Tae-Gyeong Kim1, San San Yu2, Seong-Keun Lim1, So-Jeong Ye1, Leonid N. Ten3, Seung-Yeol Lee1,3, and Hee-Young Jung1,3*
1Department of Plant Medicine, Kyungpook National University, Daegu 41566, Korea
2Department of Biotechnology Research, Ministry of Science and Technology, Kyaukse 05151, Myanmar
3Institute of Plant Medicine, Kyungpook National University, Daegu 41566, Korea
*Correspondence to heeyoung@knu.ac.kr
Korean Journal of Mycology (Kor J Mycol) 2025 December, Volume 53, Issue 4, pages 237-243.
https://doi.org/10.4489/kjm.2025.53.4.1
Received on October 17, 2025, Revised on November 10, 2025, Accepted on November 10, 2025, Published on December 31, 2025.
Copyright © The Korean Society of Mycology.
This is an Open Access article which is freely available under the Creative Commons Attribution-Non-Commercial 4.0 International License (CC BY-NC) (https://creativecommons.org/licenses/by-nc/4.0/).
Two fungal strains, designated KNUF-20-NI013 and KNUF-21-043, were isolated from soil samples collected in Jeollabuk-do and Gyeongsangbuk-do provinces, Korea. Strain KNUF-20NI013 exhibited cultural characteristics typical of the genus Arcopilus, including pinkish-red mycelia, predominantly submerged growth, and smooth colony margins. Sequence analysis of internal transcribed spacer (ITS), 28S ribosomal DNA large subunit (LSU), β-tubulin (TUB2), and the second-largest subunit of RNA polymerase II (RPB2) regions revealed over 99% similarity compared to the corresponding sequences of Arcopilus fusiformis. A phylogenetic tree constructed based on the combined sequences of these four loci confirmed the affiliation of KNUF-20-NI013 with A. fusiformis. Strain KNUF-21-043 displayed morphological characteristics consistent with Schizothecium glutinans, including grayish-green filamentous colonies with a filiform margin and powdery surface. This strain also produced hyaline smooth-walled phialides that were solitary or scattered and variably swollen at the middle or base, together with hyaline smooth phialoconidia that were subglobose to broadly obovoid and had a truncated base; these are characteristic features of S. glutinans. The phylogenetic tree based on the concatenated ITS and LSU sequences supported the identification of KNUF-21-043 as S. glutinans. To the best of our knowledge, these strains represent the first record of A. fusiformis and S. glutinans in Korea.
Arcopilus fusiformis, Phylogeny, Schizothecium glutinans, Soil fungi, Unreported species
The genus Arcopilus, classified within the class Sordariomycetes, subclass Sordariomycetidae, order Sordariales, and family Chaetomiaceae, was established by Wang et al. (2016), with A. aureus being designated as the type species [1]. In that study, Wang et al. introduced five new genera, Amesia, Arcopilus, Collariella, Dichotomopilus, and Ovatospora, based on morphological observations and multigene phylogenetic analyses using sequences of the internal transcribed spacer (ITS) region, 28S large subunit (LSU) rDNA, β-tubulin (TUB2), and the second-largest subunit of RNA polymerase II (RPB2). The genus Arcopilus was delineated through the reclassification of several species previously assigned to Chaetomium, including A. aureus (formerly Chaetomium aureum), A. cupreus (C. cupreum), A. fusiformis (C. fusiforme), A. flavigenus (C. flavigenum), and A. turgidopilosus (C. turgidopilosum) [1]. Since then, several additional species have been described as novel members of this genus, including A. megaspores [2], A. macrostiolatus [2], A. amazonicus [3], and A. albae [4]. Currently, the Index Fungorum (https:// www.indexfungorum.org) recognizes 15 species within the genus Arcopilus. Among these, A. fusiformis was originally described as Chaetomium fusiforme by Chivers in 1912 [5] and was later transferred to the genus Arcopilus by Wang et al. (2016) [1]. Members of the genus Arcopilus are characterized by arcuate ascomatal hairs and often produce colonies exhibiting various hues—ranging from yellow and orange to red and rust—due to pigmented ascomata and exudates [1]. Notably, the asexual morphs of Arcopilus species remain undescribed, highlighting the need for further biological investigations [1,2]. To date, only A. aureus has been officially reported in Korea [6]. Therefore, studies on Arcopilus species isolated from domestic environments are essential for enhancing our understanding regarding fungal biodiversity in Korea and for exploring their potential biotechnological applications.
The genus Schizothecium was first established by Corda in 1838, with S. fimicola being designated as the type species [7]. This coprophilous ascomycete is characterized by perithecia bearing agglutinated hairs and a distinctive ascomatal structure. However, Schizothecium was mistakenly synonymized with Podospora by Cesati in 1856, which resulted in decades of taxonomic confusion between the two genera [8]. This issue was resolved by Lundqvist in 1972, who reinstated Schizothecium as a distinct genus based on key morphological differences [9]. These distinguishing features include swollen—often agglutinated— ascomatal hairs, the absence of true paraphyses, early septation of ascospores, and persistent plasma-filled pedicels. Subsequently, molecular phylogenetic analyses further confirmed this taxonomy. Cai et al. (2005) used sequences from the ITS region, LSU, and TUB2 to demonstrate that Schizothecium forms a wellsupported monophyletic lineage distinct from Podospora [10]. Their findings indicated that ascomatal morphology is more informative than ascospore characteristics for delineating taxa within the order Sordariales. More recently, Marin-Felix et al. [11] formally established the family Schizotheciaceae to accommodate a well-supported lineage previously nested within the polyphyletic family Lasiosphaeriaceae. This taxonomic revision provides greater stability for the classification of Schizothecium and related taxa. According to the current taxonomic consensus, approximately 30 species of the genus Schizothecium are recognized in the Index Fungorum and MycoBank (https://www.mycobank.org) databases. These species are distributed across temperate and subtropical regions and have been isolated from various substrates, including herbivore dung, plant debris, and soil. Collectively, this genus exhibits broad ecological adaptability and is now widely recognized as an independent phylogenetic lineage through integrated morphological and molecular studies.
