1Department of Plant Medicine, Jeonbuk National University, Jeonju 54896, Korea
2Research Center for Plant Medicine, Jeonbuk National University, Jeonju 54896, Korea
3Division of Environmental Science and Ecological Engineering, Korea University, Seoul 02841, Korea
*Correspondence to hdshin@korea.ac.kr
Korean Journal of Mycology (Kor J Mycol) 2025 June, Volume 53, Issue 2, pages 101-107.
https://doi.org/10.4489/kjm.2025.53.2.5
Received on February 20, 2025, Revised on June 10, 2025, Accepted on June 11, 2025, Published on June 30, 2025.
Copyright © The Korean Society of Mycology.
This is an Open Access article which is freely available under the Creative Commons Attribution-NonCommercial 4.0 International License (CC BY-NC) (https://creativecommons.org/licenses/by-nc/4.0/).
Pseudocercospora rubigena is a recently introduced leaf spot pathogen on Rubus sp. based on single-strain identification. This study addressed the limitations of the initial description of Ps. rubigena by examining multiple isolates of Rubus corchorifolius in Korea, emphasizing its first confirmed occurrence on this host. Notably, shorter conidia with a narrow septation range are considered key characteristics that distinguish Ps. rubigena from related species. Sequences of the internal transcribed spacer (ITS) region, along with protein-coding genes such as partial actin (actA), partial translation elongation factor 1-alpha (tef1), and partial DNA-directed RNA polymerase II second-largest subunit (rpb2), were obtained from three Korean isolates. Later, these sequences were included in phylogenetic analyses using a multigene dataset of ITS, act, tef1 and rpb2. The results showed that Pseudocercospora isolates from R. corchorifolius clustered in a single clade with Ps. rubigena, thus confirming the consistency between their morphological characteristics and those of the latter. This study clarifies the morphological and molecular phylogeny of Ps. rubigena, contributing to the taxonomy of Pseudocercospora species and providing insights into its host specificity and distribution.
cercosporoid fungi, Mycosphaerellaceae, phytopathogen, Pseudocercospora rubi, Rosaceae
The genus Pseudocercospora Speg. is a diverse group of fungi known to cause leaf spot diseases on a wide range of plants globally. Using new molecular phylogenetic approaches, this genus has undergone significant taxonomic reclassification, enhancing our understanding of its biodiversity. New species continue to be discovered, and previously uncertain species classifications are being resolved [1,2]. Among these, Pseudocercospora rubi (Sacc.) Deighton was first reported on Rubus sp. in the United States in 1984 [3] and has since been recognized as the major leaf spot pathogen affecting various Rubus L. species. The genus encompasses more than 30 plants species, of them R. corchorifolius L. f., R. coreanus Miq., and R. pungens var. oldhamii (Miq.) Maxim. (= R. oldhamii Miq.) have been listed as potential hosts for this fungus in Korea [4,5]. Its classification has long been based on morphological traits, such as branched mycelia, short conidiophores, and a high range of septation in conidia.
In addition to Ps. rubi, several other Pseudocercospora species infect Rubus plants, such as Ps. heteromalla (Syd.) Deighton in India, South Africa, China, and Iran [6–9]; Ps. norchiensis Crous in Iran and New Zealand [1,10]; Ps. rubicola (Thüm.) X.J. Liu & Y.L. Guo in China [11]; and Ps. pancratii (Ellis & Everh.) U. Braun & R.F. Castaňeda in the United States [12]. Despite these findings, detailed morphological descriptions and molecular phylogenetic characterizations of some of the above-mentioned species remain incomplete, leaving gaps in our understanding of their classification and host ranges.
Recently, Ps. rubigena Crous, Y. Zhang ter. & Yuan Yuan Chen was described as a new pathogen on Rubus sp. from China [2]. Although it was phylogenetically distinct from the recorded sequences of other Pseudocercospora species, its morphological traits closely matched those of Ps. rubi, as both species share similar features. Furthermore, the conclusions in Ps. rubigena are constrained by the reliance on molecular phylogenetic results for a single strain. To address these gaps and clarify the species concept of Pseudocercospora on Rubus corchorifolius in Korea, multiple Pseudocercospora isolates from selected host plants collected across various regions were included in this study to confirm their identity and provide further insights into their morphological variability and distribution.
