Mi-Jeong Park, Aruna Jo, Kang-Hyeon Ka, and Yeongseon Jang*
Division of Forest Microbiology and Application, Department of Forest Bioresources, National Institute of Forest Science, Suwon 16631, Korea
*Correspondence to idjys@korea.kr
Korean Journal of Mycology (Kor J Mycol) 2025 December, Volume 53, Issue 4, pages 237-243.
https://doi.org/10.4489/kjm.2025.53.4.1
Received on October 17, 2025, Revised on November 10, 2025, Accepted on November 10, 2025, Published on December 31, 2025.
Copyright © The Korean Society of Mycology.
This is an Open Access article which is freely available under the Creative Commons Attribution-Non-Commercial 4.0 International License (CC BY-NC) (https://creativecommons.org/licenses/by-nc/4.0/).
As part of a study on the diversity of edible Cantharellus species in South Korea, molecular and morphological identifications were performed on specimens collected at a local market in Goesan-gun, Chungcheongbuk-do. Based on morphological characteristics and sequence information of the transcription elongation factor 1-alpha gene, the specimen was verified as Cantharellus hainanensis. This study presents the first record of C. hainanensis in South Korea and its distribution outside of China.
Chanterelle, Ectomycorrhiza, Macrofungi, Mushroom, Unreported species
The genus Cantharellus is among the most highly prized genera of wild, edible mushrooms worldwide. Because of their distinctive apricot-like aroma and excellent flavor, Cantharellus spp. have been used as popular culinary ingredients in various countries, and their commercial value has been estimated at billions of USD [1–3]. Recent pharmacological and biochemical studies have improved their market potential by elucidating their bioactive properties. For instance, extracts of C. cibarius have exhibited diverse functional activities, including antimicrobial [4], antioxidant [5,6], anti-aging [7], analgesic [8], and anticancer [9–11]. Similarly, extracts of C. applanatus have demonstrated anti-bacterial and anticancer activities [12]. A recent report added its ability to inhibit tyrosinase activity and suppress the expression of melanin synthesis-related proteins, highlighting its potential application in cosmeceutical formulations, particularly skin-whitening products [12].
In Korea, Cantharellus spp. are commonly called “Kwe-Koh-Ri (oriole)”, “Sahl-Goo (apricot)”, or “Wee-Got (cucumber flower)” mushrooms [13,14]. Several documents record that C. cibarius, C. cinnabarinus, C. ferruginascens, C. friesii, and C. tarbenensis are native to Korea, but the reliability of earlier taxonomic studies is limited by a reliance on morphological identification and weak genetic data. Since the late 2010s, collaborative efforts between the National Institute of Forest Science and Czech mycologists have resulted in the molecular verification of several additional taxa, namely C. albovenosus, C. applanatus (previously identified as C. anzutake), C. citrinus, C. curvatus, C. hongneungensis, C. koreanus, and C. subvaginatus [15–18]. Despite these advancements, species diversity within Korean Cantharellus remains under-documented when compared to neighboring China, where molecular analyses have confirmed 28 distinct species [19].
To address this gap, the present study evaluates the species diversity and improves the taxonomic resolution of wild Cantharellus. First, bulk chantarelles collected from a local market were subjected to morphological examination and molecular identification. We identified C. hainanensis, a species previously collected only from Hainan Island and Henan Province in China. We will continue to explore native Cantharellus taxa, study their conservation, and develop commercial cultivation methods.
Fresh wild chanterelles collected from Goesan-gun, Chungcheongbuk-do, were purchased in July 2024. The specimen was dried at 45℃ for 48 hours and deposited in the herbarium of the National Institute of Forest Science. Macroscopic characteristics of fresh basidiocarps were studied by examining dried materials mounted in 3% KOH using a TCS SPE confocal microscope (Leica Microsystems, Wetzlar, Germany) at 630x magnification.