This study was conducted as part of an ongoing effort to investigate the diversity of indigenous fungal species in Korea. Specifically, we aimed to identify and characterize soil-derived fungal isolates using both morphological and molecular approaches. The isolates were identified as A. fusiformis (KNUF-20-NI013) and Schizothecium glutinans (KNUF-21-043), representing the first report of these species in Korea.
The fungal isolates analyzed in this investigation were obtained from a rhizosphere soil sample of Selaginella tamariscina collected in Okdo-myeon, Gunsan-si, Jeollabuk-do province (35°48’57.8″N, 126° 23’45.5″E) and from rhizosphere soil of broadleaved forest in Uiseong-gun, Gyeongsangbuk-do province (36° 15’48.7″N, 128°41’24.5″E), Korea. The soil samples were collected from the surface to a depth of 20 cm, placed in sterile plastic bags, transported to the laboratory, and then stored at 4°C until further processing. One gram of soil sample was mixed with 10 mL of sterile distilled water, vortexed, and serially diluted. Aliquots (50–100 µL) of each suspension were subsequently spread onto potato dextrose agar (PDA; Difco, Detroit, MI, USA) in Petri dishes. The rate of fungal colony growth was evaluated by incubating the Petri dishes at 25°C until visible mycelial development was observed. To preserve the fungal strains, they were suspended in 20% (w/v) glycerol and stored at -80°C. Preliminary identification of several isolated fungal strains was performed through sequencing their ITS regions. Among them, the KNUF-20-NI013 and KNUF-21-043 strains were selected for detailed phylogenetic analysis and morphological characterization, as they appeared to represent fungal species not previously documented in Korea. Stock cultures of the isolates were deposited and maintained under inactive conditions at the National Institute of Biological Resources (NIBR).
Cultural characteristics of the fungal isolate KNUF-20-NI013 were recorded using different media, including PDA, oatmeal agar (OA; Difco, Detroit, MI), malt extract agar (MEA; Difco, Detroit, MI), potato carrot agar (PCA; Difco, Detroit, MI), and synthetic nutrient deficient agar (SNA; Difco, Detroit, MI) for f ive days at 25℃ [1]. Strain KNUF-21-043 was grown on PDA and corn meal agar (CMA; Difco, Detroit, MI) for 21 days at 24℃ [12,13]. Examination of morphological characteristics was conducted using a light microscope (BX-50; Olympus, Tokyo, Japan); the colony characteristics of the fungal isolate, including color, size, and shape of the colonies on the media, were recorded.
Total genomic DNA of strains KNUF-20-NI013 and KNUF-21-043 was extracted from 7-day-old mycelia cultured on PDA and CMA, respectively, using the HiGene Genomic DNA Prep Kit (BIOFACT, Daejeon, Korea) according to the manufacturer’s protocols. The primers used for PCR amplification included: ITS1F/ITS4 for ITS regions [14], LR0R/LR5 for LSU [15,16], T1/T4 for TUB2 [17], and rpb25F2/rpb2-7cR for RPB2 [18]. To confirm their yields, PCR products were electrophoresed on a 1% agarose gel. The amplified fragments were subsequently purified using EXOSAP-IT (Thermo Fisher Scientific, Waltham, MA, USA) and sequenced by Solgent (Daejeon, Korea) and Macrogen (Seoul, Korea). Analysis of the obtained sequence data was performed using SeqMan Lasergene software (DNAStar Inc., Madison, Wisconsin, USA). The sequences of KNUF-20-NI013 were subsequently deposited in the GenBank database under the accession numbers LC881688 (ITS), PV942136 (LSU), PV945815 (TUB2), and PV945814 (RPB2), while those of KNUF-21-043 were deposited under LC881689 (ITS) and PV942137 (LSU).
Molecular phylogenetic analyses were performed using either four or two genetic markers. The ITS, LSU, TUB2, and RPB2 loci were analyzed in KNUF-20-NI013, while the ITS and LSU regions were analyzed in KNUF-21-043. For identification, both individual and concatenated sequences were compared with reference sequences retrieved from the GenBank database of the National Center for Biotechnology Information (Table 1). Multiple sequence alignments were conducted using ClustalW integrated in MEGA version 7.0 [19]. Phylogenetic trees were constructed using the maximum likelihood (ML), neighbor joining (NJ), and maximum parsimony (MP) methods. Evolutionary distances for the NJ method were calculated using Kimura’s two-parameter model [20], and statistical support for tree topologies was assessed using 1,000 bootstrap replicates. All analyses were conducted in MEGA 7.0 [19].
Table 1. GenBank accession numbers used for the phylogenetic analyses in this study
테일블
ITS: internal transcribed spacer regions; LSU: 28S rDNA large subunit; TUB2: β-tubulin; RPB2: the second largest subunit of RNA polymerase II. TType strain. The strains isolated in this study are indicated in boldface.