In total, eight samples of R. corchorifolius infected by Pseudocercospora leaf spots collected during our field forays and preserved in the Herbaria of Korea University (KUS) and Jeonbuk National University (JBNU) were involved in the present study (Table 1). Culture isolation and molecular analysis were performed using fresh specimens, which were later deposited in relevant databases.
The morphological characteristics of the fungus were observed using an Olympus BX50 microscope (Olympus, Tokyo, Japan). Photomicrographs were captured using a Zeiss AX10 microscope equipped with an AxioCam MRc5 camera (Carl Zeiss, Oberkochen, Germany). At least thirty measurements were performed for each diagnostic structure. To obtain monoconidial isolates, conidia collected from young lesions were placed in Eppendorf tubes containing sterile water. The prepared conidial suspension was then streaked onto 2% water agar (WA; Junsei, Tokyo, Japan) plates supplemented with 100 mg/L of streptomycin sulfate and incubated at 25℃. After 2 days, germinating conidia were transferred onto 2% potato dextrose agar (PDA; Difco, France) plates. The obtained colonies were deposited at the Korean Agricultural Culture Collection (KACC), Rural Development Administration, Korea (Table 1).
Table 1. Sources of Korean fungal specimens used in this study, with addition of accession numbers of obtained strains, and sequences deposited in the NCBI and KACC, respectively
테이블
NCBI: National Center for Biotechnology Information; KACC: Korean Agricultural Culture Collection, Rural Development Administration; ITS: internal transcribed spacer; actA: actin; tef1: translation elongation factor 1-alpha; rpb2: DNA-directed RNA polymerase II second-largest subunit.
Leaf spots were amphigenous, irregular to angular, 3–6 mm in diameter, vein-limited, with a light creamy to khaki center, surrounded by a dark brown margin (Fig. 1A–B). Caespituli were amphigenous but primarily epiphyllous, fasciculate to solitary. Stromata were well-developed, composed of several swollen hyphal cells, pale to light-brown, erumpent, and up to 50 µm in diameter (Fig. 1C–D). Mycelium was internal, consisting of branched, septate, brownish hyphae. Conidiophores were numerous, forming dense fascicles, 25–57 × 3–6 µm, brown to pale brown, smooth-walled, subcylindrical, straight or slightly curved, and 1–3-septate. Conidiogenous cells were unbranched, pale to medium brown, 10–21 × 3–4 µm, with inconspicuous scars (Fig. 1E). Conidia were solitary, hyaline, narrowly obclavate to subcylindrical, straight or slightly flexuous, 1–6-septate, with an obtuse apex and truncate base, 47–80 × 3–4 µm, and featuring unthickened, non-darkened hila (Fig. 1F–G). One-week-old colonies grown on PDA at 25℃ were 7–10 mm in diameter, khaki in appearance, undulate, with a clear margin, and developed moderate aerial mycelium (Fig. 1H). The morphological characteristics of our isolates were consistent with those of Ps. rubigena [2].