Genomic DNA was isolated from fresh basidiocarps or fruiting body tissue using a DNeasy Plant Pro kit (QIAGEN, Hilden, Germany), following the manufacturer’s instructions. The nuclear ribosomal large subunit (nucLSU) and internal transcribed spacer (ITS) regions were amplified using the primer pairs LR0R/LR5 and ITS1F/ITS4, respectively [20–22]. Partial regions of transcription elongation factor 1-alpha (TEF-1α) and the second largest subunit of RNA polymerase II (RPB2) were amplified using the primer pairs tef-1Fcanth/tef-1Rcanth and RPB2-5FCanth/RPB2-7cRCanth, respectively [23]. The PCR products were sequenced by Macrogen Inc. (Seoul, Korea) using an ABI 3730xl DNA Analyzer (Applied Biosystems, CA, USA). The obtained sequences were deposited in GenBank (accession numbers: PV776357, PV777658, PV817755, and PV817756).
Phylogenetic relationships were inferred using the maximum likelihood method in MEGA 12 [24]. Multiple sequences were aligned by Multiple Sequence Comparison by Log-Expectation (MUSCLE) using the sequences of TEF-1α of 55 taxa of 31 Cantharellus species. The Tamura-Nei model was used for substitution. The robustness of the inferred phylogeny was assessed using bootstrap analysis with 1,000 replicates. The final tree was presented and annotated using MEGA 12.
The most likely tree inferred from the alignment of TEF1-α sequences of 55 taxa of Cantharellus species placed the specimen NIFoS20240725-01 within a monophyletic clade with the reference sequences of C. hainanensis, with substantial bootstrap support of 99% (Fig. 1).
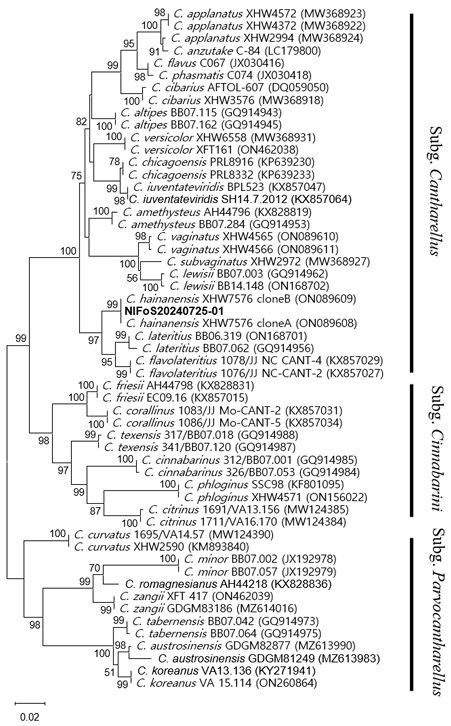
Fig. 1. The maximum-likelihood tree (-lnL = 3,672.08) constructed using transcription elongation factor 1-alpha gene sequences of Cantharellus species. Numbers on the nodes indicate bootstrap support (> 70%). The sequence obtained in this study is highlighted in bold.
Cantharellus hainanensis N. K. Zeng, Zhi Q. Liang & S Jiang, Mycoscience 58(6): 439 (2017) (Figs. 2 and 3)
Position in classification: Hydnaceae, Cantharellales, Agaricomycetes, and Basidiomycota.
Pileus 3.8-5.1 cm broad, yellow to yellowish-brown, infundibuliform; surface dry and dull; margin incurved and irregularly undulating; no change in color when injured. Hymenophore decurrent, variable development, usually smooth, weakly veined, sometimes with distinct veins, cream to yellowish-white. Stipe 3.8-4.5 × 1.0–1.6 cm, central, sub-cylindrical; yellowish white to yellowish brown. Smells weakly fruity. Spore print not obtained.
Basidiospores 5.3-6.8-7.8 × 4.3-5.2-6.4 μm, Q (length/width) = 1.11-1.33-1.65 (n = 53), smooth, oval, ellipsoid, thin-walled, hyaline in KOH. Basidia clavate to subcylindrical, (4-)5-spored, and hyaline in KOH.