Arcopilus fusiformis X. Wei Wang & Samson, Stud. Mycol. 84:217 (2016) (Fig. 1).
Specimen examined: Gunsan-si, Okdo-myeon, Jeollabuk-do province (35°48’57.8″N, 126°23’45.5″E), strain KNUF-20-NI013 (NIBRFGC000507890) isolated from the soil.
Cultural and morphological characteristics
The cultural characteristics of A. fusiformis KNUF-20-NI013 were examined on five different media: PDA, OA, MEA, SNA, and PCA, after five days of incubation at 25°C. Colonies grown on PDA reached 30–31 mm in diameter and appeared white on the surface, with reddish pigmentation on the reverse side (Fig. 1A and B). On OA, colonies measured 28–29 mm in diameter, displayed white with brown-colored exudates, and exhibited a brown to dark brown coloration on the reverse (Fig. 1C and D). On MEA, colonies were 28–30 mm in diameter, with pinkish-red mycelium visible on both the surface and reverse. The colony grew slowly, was primarily submerged in the medium, and had smooth margins (Fig. 1E and F). Colonies on SNA measured 22–24 mm exhibiting a light-yellow surface and yellow pigmentation on the reverse (Fig. 1G and H). Growth on PCA was characterized by a white surface with an ambercolored center and ginger-colored concentric rings on the reverse side (Fig. 1I and J). No anamorphic stage was observed under the examined conditions, even after incubation for up to two months. Although the closely related species A. flavigenus usually has a white aerial mycelium that turns red or orange due to a red pigment exudate, mature colonies 10–14 days develop a green-to-brown coloration in the mycelium [21]. This green-to-brown coloration was not observed in strain KNUF-20-NI013. In contrast, the cultural features of KNUF-20-NI013 aligned with those of A. fusiformis and differed from those of A. flavigenus (Table 2).
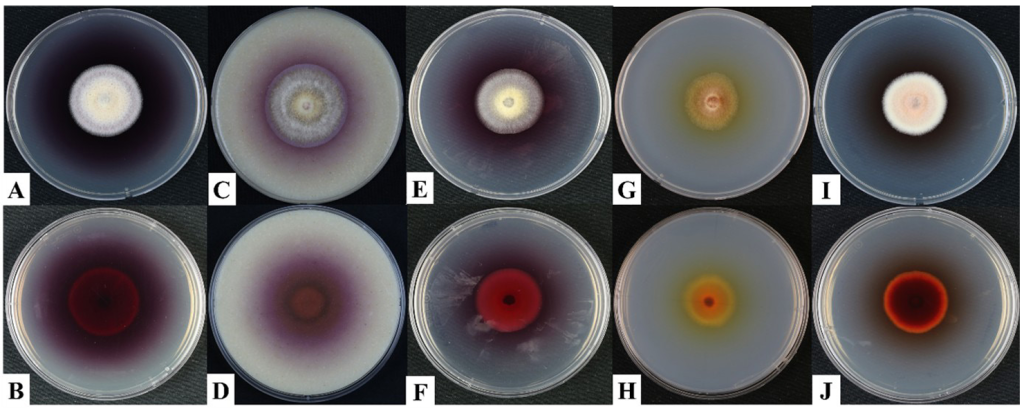
Fig. 1. Cultural characteristics of Arcopilus fusiformis KNUF-20-NI013. A: Colony on potato dextrose agar (PDA) after 5 days at 25°C; B: Reverse side on PDA; C: Colony on oatmeal agar (OA) after 5 days at 25°C; D: Reverse side on OA; E: Colony on malt extract agar (MEA) after 5 days at 25°C; F: Reverse side on MEA; G: Colony on synthetic nutrient deficient agar (SNA) after 5 days at 25°C; H: Reverse side on SNA; I: Colony on potato carrot agar (PCA) after 5 days at 25°C; J: Reverse side on PCA.
Table 2. Comparison of cultural characteristics of strain KNUF-20-0NI013 with reference Arcopilus species
테이블
PDA: potato dextrose agar; OA: oatmeal agar; MEA: malt extract agar; SNA: synthetic nutrient deficient agar; PCA: potato carrot agar; N/A: not available in previous study.
aFungal strain investigated in this study; bSources of description [1]; cSources of description [21].
