So-Jeong Ye1, Ji-yeong Jang1, Seong-Keun Lim1, Seung-Yeol Lee1,2, and Hee-Young Jung1,2*
1Department of Plant Medicine, Kyungpook National University, Daegu 41566, Korea
2Institute of Plant Medicine, Kyungpook National University, Daegu 41566, Korea
*Correspondence to heeyoung@knu.ac.kr
Korean Journal of Mycology (Kor J Mycol) 2025 December, Volume 53, Issue 4, pages 237-243.
https://doi.org/10.4489/kjm.2025.53.4.1
Received on October 17, 2025, Revised on November 10, 2025, Accepted on November 10, 2025, Published on December 31, 2025.
Copyright © The Korean Society of Mycology.
This is an Open Access article which is freely available under the Creative Commons Attribution-Non-Commercial 4.0 International License (CC BY-NC) (https://creativecommons.org/licenses/by-nc/4.0/).
A fungal strain designated KNUF-21-010 was isolated from soil obtained in Yangsan-si, Gyeongsangnam-do, Korea. Cultural characteristics, morphological features, and multilocus sequence analysis based on the internal transcribed spacer (ITS) regions, large subunit of 28S rRNA (LSU), and translation elongation factor 1-alpha (tef1-α) genes were employed to identify the strain. The strain was cultivated on potato carrot agar (PCA) and oatmeal agar (OA), and after 4 weeks of incubation at 25°C, colony diameters ranged from 58.4–64.2 mm on PCA and 68.0–72.4 mm on OA. Brown, unbranched conidiophores; hyaline to pale brown hyphae; obovate to ellipsoidal conidia; and globose to subglobose chlamydospores were observed. These features closely matched those of Chloridium chlamydosporum. BLAST analysis of the ITS, LSU, and tef1-α sequences reveals that strain KNUF-21-010 shared > 99% sequence similarity with Chl. chlamydosporum. Phylogenetic analysis further confirmed that the strain formed a highly supported clade with Chl. chlamydosporum. Based on cultural, morphological, and molecular phylogenetic analyses, strain KNUF-21-010 was identified as Chl. chlamydosporum. This study represents the first report of Chl. chlamydosporum in Korea.
Chloridium chlamydosporum, Saprophytic fungi, Soil fungi
The genus Chloridium was established by Link in 1809, with Chloridium virescens designated as the type species [1]. Chloridium comprises saprophytic fungi and is classified in the family Chaetosphaeriaceae, order Chaetosphaeriales, and class Sordariomycetes. Chaetosphaeriaceae, one of the largest families within Sordariomycetes, is commonly found on various decaying plant substrates, including leaves, fruits, branches, bark, and wood [2]. In this family, Chloridium also thrives on decaying plant materials in terrestrial and freshwater habitats [3]. Traditionally, species in Chloridium are regarded as saprophytic fungus that primarily grows on wood or other plant debris [4]. However, recent studies show that some species are frequently isolated from soil, indicating that members of Chloridium can also be classified as soilborne fungi [3,5].
The genus Chloridium was originally proposed for a single species, Chl. viride [1], which is morphologically characterized based on pigmented, unbranched, erect, caespitose conidiophores and hyaline, globose conidia that adhere in green masses at the tip [3]. Subsequent taxonomic studies show that Chl. viride is conspecific with Dematium virescens (1794) [3]. According to the principle of nomenclatural priority, the epithet “virescens” takes precedence over Chl. viride, and the species name is accordingly revised to Chl. virescens [4].
Chl. chlamydosporum is initially described under the genus Bisporomyces based on its distinct morphological characteristics, including conidiophores that produce pairs of conidia at the tip of the phialide [6,7]. In subsequent years, the taxonomic relationships among Bisporomyces chlamydosporum, Chl. virescens, and Cirrhomyces caudiger was debated owing to their morphological similarities. Hughes later synonymizes the genera Cirrhomyces and Bisporomyces with Chloridium, recognizing five species within the Chloridium, including Chl. chlamydosporum [8].
