3.5) μm, with an ovoid basal portion slightly tapering into a neck, 1 μm in width. Conidia were cylindrical to ellipsoidal, unicellular, and hyaline, measuring (3–)3.5–6(–7) ×(1–)1.5–2(–2.5) μm.
Strain examined: KACC 410975
Diagnosis: C. koratensis is morphologically characterized by obovoid-to-ellipsoid phialides, whereas KACC 410975 produces flask-shaped phialides, similar to C. javanica.
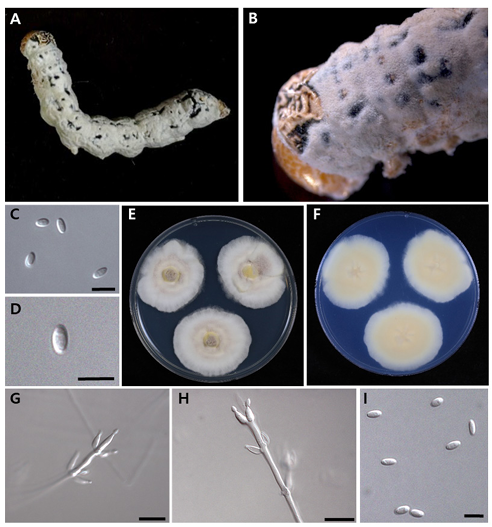
Fig. 2. Morphology of Cordyceps koratensis (KACC 410975). A, B: Mycelium arising from larva of Lepidoptera. C, D: Conidia on the host. E: Obverse of colonies on potato dextrose agar (PDA) at 14 d. F: Reverse of colonies on PDA at 14 d. G, H: Phialides on PDA. I: Conidia on PDA culture. Scale bars: C, D = 5 µm, G, H = 10 µm, I = 5 µm.
We aimed to identify an entomopathogenic fungus infecting the larvae of Lepidoptera based on a combination of morphological characteristics and multi-locus sequence analysis. Cordyceps koratensis was originally described as Akanthomyces koratensis [24], later transferred to Hevansia koratensis [15] and then to Parahevansia koratensis [25]. Mongkolsamrit et al. [25] concluded that two strains of P. koratensis (NHJ 2662 and NHJ 666.01) formed an independent clade with strong support from the Hevansia clade and in proximity to Cordyceps species. The morphology of C. koratensis described by Hywel-Jones [24] exhibited significant variation compared to strains NTUPPMCC 18-115, NTUPPMCC 18-116, TBRC 7259, TBRC 7260, TBRC 7261, and TBRC 7262, previously identified as C. javanica by Chuang et al. [26]. However, the phylogenetic classification of cordycepitoid fungi showed that most diagnostic characteristics used in the current classifications of Cordyceps species (e.g., host, arrangement of perithecia, ascospore fragmentation, conidiogenous structures, conidial shape, and size) are not phylogenetically informative [15,17,18,27]. Therefore, strain KACC 410975 and six previously studied strains (NTUPPMCC 18-115, NTUPPMCC 18-116, TBRC 7259, TBRC 7260, TBRC 7261, and TBRC 7262) were identified as C. koratensis by multilocus sequence analysis (Fig. 1).
In summary, the current study identified an entomopathogenic fungus infecting the larvae of lepidopterans as Cordyceps koratensis. To date, 59 Cordyceps species have been identified worldwide. Among these, C. ampullacea, C. bifusispora, C. brongniartii, C. coccidiocapitata, C. isarioides, C. kyusyuensis, C. militaris, C. ninchukispora, C. ochraceostromata, C. ootakiensis, C. pruinosa, C. rosea, and C. tenuipes have been recorded in Korea [28]. To our knowledge, this study represents the first record of C. koratensis in Korea. These results help expand our current knowledge of Cordyceps diversity in Korea and supplement the available bioresources in Korea from the Cordyceps. Moreover, this finding will help develop biocontrol agents within the framework of ecofriendly pest management for C. koratensis.