Molecular phylogeny analysis
For the molecular identification of isolated strain KNUF-20-NI013, total genomic DNA was amplified to obtain the ITS region, LSU, TUB2, and RPB2 sequences, yielding lengths of 611, 870, 657, and 1048 bp, respectively. The ITS region of strain KNUF-20-NI013 showed high similarity to A. cupreus TMDF101 (100%, JQ676206), A. flavigenus CBS 337.67T (99.84%, MH858989), A. tangerinicapillus WZ-951 (99.83%, OP163629), and A. fusiformis CBS 484.85 (99.64%, KX976585). Based on the LSU sequence, the species with high sequence similarity to strain KNUF-20-NI013 were A. aureus CBS 140.3 (99.65%, MH867117), A. navicularis CCF 3252 (99.54%, MW798181.1), A. fusiformis CBS 788.71 (99.54%, MH872105.1), and A. cupreus CBS 560.80 (99.54%, MH873060.1). The TUB2 sequence of strain KNUF-20-NI013 showed high similarity to A. fusiformis CBS 484.85 (99.54%, KX976927.1) and A. f lavigenus CBS 337.67T (98.93%, KX976929.1), while notably lower similarities were observed with other Arcopilus species including A. megasporus CBS 127650 (86.45%, MZ343010), A. purpurascens CBS 287.83 (86.23%, MZ343021), A. macrostiolatus CBS 102435T (85.95%, MZ343006), A. turgidopilosus CBS 169.52T(83.48%, KX976930), and A. amazonicus AM2411 (82.33%). The partial RPB2 sequence of the isolate exhibited the highest similarity to A. flavigenus CBS 337.67T (99.43%), A. eremanthusum CML 3766 (99.41%), and A. fusiformis CBS 484.85 (99.04%), whereas all other Arcopilus species showed similarity values below 93%. These results indicated that several Arcopilus species share high genetic similarity with strain KNUF-20-NI013, making it difficult to achieve precise species-level identification using any single locus. The ML phylogenetic tree based on the combined dataset (Fig. 2) showed that strain KNUF-20-NI013 consistently clustered with A. fusiformis, indicating its closest phylogenetic relationship at the species level. The tree topologies generated using NJ and MP methods were congruent with those of the ML tree, providing additional support for the identification of the isolate as A. fusiformis. Based on the combined molecular evidence and culture characteristics (Table 2), strain KNUF-20-NI013 was identified as A. fusiformis. To the best of our knowledge, this is the first report of this species in Korea.
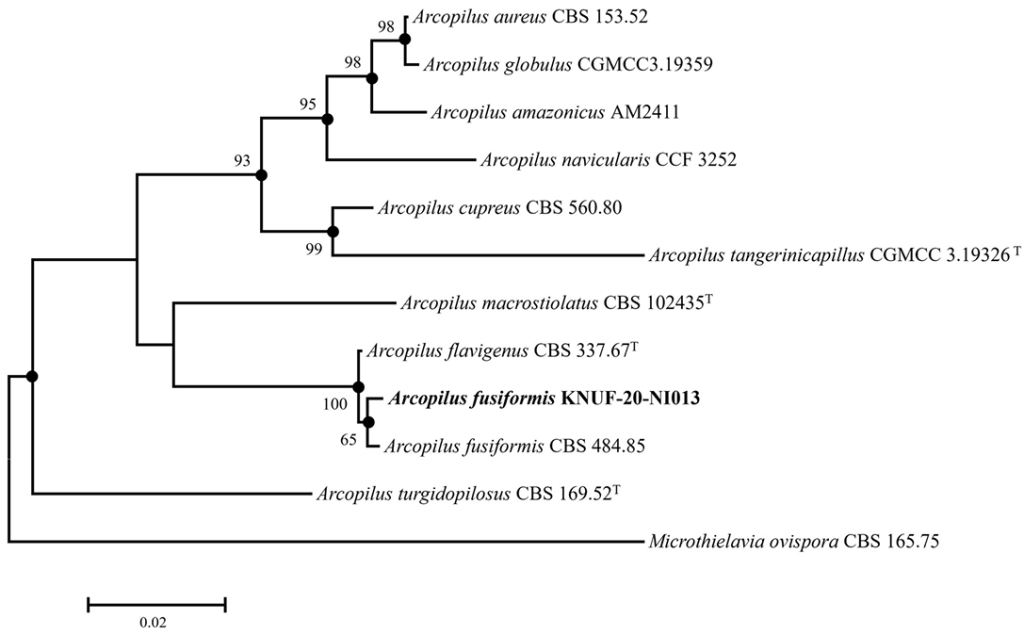
Fig. 2. Maximum-likelihood phylogenetic tree based on the combined sequences of internal transcribed spacer (ITS) regions, the 28S ribosomal RNA (LSU), β-tubulin (TUB2) and RNA polymerase II second largest subunit gene (RPB2) genes showing the phylogenetic position of strain KNUF-20-NI013 among Arcopilus species. Bootstrap values greater than 60% (based on 1,000 replications) are shown at branch points. The filled circles indicate that the corresponding nodes were also recovered in the trees generated using the neighbor joining and maximum parsimony algorithms. The isolated strain is indicated in bold. Microthielavia ovispora CBS 165.75 was used as an outgroup. Bar, 0.02 substitutions per nucleotide position.
Schizothecium glutinans N. Lundq., Acta Univ. Upsal., Symb. Bot. Upsal. 20(1): 254 (1972)
Specimen examined: Uiseong-gun, Gyeongsangbuk-do province (36°15’48.7″N, 128°41’24.5″E), strain KNUF-21-043 (NIBRFGC000509194) isolated from the soil.
Cultural and morphological characteristics
After 21 days at 24°C on PDA, the colony diameter of strain KNUF-21-043 ranged from 66–69 mm, exhibiting a grayish greenish surface and black pigmentation on the reverse side (Fig. 3A and B). The colonies were flat, velvety-to-powdery in texture, filamentous in form, and had filiform margins (Fig. 3A and B). On CMA, colonies were 63–69 mm in diameter and olivaceous grey in color; they showed gradually spreading, appressed growth with mycelia mostly submerged in the medium, closely resembling that of the reference species S. glutinans CBS 134.83 (Fig. 3C and D). The phialides of KNUF-21-043 were hyaline, smooth-walled, and occurred solitarily or scattered directly on the substrate hyphae without conidiophores. They measured 18.7–25.6 × 3.4–6.2 µm and varied in shape from long, narrow, hypha-like to slightly swollen at the middle or base, while being occasionally pyriform. A flared apical collarette was consistently observed (Fig. 3E and F). Phialoconidia were hyaline, smooth-walled, and measured 3.2–4.3 × 2.5–3.4 µm. They were typically produced singly and aggregated in a glioid mass at the apex of the phialide, with shapes ranging from subglobose to broadly obovoid and exhibiting truncate bases. Short chains of conidia were occasionally observed (Fig. 3E and F). These results confirmed that the morphological characteristics of KNUF-21-043 closely aligned with those of the reference species S. glutinans CBS 134.83 (Table 3).