The application of molecular phylogenetic methods in recent fungal taxonomy studies enables a more accurate understanding of taxonomic and phylogenetic relationships [2,3]. Traditionally circumscribed based on morphological characteristics, the genus Chloridium undergoes continuous taxonomic revisions at the genus and species levels with the advent of molecular phylogenetics. This approach addresses the limitations of morphology-based classification, and subsequent studies reveal that Chloridium is a polyphyletic group [2,3,9,10]. Since the genus Chloridium is revealed to be polyphyletic, researchers collect genetic data from Chloridium species and conduct phylogenetic analyses to revise its classification [2,3]. These studies highlight the importance of accurate classification through molecular approaches in fungal taxonomy.
This study aims to isolate the strain KNUF-21-010, presumed to be Chloridium chlamydosporum, from soil collected in Yangsan-si, Gyeongsangnam-do, which has not previously been revealed in Korea. To achieve accurate identification of this fungus, cultural, morphological, and molecular phylogenetic analyses were conducted. Multilocus sequence analysis (MLSA) was performed based on the internal transcribed spacer (ITS) regions, large subunit of 28S rRNA (LSU), and translation elongation factor 1-alpha (tef1-α).
The strain KNUF-21-010 was collected from the soil in Yangsan-si, Gyeongsangnam-do, Korea (35°30’53.1″N 129°03’08.1″E). The standard serial dilution method was performed to isolate fungal strains from the soil [11]. One gram of collected soil sample was added to 10 mL of sterile distilled water and mixed thoroughly by vortexing. A 100 μL of the suspension was spread onto potato dextrose agar (PDA; Difco, Detroit, MI, USA) plates, which were incubated in the dark at 25°C for approximately 3 days. Single colonies were transferred to a fresh PDA plate and incubated at 25°C. This step was repeated to obtain pure cultures, and several fungal strains were isolated. DNA was extracted, and polymerase chain reaction (PCR) amplification of the ITS regions was conducted. Among these strains, the ITS sequence of KNUF-21-010 was selected for further taxonomic study because it indicated a potentially unreported fungal species in Korea. Subsequently, cultural, morphological, and molecular phylogenetic analyses were conducted to identify the strain. The isolate KNUF-21-010 was deposited at the National Institute of Biological Resources as a metabolically inactive culture (NIBRFGC000509190).
The cultural characteristics of strain KNUF-21-010 were examined on potato carrot agar (PCA; HiMedia, Mumbai, India) and oatmeal agar (OA; Difco, Detroit, MI, USA) plates. Colonies on both media were incubated at 25°C for 4 weeks in the dark. The features of the colonies on each medium, including color, shape, and size, were compared after 4 weeks of incubation. Colonies were imaged after 3 weeks using a Canon EOS 5D Mark III digital camera (Canon, Tokyo, Japan). To observe morphological characteristics, slide cultures were prepared from colonies grown on PCA. The morphological features, including hyphae, conidiophore, chlamydospores, conidia, and conidiogenous cells, were observed after 1~2 weeks of incubation at 25°C in the dark, under a light microscope (BX50, Olympus, Tokyo, Japan).
The strain KNUF-21-010 was incubated on PDA at 25°C for 2 weeks. DNA was extracted from the hyphae of this colony using a HiGene Genomic DNA Prep Kit (Biofact, Daejeon, South Korea), following the protocol of the manufacturer. The extracted DNA was used to amplify the ITS regions, LSU, and tef1-α. The ITS regions were amplified with the primer pair V9G/LR8 [12,13], and the tef1-α was amplified with the primer pair EF1-983F/EF1-2218R [14]. PCR amplicons were purified using the ExoSAP-IT™ PCR Product Cleanup Reagent (Thermo Fisher Scientific, Waltham, MA, USA). The purified amplicons were sequenced using Macrogen (Sejong, South Korea). Sequencing of the LSU was performed using primer JS1, JS7, JS8, and LR7 [15]. The ITS, LSU, and tef1-α sequences of strain KNUF-21-010 were deposited in GenBank under the accession numbers LC886056 (ITS), LC886057 (LSU), and LC886058 (tef1-α).