Pileipellis 3.3-9.2 μm wide, cylindrical or subclavate, infrequently branched, hyaline to colorless in KOH.
Clamp connections present in all the tissues.
Edibility: Edible
Distribution: South Korea; China
Specimen examined: Cheongcheon-myeon, Goesan-gun, Chungcheongbuk-do, Republic of Korea; July 25, 2024 (NIFoS20240725-01).
Notes: Phylogenetically, C. hainanensis is related to C. applanatus and C. subvaginatus among the previously recorded Korean species. However, C. hainanensis is differentiated by its infundibuliform basidiomata and decurrent hymenophores with smooth veins.
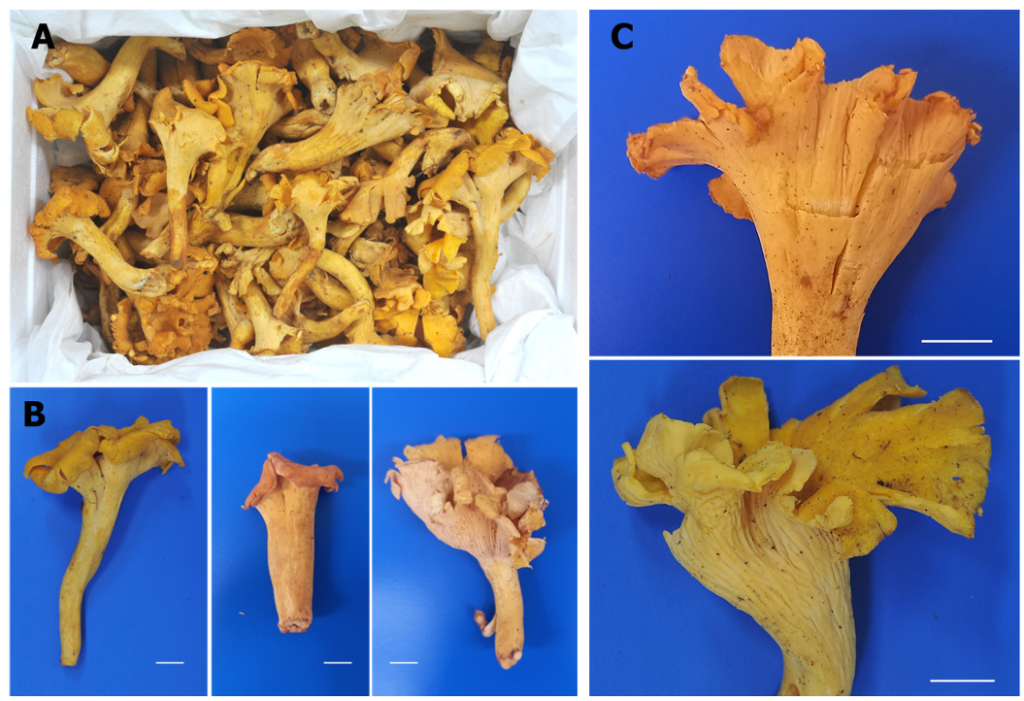

Fig. 2. Basidiomata of Cantharellus hainanensis (NIFoS20240725-01) collected at a local market in Goesan-gun, Chungcheongbuk-do. Bars = 1 cm.
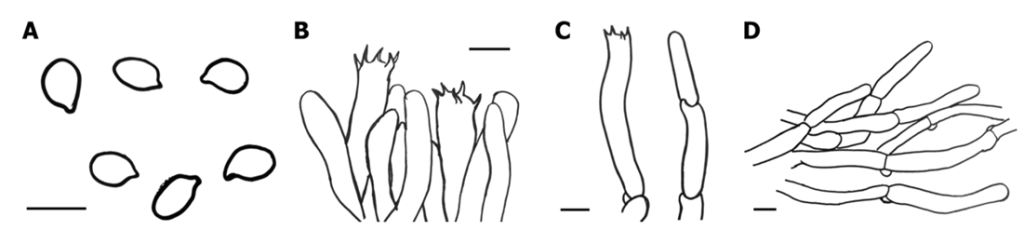
Fig. 3. Microscopic features of Cantharellus hainanensis (NIFoS20240725-01). A, Basidiospores; B and C, Basidia; D, Pileipellis. Bars = 10 µm.