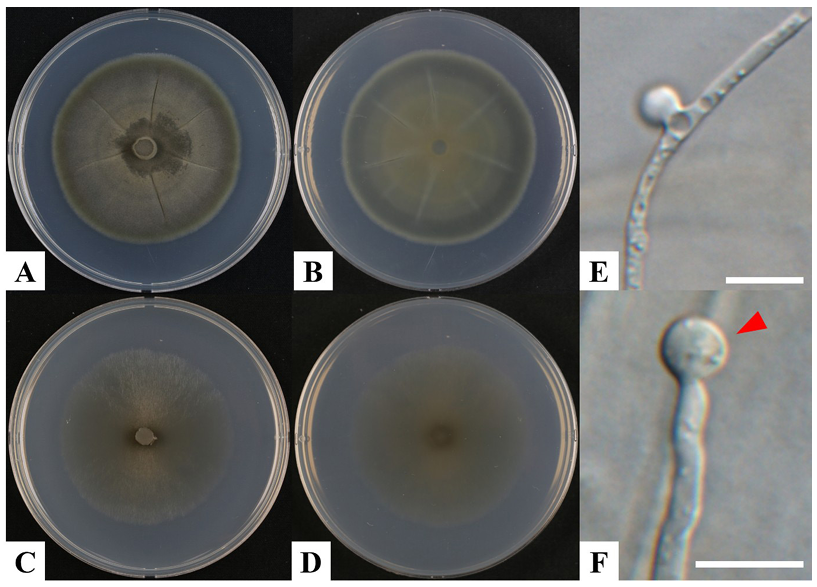
Fig. 3. Cultural and morphological characteristics of Schizothecium glutinans KNUF-21-043. A, B: Colony on potato dextrose agar (PDA) after 21 days at 24°C; C, D: Colony on corn meal agar (CMA) after 21 days at 24°C; E, F: Phialide, Phialoconidia (indicated by arrowhead). Scale bars E– F = 10 µm.
Table 3. Morphological characteristics of the isolated strain KNUF-21-043 with reference to Schizothecium gltuinans
table
PDA: potato dextrose agar; CMA: corn meal agar; N/A: not available in previous study.
aFungal strain investigated in this study; bSources of description [12,13].
Molecular phylogeny analysis
The amplicons obtained from the ITS and LSU regions of strain KNUF-21-043 were 562 and 897 bp, respectively. The ITS sequence of the isolate exhibited high similarity (99.43–98.52%) with several strains of S. glutinans, including A620 (MK247326), CBS 113105 (MH862915), CBS 11310 (AY615208.1), S82 (KT224860.1), and TB64 (PQ032307.1). In contrast, the next closest species, Schizothecium vesticola CBS 366.69 (MH859325) and Schizothecium miniglutinans CBS 131.94 (AY515362), showed lower similarities of 96.11% and 96.03%, respectively. Based on the LSU sequence of strain KNUF-21-043, S. glutinans CBS 113105 (MH874488.1) exhibited 100% sequence similarity, confirming it as the closest relative. Additionally, high LSU sequence similarities were observed with other Schizothecium species, including S. minicauda (CBS 227.87T, MH873757.1; 99.66%), S. tetrasporum (CBS 394.87, MH873776.1; 99.55%), S. inaequale (CBS 239.71, MH871870.1; 99.55%; CBS 226.87, AY999094.1; 99.41%), S. carpinicola (CBS 228.87, NG_057741.1; 99.40%), and S. vesticola (SMH3187, AY780076.1; 99.32%). These results indicated that strain KNUF-21-043 is most closely related to S. glutinans based on both ITS and LSU sequence data. However, several other Schizothecium species exhibited LSU sequence similarities exceeding the commonly accepted threshold for species-level distinction, underscoring the limitations of relying on a single genetic marker for accurate identification. Recent studies have demonstrated the effectiveness of combined sequence phylogenetic approaches in resolving complex relationships within Schizothecium [11,22]. A multilocus sequence analysis (MLSA) was performed using concatenated ITS and LSU sequences to accurately determine the taxonomic position of strain KNUF-21-043 (Table 1). The ML phylogenetic tree based on the combined dataset (Fig. 4) firmly placed strain KNUF-21-043 within the genus Schizothecium, and the resulting topology clearly identified it as S. glutinans. Moreover, the NJ and MP methods produced similar tree topologies, as indicated by the filled circles in Fig. 4, further supporting the phylogenetic placement of the isolate. The combined results of the morphological and molecular analyses confirmed that strain KNUF-21-043 belonged to S. glutinans. To the best of our knowledge, this is the first report on this fungal species in Korea.