Sequences of each region were compared with those of other species using National Center for Biotechnology Information (NCBI) Basic Local Alignment Search Tool (BLAST). Table 1 presents the assembled DNA data registered in NCBI, used to determine the phylogenetic position of strain KNUF21-010 relative to other species. The collected gene data from each region were aligned using Clustal X (version 2.0). Phylogenetic trees were constructed using the maximum likelihood method with the Kimura 2-parameter model and 1,000 bootstrap replicates [16]. The trees were generated using MEGA software (version 12.0) [17].
Table 1. List of species used in phylogenetic analysis along with their GenBank accession numbers
테일블
Colonies were grown on PCA and OA to compare the cultural characteristics of strain KNUF-21-010 with those of Chl. chlamydosporum and Chl. peruense. On PCA, colonies of strain KNUF-21-010 were whitish beige at the center, cinnamon to brown towards the periphery, and dark olivaceous grey towards the margin, reaching 58.4–64.2 mm in diameter after 4 weeks of incubation. They were circular, flat, margin fimbriate, velvety to cobwebby in texture, with a brown reverse (Fig. 1A, 1B). On OA, colonies were olivaceous dark grey, sometimes producing brown pigment. They were circular, flat, velvety to cobwebby, with an entire margin and a brown reverse (Fig. 1C, 1D). The colonies peaked at 68.0–72.4 mm in diameter after 4 weeks of incubation. Following microscopic observation of KNUF-21-010, the features of its conidiophores, conidia, chlamydospores, and vegetative hyphae were examined and compared. The conidiophores of KNUF-21-010 were brown, becoming paler toward the apex, macronematous, straight, unbranched, septate, and cylindrical, measuring 34–152(–245) × 2.7–3.5(–4.07) μm (Fig. 2A, 2B). Phialides that form conidia were present at the apices of the conidiophores (Fig. 2C). KNUF-21010 produced chlamydospores that were lateral, sessile, intercalary, terminal, solitary, or arranged in short chains; globose to subglobose, thick-walled, smooth, and brown (Fig. 2D–F). The chlamydospores peaked at 4.75–6.21 × 4.16–5.41 μm. The strain also produced hyaline conidia that turned pale brown with age (Fig. 2G). The conidia were obovate to ellipsoidal, smooth, aseptate, with dimensions of 4.32–5.00 × 2.81–3.80 μm. The vegetative hyphae were hyaline to pale brown, septate, smooth, and 2.16–3.86 μm wide. Most microscopic features observed in KNUF-21-010 corresponded to those of Chl. chlamydosporum, the reference species (Table 2). Chl. peruense exhibited greater growth on PCA and OA media and produced smaller conidia than KNUF-21-010 and Chl. chlamydosporum.
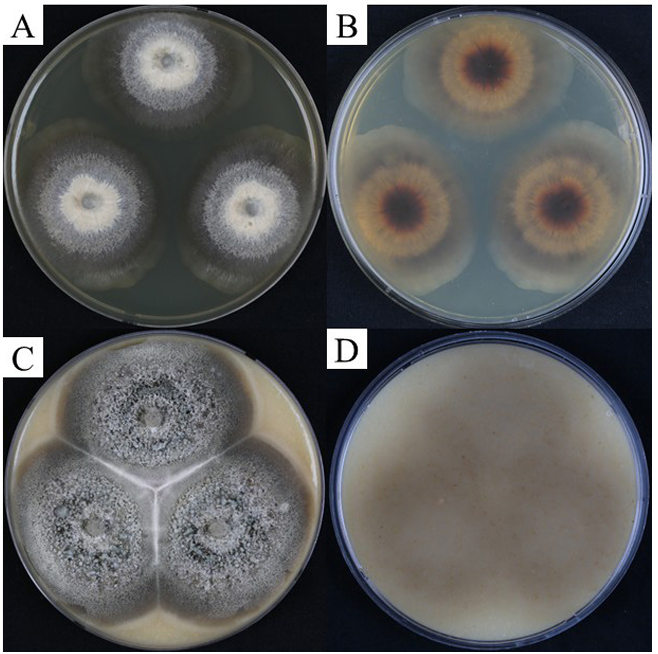
Fig. 1. Colony of KNUF-21-010 (Chloridium chlamydosporum) on oatmeal agar (OA) and potato carrot agar (PCA) after 28 days at 25°C. A, B: Colony on PCA; C, D: Colony on OA.