The authors pronounce that they have no potential conflict of interest.
This study was supported by grants (PJ017286) from the Rural Development Administration and (M3H9A1081254) from the Ministry of Science and ICT of Korea.
1. Lacey L. Entomopathogens used as microbial control agents. Microbial control of insect and mite pests. Cambridge, MA: Academic Press; 2017. p. 3-12. [DOI]
2. Wang C, Wang S. Insect pathogenic fungi: Genomics, molecular interactions, and genetic improvements. Annu Rev Entomol 2017;62:73-90. [DOI]
3. Hajek AE, Shapiro-Ilan DI. Ecology of invertebrate diseases. Hoboken, NJ: John Wiley & Sons; 2018. [DOI]
4. Wu X, Wu T, Huang A, Shen Y, Zhang X, Song W, Wang S, Ruan H. New insights into the biosynthesis of typical bioactive components in the traditional Chinese medicinal fungus Cordyceps militaris. Front Bioeng Biotechnol 2021;9:801721. [DOI]
5. Ortiz-Urquiza A, Keyhani NO. Action on the surface: Entomopathogenic fungi versus the insect cuticle. Insects 2013;4:357-74. [DOI]
6. Lacey LA, Kaya HK. Field manual of techniques in invertebrate pathology: Application and evaluation of pathogens for control of insects and other invertebrate pests. Dordrecht: Springer Dordrecht; 2007. [DOI]
7. Wu S, Reddy GV, Jaronski ST. Advances in microbial insect control in horticultural ecosystem. In: Nandwani D, editor. Sustainable horticultural systems: Issues, technology and innovation. Cham: Springer; 2014. p. 223-52. [DOI]
8. Sain SK, Monga D, Hiremani NS, Nagrale DT, Kranthi S, Kumar R, Kranthi KR, Tuteja OP, Waghmare VN. Evaluation of bioefficacy potential of entomopathogenic fungi against the whitefly (Bemisia tabaci Genn.) on cotton under polyhouse and field conditions. J Invertebr Pathol 2021;183:107618. [DOI]
9. Wu S, Toews MD, Castrillo LA, Barman AK, Cottrell TE, Shapiro-Ilan DI. Identification and virulence of Cordyceps javanica strain wf GA17 isolated from a natural fungal population in sweetpotato whiteflies (Hemiptera: Aleyrodidae). Environ Entomol 2021;50:1127-36. [DOI]
10. Mora MAE, Castilho AMC, Fraga ME. Classification and infection mechanism of entomopathogenic fungi. Arq Inst Biol 2017;84:e0552015. [DOI]
11. Olatunji OJ, Tang J, Tola A, Auberon F, Oluwaniyi O, Ouyang Z. The genus Cordyceps: An extensive review of its traditional uses, phytochemistry and pharmacology. Fitoterapia 2018;129:293-316. [DOI]
12. Zha LS, Huang SK, Xiao YP, Boonmee S, Eungwanichayapant PD, McKenzie EHC, Kryukov V, Wu XL, Hyde KD, Wen TC. An evaluation of common Cordyceps (Ascomycetes) species found in Chinese markets. Int J Med Mushrooms 2018;20:1149-62. [DOI]
13. Chen WH, Liu C, Han YF, Liang JD, Tian WY, Liang ZQ. Three novel insect-associated species of Simplicillium (Cordycipitaceae, Hypocreales) from Southwest China. MycoKeys 2019;58:83-102. [DOI]
14. Shrestha B, Tanaka E, Hyun MW, Han JG, Kim CS, Jo JW, Han SK, Oh J, Sung GH. Coleopteran and lepidopteran hosts of the entomopathogenic genus Cordyceps sensu lato. J Mycol 2016;2016:7648219. [DOI]