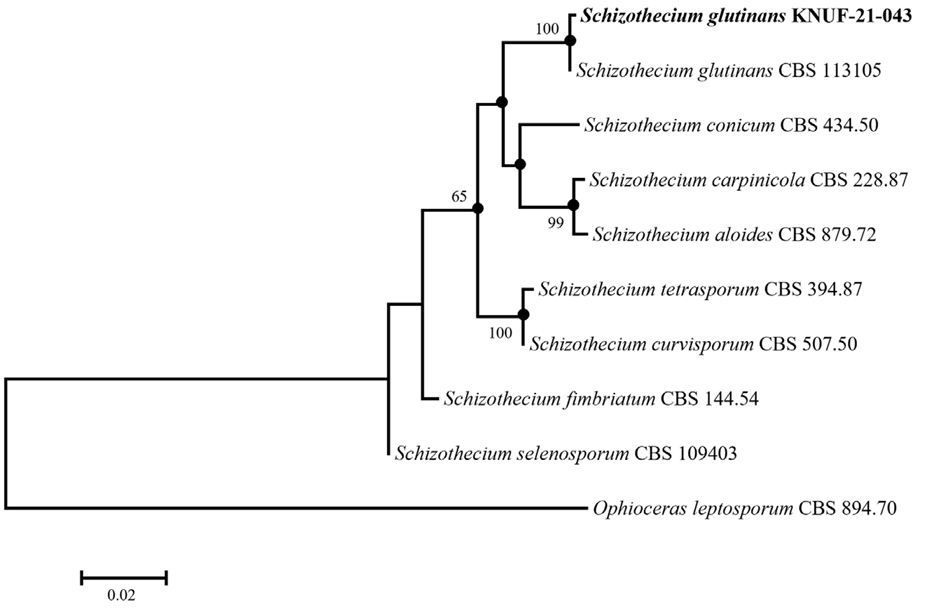
Fig. 4. Maximum-likelihood phylogenetic analysis of KNUF-21-043 based on the 28S ribosomal RNA gene (LSU) and the internal transcribed (ITS) sequence showing the phylogenetic position of the closest species in the genus Schizothecium. Bootstrap values greater than 60% (percentage of 1,000 replications) are shown at branching points. The filled circles indicate that the corresponding nodes were also recovered in the trees generated using the neighbor joining and maximum parsimony algorithms. The strain isolated in this study was highlighted in bold. Ophioceras leptosporum CBS 894.70 was used as an outgroup. Bar, 0.02 substitutions per nucleotide position.
The genus Arcopilus is known to inhabit a broad range of cellulose-rich substrates, including soil, marine environments, animal dung, plant tissues, hair, seeds, textiles, and wood [1,2]. According to the Mycobank database (https://www.mycobank.org/), A. fusiformis has been reported in diverse environments, such as soil, decaying wood, animal feces, and discarded textiles, with occurrences documented in Sri Lanka, Suriname, the United States, and Kenya. These findings suggest that A. fusiformis possesses considerable ecological adaptability, enabling it to thrive under varied environmental conditions. Arcopilus species are being increasingly recognized as prolific producers of diverse secondary metabolites with a wide range of biological activities. Among them, A. aureus is the most extensively studied. Several A. aureus strains have been investigated for the production of resveratrol, a polyphenolic compound known for its anti-inflammatory, antioxidant, anti-aging, and chemopreventive properties, including protection against diabetes, cardiovascular diseases, and cancer [23]. Additionally, A. aureus has been identified as a non-toxic endophytic fungus capable of producing the yellow pigment cochlioquinol II, which demonstrates excellent thermal stability and holds considerable promise as a natural food colorant for future industrial applications [3]. Another strain, A. aureus HJ-7, was reported to synthesize a novel compound, xanthoradone D, which exhibits potent antibacterial activity against Staphylococcus aureus and Xanthomonas axonopodis as well as antifungal effects against Cochliobolus miyabeanus and Ceratocystis paradoxa [24]. Other species in this genus have also demonstrated notable bioactivities. For instance, A. navicularis was recently found to produce a series of novel tetramic acid derivatives named arcopilins A–G. Notably, arcopilin A was shown to effectively disrupt biofilms formed by S. aureus [25]. Furthermore, the A. aureus strain YZXR has been explored as a plant-beneficial fungus that exhibits strong antifungal activity against Fusarium fujikuroi, the causal agent of leaf spot disease in Polygonatum odoratum, thereby demonstrating its potential as a biological control agent [4]. Collectively, these findings highlight the genus Arcopilus as a promising source of bioactive secondary metabolites with potential applications in agriculture, medicine, and industry. Despite this growing interest, A. fusiformis has not been investigated for its secondary metabolite production, pigment biosynthesis, or potential as a biocontrol agent. Therefore, the identification of strain KNUF-20-NI013 in this study not only marks the first record of A. fusiformis in Korea but also introduces a promising biological resource for future functional and industrial research.
S. glutinans has been previously reported in various regions across Europe, North America, and Asia— including Belgium, Canada, Denmark, Finland, France, Ireland, Japan, Mexico, Poland, Spain, Sweden, the United Kingdom, and the United States [9,10,26–28]. It is most frequently isolated from coprophilous substrates, particularly herbivore dung of hares and goats, as originally described in Japanese collections [27]. These nutrient-rich substrates support the growth and reproduction of coprophilous fungi such as S. glutinans. However, sporadic isolation from soil and decaying plant material suggests a broader ecological range than previously recognized [9,28]. The isolation of strain KNUF-21-043 from soil in Uiseong-gun, Korea, extends the known ecological and geographical distribution of S. glutinans and suggests a higher degree of ecological plasticity. While closely related genera such as Podospora are largely dung-restricted, Schizothecium species appear capable of colonizing both dung and soil habitats. This ecological versatility may reflect evolutionary differentiation within the order Sordariales and justifies the taxonomic separation of Schizothecium from other genera. As no species of Schizothecium had previously been officially recorded in Korea, this study represents the first confirmed occurrence of S. glutinans in the country, thereby enriching the catalog of indigenous fungal diversity. Both A. fusiformis KNUF-20-NI013 and S. glutinans KNUF-21-043 are part of a broader initiative to document novel native fungal species in Korea, through which several previously unreported taxa have recently been described [29—33]. However, from a practical standpoint, the genus Schizothecium remains poorly studied for industrial or biotechnological applications. To date, no commercial uses of Schizothecium species have been documented. Nevertheless, recent studies have indicated their potential ecological relevance. For instance, Schizothecium has been detected in oil-contaminated soils [34,35], and S. glutinans has been proposed as a potential degrader of polycyclic aromatic hydrocarbons [35]. These findings suggest that S. glutinans may harbor enzymatic pathways useful for bioremediation. Its ability to survive on diverse substrates and tolerate environmental contaminants also makes it a promising subject for studies on soil health, organic matter recycling, and environmental biotechnology.