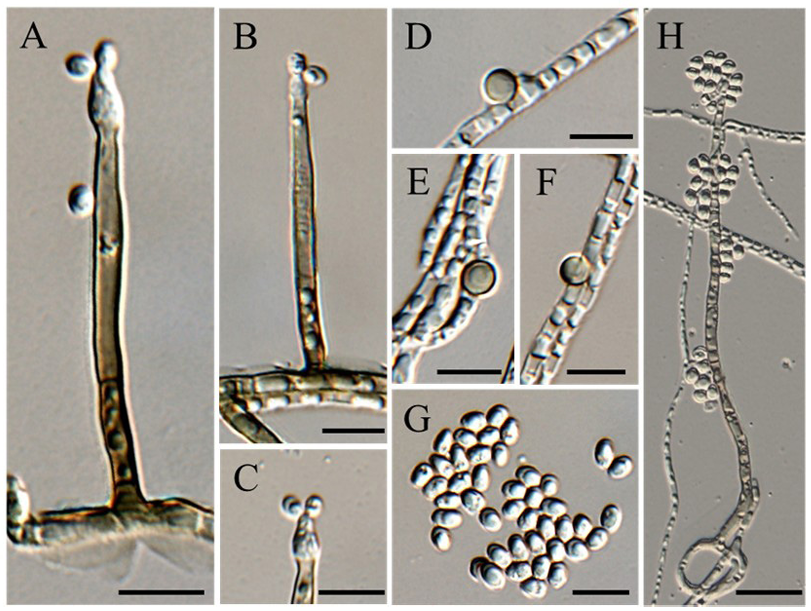
Fig. 2. Morphological characteristics of Chloridium chlamydosporum. A–B: Conidiophores; C: Tip of the phialide; D–F: Chlamydospores; G: Conidia; H: Conidiophores with multiple conidia. Scale bars: AG, 10 μm; H, 20 μm.
Table 2. Comparison of morphological characteristics of KNUF–21–010 with Chloridium chlamydosporum and Chloridium peruense
table
PCA: potato carrot agar; OA: oatmeal agar.
Sequencing of the ITS regions, LSU, and tef1-α yielded sequences of 581 bp, 1,765 bp, and 966 bp, respectively. Sequences from KNUF-21-010 were analyzed using the NCBI BLAST tools and showed a high level of similarity within the three regions. The ITS regions of KNUF-21-010 showed 99.79% identity with Chl. chlamydosporum (CBS 114.41; OP455385), 99.60% with Chl. humicola (WZ-918; OP163808), and 99.16% with Chl. virescens (KUNCC 24-17978; PQ22234). The LSU of KNUF-21-010 exhibited 99.89% identity with Chl. chlamydosporum (CBS 114.41; OP455492), 99.66% with Chl. detriticola (M.R. 3774; OP455508), and 99.49% with Chl. peruense (CBS 126074; OP455531). The tef1-α of the strain exhibited 99.88% identity with Chl. chlamydosporum (CBS 149052; OP464962), 98.45% with Chl. peruense (CBS 126074; OP464999), and 98.24% with Chl. guttiferum (CBS 126073; OP464991). BLAST results for all sequence regions of the strain showed > 99% identity with Chl. chlamydosporum. These results reveal that the strain KNUF-21-010 is Chl. chlamydosporum. To clarify this identification, MLSA was conducted using the ITS regions, LSU, and tef1-α. The phylogenetic tree (Fig. 3), constructed from the combined three loci (ITS, LSU, and tef1-α), placed KNUF-21-010 in a clade with Chl. chlamydosporum, Chl. peruense, and Chl. detriticola (90% bootstrap). Among the species analyzed, KNUF-21-010 was clearly distinct from Chl. peruense and Chl. detriticola. Within this clade, the strain formed a small clade with Chl. chlamydospore (CBS 149052) and Chl. chlamydosporum (CBS 114.41T) with 95% bootstrap. KNUF-21-010 and Chl. chlamydosporum (CBS 149052) clustered with 100% bootstrap, indicating that KNUF-21-010 represents a new strain of Chl. chlamydosporum.