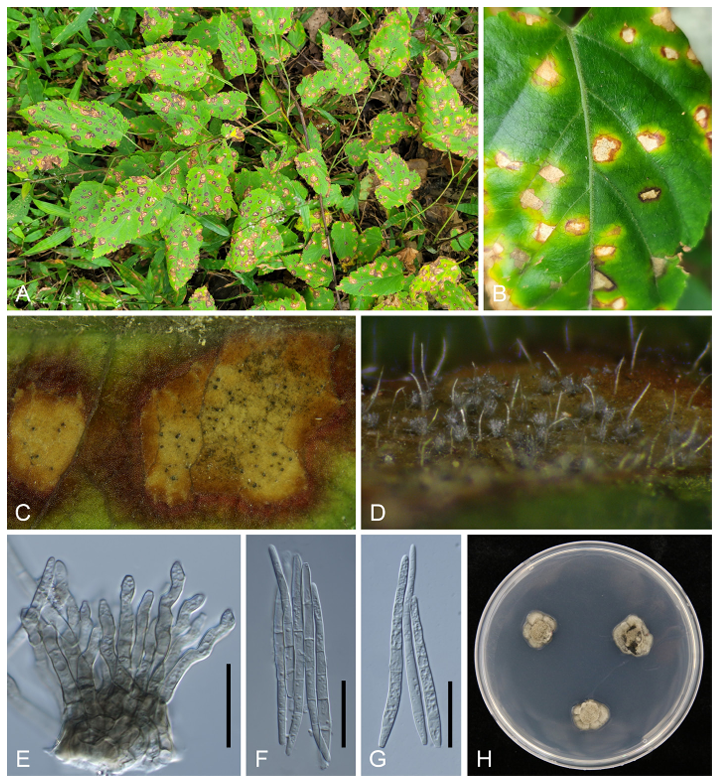

Fig. 1. Leaf spot disease caused by Pseudocercospora rubigena on Rubus corchorifolius. (A) Symptoms on infected plants in the field. (B) Close-up of leaf spots. (C) Lesions with distinct brown margins and abundant caespituli in the creamy center. (D) Heavy fructification showing greyish conidial masses. (E) Conidiophores. (F, G) Conidia. (H) One-week-old colony of Ps. rubigena grown on potato dextrose agar at 25℃. Scale bar indicates 20 µm.
To confirm the morphology-based identification, DNA was extracted from three two-week-old colonies grown on PDA to perform molecular phylogenetic analyses. Nucleotide sequences of the internal transcribed spacer (ITS) region, along with protein-coding genes such as partial actin (actA), partial translation elongation factor 1-alpha (tef1), and partial DNA-directed RNA polymerase II second-largest subunit (rpb2), were determined using the primer pairs V9G/ITS4, ACT-512F/ACT-783R, EF1-728F/EF1986R, and P_RPB2-F/P_RPB2-R, respectively [13,14]. Assembled sequences were submitted to GenBank (Table 1). For phylogenetic analyses, a data matrix was generated for each gene individually and aligned using the MUSCLE algorithm in MEGA 11 [15]. All four datasets were then concatenated into a single multigene dataset in the order ITS, act, tef1 and rpb2 using SequenceMatrix [16]. Trochophora simplex (CBS 214744) and Pallidocercospora heimioides (CBS 111190) were selected as outgroups [13]. Maximum parsimony (MP) analysis was performed in PAUP* 4.0a using a heuristic search with the tree-bisection algorithm [17], while maximum likelihood (ML) analysis was conducted in raxmlGUI 2.0.13 based on the GTR substitution model with GAMMA distribution [18]. Tree robustness was assessed via 1,000 bootstrap replications. The final alignment included 31 sequences and 1,912 characters, of which 206 (10.77%) were variable and parsimony-uninformative, while 535 (29.55%) were informative for parsimony analysis.
BLASTn search results showed 99.85% similarity in the ITS region with Pseudocercospora sp. on Syzygium cordatum (KX287305), differing by a single base (adenine in all three Korean isolates vs. thymine in the reference strain). The actA and rpb2 sequences were 100% identical to Ps. rubigena (PP404605), while tef1 showed 99% identity with Ps. rubigena (PP404923). In the phylogenetic tree, all three Korean isolates clustered in a distinct, highly supported branch (100/100 BS values) with Ps. rubigena (Fig. 2). The phylogenetic separation of Ps. rubi, Ps. rubigena, Ps. norchiensis, and Ps. pancratii into distinct clades indicates genetic divergence among Rubus leaf spot pathogens, suggesting adaptation to specific environments and hosts.