In conclusion, this study documents the first occurrence of A. fusiformis and S. glutinans in Korea, broadening our understanding of their phylogeographic range and ecological diversity. While single-locus analyses provided preliminary insights, they were insufficient to fully resolve species boundaries; therefore, an MLSA was conducted using concatenated ITS, LSU, TUB2, and RPB2 sequences [1,2]. This approach offered improved phylogenetic resolution, reducing ambiguity and enhancing species delimitation. These findings add to the national fungal inventory and lay the groundwork for future studies exploring the ecological roles, physiological properties, and potential applications of these taxa. Continued exploration of understudied fungi remains crucial for unlocking their hidden potential in environmental and industrial sciences.
The authors declare that they have no potential conflicts of interest.
This work was supported by a grant from the National Institute of Biological Resources, funded by the Ministry of Environment of the Republic of Korea [NIBR202002104, NIBR202102107].
1. Wang XW, Houbraken J, Groenewald JZ, Meijer M, Andersen B, Nielsen KF, Crous PW, Samson RA. Diversity and taxonomy of Chaetomium and chaetomium-like fungi from indoor environments. Stud Mycol 2016;84:145-224. https://doi.org/10.1016/j.simyco.2016.11.005
[DOI]
2. Wang XW, Han PJ, Bai FY, Luo A, Bensch K, Meijer M, Kraak B, Han DY, Sun BD, Crous PW, et al. Taxonomy, phylogeny and identification of Chaetomiaceae with emphasis on thermophilic species. Stud Mycol 2022;101:121-243. https://doi.org/10.3114/sim.2022.101.03
[DOI]
3. Silva LADF, Alves MF, Florêncio Filho D, Takahashi JA, Santos LS, De Carvalho SA. Pigment produced from Arcopilus aureus isolated from grapevines: Promising natural yellow colorants for the food industry. Food Chem 2022;389:132967. https://doi.org/10.1016/j.foodchem.2022.132967
[DOI]
4. Xu R, Song S, Wu G, Hu B, Xie J, Wang X, Xia Y, Zheng S, Peng S, Yuan Y, et al. Biocontrol ability of Arcopilus aureus YZXR against Fusarium fujikuroi causing leaf spot on Polygonatum odoratum. Phytopathol Res 2024;6:68. https://doi.org/10.1186/s42483-024-00285-2
[DOI]
5. Chivers AH. Preliminary diagnoses of new species of Chaetomium. Proc Am Acad Arts Sci 1912;48:83-8. https://doi.org/10.2307/20022812
[DOI]
6. Lee SH, Park HS, Nguyen TTT, Lee HB. Characterization of three species of Sordariomycetes isolated from freshwater and soil samples in Korea. Mycobiology 2019;47:20-30. https://doi.org/10.1080/12298093.2019.1574372
[DOI]
7. Corda ACJ. Icones fungorum hucusque cognitorum. Pragae: J. G. Calve; 1838. p. 43.
8. Cesati V. Explicatio iconum. Hedwigia 1856;1:103-4.
9. Lundqvist N. Nordic Sordariaceae s. lat. Symb Bot Ups 1972;20:1-374.
10. Cai L, Jeewon R, Hyde KD. Phylogenetic investigations of Sordariaceae based on multiple gene sequences and morphology. Mycol Res 2006;110:137-50. https://doi.org/10.1016/j.mycres.2005.09.014
[DOI]
11. Marin-Felix Y, Miller AN, Cano-Lira JF, Guarro J, García D, Stadler M, Huhndorf SM, Stchigel AM. Re-evaluation of the order Sordariales: Delimitation of Lasiosphaeriaceae s. str., and introduction of the new families Diplogelasinosporaceae, Naviculisporaceae, and Schizotheciaceae. Microorganisms 2020;8:1430. https://doi.org/10.3390/microorganisms8091430
[DOI]
12. Bell A, Mahoney DP. Coprophilous fungi in New Zealand. I. Podospora species with swollen agglutinated perithecial hairs. Mycologia 1995;87:375-96. https://doi.org/10.1080/00275514.1995.12026544
[DOI]
13. Zambrano Sánchez PE. Aislamiento y caracterización de hongos presentes en orquídeas del Jardín Botánico de Quito [undergraduate thesis]. Quito: Pontificia Universidad Católica del Ecuador; 2023.
14. White TJ, Bruns TD, Lee SB, Taylor JW. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, editors. PCR protocols: A guide to methods and applications. New York: Academic Press; 1990. p. 315-22.