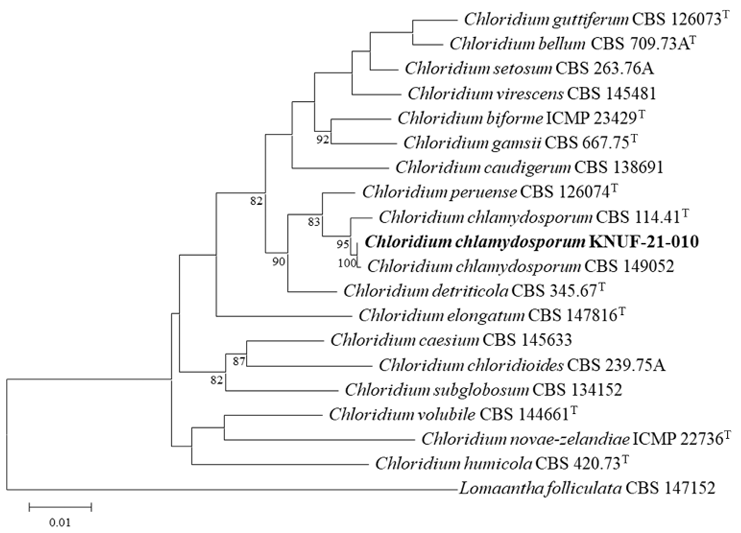
Fig. 3. The maximum likelihood phylogenetic tree based on the internal transcribed spacer (ITS) regions, large subunit of 28S rRNA (LSU), and translation elongation factor 1-alpha (tef1-α) sequences. The tree shows the position of the KNUF-21-010 within the genus Chloridium. The fungal isolate newly determined in this study is highlighted in bold. Bootstrap values (> 80%) based on 1000 replicates were shown at nodes. Bar, 0.01 substitutions per nucleotide position.
Chloridium is a saprophytic fungal genus, with Chl. virescens which is designated as the type species [3]. In this study, Chl. chlamydosporum, a species not previously observed in Korea, was isolated from soil samples collected in Yangsan-si, Gyeongsangnam-do. For identification, its cultural and morphological characteristics, and genetic data were examined. Based on MLSA of the ITS regions, LSU, and tef1-α, the phylogenetic tree shows that our strain clustered in a distinct clade with the two reference strains of Chl. chlamydosporum (CBS 114.41T and CBS 149052), thereby supporting its identification as Chl. chlamydosporum. Fungi of the family Chaetosphaeriaceae, which is one of the largest families within the class Sordariomycetes, are widely distributed and typically inhabit decaying leaves, fruits, and woody substrates in terrestrial and aquatic ecosystems [2,3].
The family Chaetosphaeriaceae was first reveal by Locquin in 1984 [18]. However, because it lacked a description compliant with the International Code of Botanical Nomenclature, it was considered invalidly published and was subsequently validated by Réblová et al. (1999) [19]. The introduction of molecular phylogenetic approaches into fungal classification has significantly enhanced taxonomic and phylogenetic understanding, resulting in continuous reclassification within this family [2]. Owing to the presence of closely related polyphyletic genera within the Chaetosphaeriaceae family, accurate classification of genera, including Chaetosphaeria, Chloridium, Codinaea, Cryptophiale, Dictyochaeta and Kionochaeta remains necessary [2].