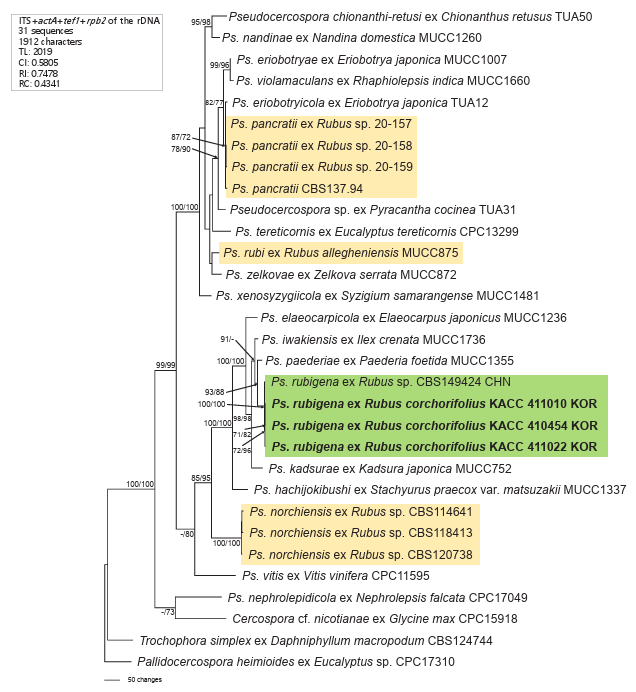
Fig. 2. Phylogenetic tree of Pseudocercospora rubigena on Rubus corchorifolius constructed using maximum parsimony (MP) based on a combined multigene dataset (ITS + actA + tef1 + rpb2) comprising 31 sequences. Isolates obtained in this study are shown in bold. Bootstrap values (> 70%) from MP and maximum likelihood (ML) analyses are indicated on corresponding branches, respectively. Tree scores, such as tree length (TL), consistency index (CI), retention index (RI), and rescaled consistency index (RC), are provided in the box on the left. ITS: internal transcribed spacer; actA: actin; tef1: translation elongation factor 1-alpha; rpb2: DNA-directed RNA polymerase II secondlargest subunit.
The morphology of Korean specimens on R. corchorifolius was compared with Ps. rubigena and Ps. rubi previously reported in Korea (Table 2). Conidia of Ps. rubi are longer (up to 132 µm), with more septa (3–11), and conidiophores are narrower (up to 4 µm) than those of Ps. rubigena and the Korean isolates. However, under in vitro conditions, conidia of Ps. rubigena have been measured up to 123.5 µm in length, with (3–)5–10 septa [2], closely resembling the characteristics of Ps. rubi reported in Korea. Owing to the lack of reliable data, colony morphology comparisons between Ps. rubigena and Ps. rubi were not provided. Nevertheless, these findings support the conclusion that the previously reported Ps. rubi on R. corchorifolius in Korea is actually Ps. rubigena. Separately, ITS, LSU, rpb2, actA, and tef1 sequences obtained from MUCC 875 on Rubus allegheniensis in Japan have been deposited in GenBank. However, owing to the absence of molecular-phylogenetic confirmation of Ps. rubi in Korea, further studies using updated molecular-phylogenetic approaches are required to verify its presence.
Table 2. Morphological comparison of Korean isolates with Pseudocercospora rubigena and Ps. rubi
테이블
The genus Rubus (Rosaceae; Rosoideae) encompasses approximately 700 species distributed across temperate and subtropical regions, including various shrubs, subshrubs, and herbs [19]. Many are economically important plants, commonly known as brambles, raspberries, or blackberries. R. corchorifolius is native to East Asia, including China, Korea, and Japan [20]. The phylogenetic and geographic range of R. corchorifolius closely aligns with the distribution of Ps. rubigena, first reported on Rubus sp. in China [2] and now confirmed in Korea. This overlap suggests a potential evolutionary relationship between host and pathogen, supporting the hypothesis that Ps. rubigena is a distinct species adapted to specific Rubus hosts.
This study reports a new finding of Ps. rubigena in Korea and identifies a new host record for this species globally. The findings provide strong evidence for the identification of Ps. rubigena, and the observed morphological variations may reflect environmental influences. Consequently, this work contributes to a deeper understanding of the taxonomy, phylogeny, and host range of Ps. rubigena for future research.
The authors declare no competing interests.