[DOI]
15. Vilgalys R, Sun BL. Ancient and recent patterns of geographic speciation in the oyster mushroom Pleurotus revealed by phylogenetic analysis of ribosomal DNA sequences. Proc Natl Acad Sci 1994;91:4599-603. https://doi.org/10.1073/pnas.91.10.4599
[DOI]
16. Vilgalys R, Hester M. Rapid genetic identification and mapping of enzymatically amplified ribosomal DNA from several Cryptococcus species. J Bacteriol 1990;172:4238-46. https://doi.org/10.1128/jb.172.8.4238-4246.1990
[DOI]
17. O’Donnell K, Cigelnik E. Two divergent intragenomic rDNA ITS2 types within a monophyletic lineage of the fungus Fusarium are nonorthologous. Mol Phylogenet Evol 1997;7:103-16. https://doi.org/10.1006/mpev.1996.0376
[DOI]
18. Liu YJ, Whelen S, Hall BD. Phylogenetic relationships among ascomycetes: Evidence from an RNA polymerase II subunit. Mol Biol Evol 1999;16:1799-808. https://doi.org/10.1093/oxfordjournals.molbev.a026092
[DOI]
19. Kumar S, Stecher G, Tamura K. MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 2016;33:1870-4. https://doi.org/10.1093/molbev/msw054
[DOI]
20. Kimura M. A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 1980;16:111-20. https://doi.org/10.1007/BF01731581
[DOI]
21. Pornsuriya C, Lin FC, Kanokmedhakul S, Soytong K. New record of Chaetomium species isolated from soil under pineapple plantation in Thailand. J Agric Technol 2008;4:91-103.
22. Atashi Khalilabad A, Fotouhifar KB. Two new species of Ophioceras and Schizothecium for funga of Iran. Mycol Iran 2024;11:33-40. https://doi.org/10.22092/mi.2024.367836.1295
[DOI]
23. Dwibedi V, Saxena S. Arcopilus aureus, a resveratrol-producing endophyte from Vitis vinifera. Appl Biochem Biotechnol 2018;186:476-95. https://doi.org/10.1007/s12010-018-2755-x
[DOI]
24. Liang LQ, He XH, Wei CY, Zhao JX, Wu Q, Liang M, Li J, Luo Y, Xu WF, Yang RY. Antimicrobial polyketides from an endophytic fungus Arcopilus aureus HJ-7 from Ardisia mamillata Hance. Nat Prod Res 2024;19:1-8. https://doi.org/10.1080/14786419.2024.2429119
[DOI]
25. Charria-Girón E, Zeng H, Gorelik TE, Pahl A, Truong KN, Schrey H, Surup F, MarinFelix Y. Arcopilins: A new family of Staphylococcus aureus biofilm disruptors from the soil fungus Arcopilus navicularis. J Med Chem 2024;67:15029-40. https://doi.org/10.1021/acs.jmedchem.4c00585
[DOI]
26. Barrasa JM, Soláns MJ. Introducción al estudio del género Schizothecium Corda emend. Lundq. en España. Rev Iberoam Micol 1989;6:1-11.
27. Furuya K, Udagawa SI. Coprophilous pyrenomycetes from Japan I. J Gen Appl Microbiol 1972;18:433-54. https://doi.org/10.2323/jgam.18.433
[DOI]
28. Mirza JH, Cain RF. Revision of the genus Podospora. Can J Bot 1969;47:1999-2048. https://doi.org/10.1139/b69-293
[DOI]
29. Kim YJ, Lim SK, Nam SW, Back CG, Ten LN, Lee SY, Jung HY. First report of Varicosporellopsis shangrilaensis isolated from soil in Korea. Kor J Mycol 2025;53:21-8.
[DOI]
30. Kim TG, Nam SW, Lim SK, Back CG, Lee SY, Ten LN, Jung HY. First report of Magnusiomyces magnusii isolated in Korea. Kor J Mycol 2025;53:11-20.
[DOI]
31. Jung HY, Hong SM, Lim SK, Nam SW, Ten LN, Jung WK, Lee SY. Morphological and molecular characterization of unreported Fusarium mangiferae isolated from mango in Korea. Kor J Mycol 2024;52:223-32.
[DOI]
32. Azmi MH, Lim SK, Lim GJ, Choi JS, Choi JW, Kim MG, Lee SY, Jung HY. Morphological and phylogenetic analyses of an unreported species Diatrype rubi isolated from soil in Korea. Kor J Mycol 2024;52:391-8.
[DOI]
33. Kim TG, Ten LN, Hong SM, Lim SK, Lee SY, Jung HY. First report of Hamigera ingelheimensis isolated from Cheoltan Mountain in Korea. Kor J Mycol 2024;52:155-63.
[DOI]
34. Covino S, Stella T, D’Annibale A, Lladó S, Baldrian P, Čvančarová M, Cajthaml T, Petruccioli M. Comparative assessment of fungal augmentation treatments of a fine-textured and historically oil-contaminated soil. Sci Total Environ 2016;566-7:250-9. https://doi.org/10.1016/j.scitotenv.2016.05.018
[DOI]
35. Gałązka A, Grządziel J, Gałązka R, Gawryjołek K, Ukalska-Jaruga A, Smreczak B. Fungal community, metabolic diversity, and glomalin-related soil proteins (GRSP) content in soil contaminated with crude oil after long-term natural bioremediation. Front Microbiol 2020;11:572314. https://doi.org/10.3389/fmicb.2020.572314
[DOI]