After Link established the genus Chloridium, von Höhnel (1903) identified a new genus, Cirrhomyces, with C. caudiger designated as the type species [3]. Cirrhomyces was distinguished from Chloridium based on the formation of multiple conidia. C. caudiger is characterized based on dematiaceous, fasciculate conidiophores and hyaline, oblong conidia adhering in white cirrhi. In 1940, van Beyma described Bisporomyces chlamydosporum, a fungus distinguished by its unique structure of paired conidia at the tip of the phialide [7]. Owing to the morphological similarities among B. chlamydosporum, Chl. virescens, and C. caudiger, considerable debate arose regarding their taxonomic placement. Hughes (1958) synonymized Cirrhomyces and Bisporomyces with Chloridium, recognizing five species within the genus: Chl. botryoideum, Chl. caudigerum, Chl. chlamydosporum, Chl. minutum, and Chl. virescens [8]. Gams and Holubová-Jechová (1976) subsequently classify Chloridium into three sections—Chloridium, Gongromeriza, and Psilobotrys—based on morphological characteristics such as the conidiogenous locus and the presence of chlamydospores [4].
In Chloridium, molecular data for some species have only been studied in the 21st century [3]. Traditionally, Chloridium was classified based on morphological traits; the recent application of molecular phylogenetics has prompted ongoing revisions at the genus and species levels. Based on molecular analyses of the LSU sequence, the hypothesis dividing the genus into three sections was rejected [20], and several studies have demonstrated the polyphyletic nature of Chloridium [2,3,9,10]. Wu and Diao (2022) attempt to redefine Chloridium based on ITS and LSU data, considering it paraphyletic, including parts of Adautomilanezia and Sporoschisma [2]. However, Réblová et al. (2022) demonstrate that Chloridium forms a separate monophyletic clade distinct from the Adautomilanezia and Sporoschisma group, using ITS and LSU data refined with Gblocks [3].
Chloridium has traditionally been regarded as a saprophytic genus primarily occurring on wood or other plant debris [4], but some species have also been reported to be isolated from soil [3]. In particular, Chloridium may contribute to increasing nitrogen-use efficiency and agricultural productivity in acidic soil ecosystems through heterotrophic nitrification [21]. Therefore, given its isolation from Korean soils, Chl. chlamydosporum warrants further investigation to clarify its potential roles in organic matter decomposition and contribution to the nitrification process within these soil ecosystems.
This study provides the first evidence that Chl. chlamydosporum occurs in Korean soils, contributing to the ecological understanding and biogeographical expansion of the genus Chloridium. Chloridium is a globally distributed fungal genus; however, only a few species have been revealed in Korea, indicating that research on this group remains in its early stages [22]. Therefore, additional research is required to identify previously unrecorded or novel Chloridium species and to better understand their distribution and diversity in Korean ecosystems.
The authors declare no potential conflicts of interest.
This work was supported by a grant from the National Institute of Biological Resources, funded by the Ministry of Environment of the Republic of Korea (NIBR202102107).
1. Link HF. Observationes in ordines plantarum naturales. Dissertatio Ima. Mag Neuesten Entdeck Gesammten Naturk Ges Naturf Freunde Berlin 1809;3:3-42.
2. Wu W, Diao Y. Anamorphic chaetosphaeriaceous fungi from China. Fungal Divers 2022;116:1-546. https://doi.org/10.1007/s13225-022-00509-w [DOI]
3. Réblová M, Hernández-Restrepo M, Sklenář F, Nekvindová J, Réblová K, Kolařík M. Consolidation of Chloridium: New classification into eight sections with 37 species and reinstatement of the genera Gongromeriza and Psilobotrys. Stud Mycol 2022;103:87-212. https://doi.org/10.3114/sim.2022.103.04 [DOI]
4. Gams W, Holubová-Jechová V. Chloridium and some other dematiaceous hyphomycetes growing on decaying wood. Stud Mycol 1976;13:1-99. [DOI]