15. Kepler RM, Luangsa-Ard JJ, Hywel-Jones NL, Quandt CA, Sung GH, Rehner SA, Aime MC, Henkel TW, Sanjuan T, Zare R, et al. A phylogenetically-based nomenclature for Cordycipitaceae (Hypocreales). IMA fungus 2017;8:335-53. [DOI]
16. Mongkolsamrit S, Noisripoom W, Tasanathai K, Khonsanit A, Thanakitpipattana D, Himaman W, Kobmoo N, Luangsa-Ard JJ. Molecular phylogeny and morphology reveal cryptic species in Blackwellomyces and Cordyceps (Cordycipitaceae) from Thailand. Mycol Progress 2020;19:957-83. [DOI]
17. Wang YB, Wang Y, Fan Q, Duan DE, Zhang GD, Dai RQ, Dai YD, Zeng WB, Chen ZH, Li DD, et al. Multigene phylogeny of the family Cordycipitaceae (Hypocreales): New taxa and the new systematic position of the Chinese cordycipitoid fungus Paecilomyces hepiali. Fungal Divers 2020;103:1-46. [DOI]
18. Sung GH, Hywel-Jones NL, Sung JM, Luangsa-Ard JJ, Shrestha B, Spatafora JW. Phylogenetic classification of Cordyceps and the clavicipitaceous fungi. Stud Mycol 2007;57:5-59. [DOI]
19. Tasanathai K, Thanakitpipattana D, Noisripoom W, Khonsanit A, Kumsao J, Luangsa-ard JJ. Two new Cordyceps species from a community forest in Thailand. Mycol Progress 2016;15:18. [DOI]
20. Zha LS, Wen TC, Huang SK, Boonmee S, Eungwanichayapant PD. Taxonomy and biology of Cordyceps qingchengensis sp. nov. and its allies. Phytotaxa 2019;416:14-24. [DOI]
21. Li YP, Chen WH, Han YF, Liang JD, Liang ZQ. Cordyceps yinjiangensis, a new antpathogenic fungus. Phytotaxa 2020;453:284-92. [DOI]
22. Li ZZ, Luan FG, Hywel-Jones NL, Zhang SL, Chen MJ, Huang B, Sun CS, Chen ZA, Li CR, Tan YJ, et al. Biodiversity of cordycipitoid fungi associated with Isaria cicadae Miquel II: Teleomorph discovery and nomenclature of chanhua, an important medicinal fungus in China. Mycosystema 2021;40:95-107.
23. Tamura K, Stecher G, Kumar S. MEGA11: Molecular evolutionary genetics analysis version 11. Mol Biol Evol 2021;38:3022-7. [DOI]
24. Hsieh LS, Tzean SS, Wu WJ. The genus Akanthomyces on spiders from Taiwan. Mycologia 1997;89:319-24. [DOI]
25. Mongkolsamrit S, Noisripoom W, Tasanathai K, Kobmoo N, Thanakitpipattana D, Khonsanit A, Petcharad B, Sakolrak B, Himaman W. Comprehensive treatise of Hevansia and three new genera Jenniferia, Parahevansia and Polystromomyces on spiders in Cordycipitaceae from Thailand. MycoKeys 2022;91:113-49. [DOI]
26. Chuang WY, Lin YC, Shrestha B, Luangsa-Ard JJ, Stadler M, Tzean SS, Wu S, Ko CC, Hsieh SY, Wu ML, et al. Phylogenetic diversity and morphological characterization of cordycipitaceous species in Taiwan. Stud Mycol 2024;109:1-56. [DOI]
27. Mongkolsamrit S, Noisripoom W, Thanakitpipattana D, Wutikhun T, Spatafora JW, LuangsaArd J. Disentangling cryptic species with isaria-like morphs in Cordycipitaceae. Mycologia 2018;110:230-57. [DOI]
28. National Institute of Biological Resources. 2024 National Species List of Korea [Internet]. Incheon: National Institute of Biological Resources; 2024 [cited 2024 December 16]. Available from: https://species.nibr.go.kr/home/mainHome.do?cont_ link=002&subMenu=002001&contCd=002001001