1. Crous PW, Braun U, Hunter GC, Wingfield MJ, Verkley GJM, Shin HD, Nakashima C, Groenewald JZ. Phylogenetic lineages in Pseudocercospora. Stud Mycol 2013;75:37-114. [DOI]
2. Groenewald JZ, Chen YY, Zhang Y, Roux J, Shin HD, Shivas RG, Summerell BA, Braun U, Alfenas AC, Ujat AH, et al. Species diversity in Pseudocercospora. Fung Syst Evol 2024;13:29-89. [DOI]
3. Alfieri SA Jr, Langdon KR, Wehlburg C, Kimbrough JW. Index of plant diseases in Florida. Revised ed. Gainesville, FL: Florida Department of Agriculture and Consumer Services, Division of Plant Industry; 1984. p. 1-389.
4. Farr DF, Rossman AY. Fungal databases, U.S. National Fungus Collections [Internet]. Maryland: ARS, USDA; 2025 [cited 2025 Feb 6]. Available from https://fungi.ars.usda.gov/.
5. Shin HD, Kim JD. Cercospora and allied genera from Korea. Suwon: National Institute of Agricultural Science and Technology; 2001. p. 1-302.
6. Kamal. Cercosporoid fungi of India. Dehra Dun, India: Bishen Singh Mahendra Pal Singh; 2010. p. 1-349.
7. Crous PW, Phillips AJL, Baxter AP. Phytopathogenic fungi from South Africa. Stellenbosch, South Africa: University of Stellenbosch, Department of Plant Pathology Press; 2000. p. 1-358.
8. Liu XJ, Guo YI. Flora Fungorum Sinicorum. Vol. 9. Pseudocercospora. Beijing: Science Press; 1998. p. 1-474.
9. Pirnia M, Zare R, Zamanizadeh HR, Khodaparast A. New records of cercosporoid hyphomycetes from Iran. Mycotaxon 2012;120:157-69. [DOI]
10. Guatimosim E, Schwartsburd PB, Barreto RW, Crous PW. Novel fungi from an ancient niche: Cercosporoid and related sexual morphs on ferns. Persoonia 2016;37:106-41. [DOI]
11. Zhuang WY. Higher fungi of tropical China. Ithaca, NY: Mycotaxon Ltd; 2001.
12. Marin MV, Wang NY, Seijo T, Druffel KL, McGuin RL, Deng Z, Peres NA. Pseudocercospora pancratii causing leaf spots on commercial blackberry (Rubus sp.) in Florida. Plant Dis 2023;107:131-5. [DOI]
13. Nakashima C, Motohashi K, Chen CY, Groenewald JZ, Crous PW. Species diversity of Pseudocercospora from Far East Asia. Mycol Prog 2016;15:1093-117. [DOI]
14. Lee KJ, Choi IY, Abasova L, Choi JH, Park JH, Shin HD. Morphology and molecular phylogeny of Pseudocercospora fatouae associated with hypophyllous leaf mold on Fatoua villosa in Korea. Kor J Mycol 2023;51:147-53.
15. Tamura K, Stecher G, Kumar S. MEGA11: Molecular evolutionary genetics analysis version 11. Mol Biol Evol 2021;38:3022-7. [DOI]
16. Vaidya G, Lohman DJ, Meier R. SequenceMatrix: Concatenation software for the fast assembly of multi-gene datasets with character set and codon information. Cladistics 2011;27:171-80. [DOI]
17. Swofford DL. PAUP: Phylogenetic analysis using parsimony (and other methods) 4.0b10. Sunderland, MA: Sinauer Associates; 2002.
18. Silvestro D, Michalak I. raxmlGUI: A graphical front-end for RAxML. Org Divers Evol 2012;2:335-7. [DOI]
19. Yu J, Fu J, Fang Y, Xiang J, Dong H. Complete chloroplast genomes of Rubus species (Rosaceae) and comparative analysis within the genus. BMC Genomics 2022;23:1-14. [DOI]
20. Huang JY, Hu JM. Revision of Rubus (Rosaceae) in Taiwan. Taiwania 2009;54:285-310.