5. Domsch KH, Gams W, Anderson TH. Compendium of soil fungi. Volume 1. London: Academic Press; 1980.
6. Omvik A. Morphology and Nutrition of Chloridium chlamydosporis (Bisporomyces chlamydosporis). Mycologia 1970;62:209-26. https://doi.org/10.1080/00275514.1970.12018960 [DOI]
7. van Beyma Thoe Kingma FH. Beschreibung einiger neuer Pilzarten aus dem Centraalbureau voor Schimmelcultures, Baarn (Nederland) VI. Mitteilung. Antonie Van Leeuwenhoek 1939;6:263-90. [DOI]
8. Hughes SJ. Revisiones hyphomycetum aliquot cum appendice de nominibus rejiciendis. Can J Bot 1958;36:727-836. https://doi.org/10.1139/b58-067 [DOI]
9. Fehrer J, Réblová M, Bambasová V, Vohník M. The root-symbiotic Rhizoscyphus ericae aggregate and Hyaloscypha (Leotiomycetes) are congeneric: Phylogenetic and experimental evidence. Stud Mycol 2019;92:195-225. https://doi.org/10.1016/j.simyco.2018.10.004 [DOI]
10. Crous PW, Wingfield MJ, Schumacher RK, Akulov A, Bulgakov TS, Carnegie AJ, Jurjević Ž, Decock C, Denman S, Lombard L, et al. New and interesting fungi. 3. Fungal Syst Evol 2020;6:157-231. https://doi.org/10.3114/fuse.2020.06.09 [DOI]
11. Sinclair JB, Dhingra OD. Basic plant pathology methods. 2nd ed. Boca Raton: CRC Press; 1995.
12. de Hoog GS, Gerrits van den Ende AH. Molecular diagnostics of clinical strains of filamentous Basidiomycetes. Mycoses 1998;41:183-9. https://doi.org/10.1111/j.1439-0507.1998.tb00321.x [DOI]
13. Vilgalys R, Hester M. Rapid genetic identification and mapping of enzymatically amplified ribosomal DNA from several Cryptococcus species. J Bacteriol 1990;172:4238-46. https://doi.org/10.1128/jb.172.8.4238-4246.1990 [DOI]
14. Rehner SA, Buckley E. A Beauveria phylogeny inferred from nuclear ITS and EF1alpha sequences: Evidence for cryptic diversification and links to Cordyceps teleomorphs. Mycologia 2005;97:84-98. https://doi.org/10.3852/mycologia.97.1.84 [DOI]
15. Landvik S. Neolecta, a fruit-body-producing genus of the basal ascomycetes, as shown by SSU and LSU rDNA sequences. Mycol Res 1996;100:199-202. https://doi.org/10.1016/S0953-7562(96)80122-5 [DOI]
16. Kimura M. A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 1980;16:111-20. https://doi.org/10.1007/BF01731581 [DOI]
17. Kumar S, Stecher G, Suleski M, Sanderford M, Sharma S, Tamura K. MEGA12: Molecular evolutionary genetic analysis version 12 for adaptive and green computing. Mol Biol Evol 2024;41:1-9. https://doi.org/10.1093/molbev/msae263 [DOI]
18. Zhang JY, Hyde KD, Ma J, Wu N, Al-Otibi F, Zhang LJ, Lu YZ. Morpho-phylogenetic evidence reveals Pseudolomaan thathailandica gen. et sp. nov. and Submultiguttulispora multiseptata gen. et sp. nov. in Chaetosphaeriaceae. MycoKeys 2025;113:123-46. https://doi.org/10.3897/mycokeys.113.142643 [DOI]
19. Réblová M, Barr ME, Samuels GJ. Chaetosphaeriaceae, a new family for Chaetosphaeria and its relatives. Sydowia 1999;51:49-70. [DOI]
20. Réblová M, Winka K. Phylogeny of Chaetosphaeria and its anamorphs based on morphological and molecular data. Mycologia 2000;92:939-54. https://doi.org/10.1080/00275514.2000.12061238 [DOI]
21. Zhang Y, Dai S, Huang X, Zhao Y, Zhao J, Cheng Y, Cai Z, Zhang J. pH-induced changes in fungal abundance and composition affects soil heterotrophic nitrification after 30 days of artificial pH manipulation. Geoderma 2020;366:114255. https://doi.org/10.1016/j.geoderma.2020.114255 [DOI]
22. Azmi MH, Lim SK, Lee SM, Kang DH, Choi JS, Lim GJ, Choi JW, Kim MK, Lee SY, Jung HY. New records of two unreported fungal species: Cladophialophora floridana and Chloridium setosum isolated from soil in Korea. Kor J Mycol 2024;52:165-75. http://doi.org/10.4489/kjm.520302 [DOI]