Cantharellus spp. are widely consumed as food ingredients in Korea, and are distributed as fresh mushrooms in local and online markets. Based primarily on morphological characteristics, more than ten species, including C. cibarius, are presumed to be native to Korea [13,14]. Due to the inherent difficulty in distinguishing species within the genus, identification “in the pre-molecular era” relied predominantly on general attributes such as size and color [19]. Consequently, as in other Asian countries, the taxonomic identification of these fungi may have significant flaws. In support of this, molecular identification conducted in Korea during the 2010s by the National Institute of Forest Science and Czech mycologists revealed six novel species [15–18]; previously recorded species were not found, indicating the urgent need for accurate species identification.
The ITS region is commonly used for fungal identification owing to its high copy number, which facilitates amplification. However, in Cantharellus spp., the ITS region exceeds 1 kb and exhibits substantial heterogeneity (e.g., the C. applanatus strain KUN-HKAS109695 harbors a 131 bp deletion), complicating molecular identification. Recent comprehensive analyses of Asian Cantharellus spp. incorporating both morphological and genetic data has led to significantly advanced species identification [19]. To our knowledge, eight species, C. hainanensis, C. albovenosus, C. applanatus, C. citrinus, C. curvatus, C. hongneungensis, C. koreanus, and C. subvaginatus, have been molecularly verified in Korea [15–18]. This number is considerably lower than previous estimates based on morphological classification, and starkly contrasts with the 28 species reported in China, highlighting the necessity for further taxonomic studies on Korean Cantharellus species.
As ectomycorrhizal basidiomycetes, Cantharellus spp. relies on symbiotic associations with host plants for energy acquisition, which makes artificial cultivation more challenging than for saprotrophic fungi. However, Korean researchers have successfully cultivated the ectomycorrhizal basidiomycete Tricholoma matsutake [25–28], inducing mycorrhizal formation in the host plant Pinus densiflora in the forest. Subsequently, they facilitated the spread of mycorrhizae to neighboring hosts, leading to the production of T. matsutake fruiting bodies. This method was scientifically verified using molecular marker microsatellite or simple sequence repeat markers, demonstrating that fruiting bodies produced at the pine tree transplant site indeed originated from the same site where the T. matsutake-infected pine trees were made [28]. Moreover, continuous fruiting was observed for over eight years. Reports indicate that artificial fruiting of certain Cantharellus spp. has also been achieved under laboratory conditions [29,30]. Using the recently obtained pure isolates of C. hainanensis and C. subvaginatus, we will further develop cultivation methods for Cantharellus spp.
The authors declare no potential conflict of interest for this study.
This study was supported by a grant from the General Project (FP0800-2023-01) of the National Institute of Forest Science, Republic of Korea.
1. Watling R. The business of fructification. Nature 1997;385:299-300. https://doi.org/10.1038/385299a0
[DOI]
2. Pilz D, Norvell L, Danell E, Molina R. Ecology and management of commercially harvested chanterelle mushrooms. Portland: US Department of Agriculture, Forest Service, Pacific Northwest Research Station; 2003. https://doi.org/10.2737/PNW-GTR-576
[DOI]
3. Kumar A, Devi R, Dhalaria R, Tapwal A, Verma R, Rashid S, Elossaily GM, Khan KA, Chen KT, Verma T. Nutritional, nutraceutical, and medicinal potential of Cantharellus cibarius Fr.: A comprehensive review. Food Sci Nutr 2024;13:e4641. https://doi.org/10.1002/fsn3.4641
[DOI]
4. Kolundžić M, Stanojković T, Radović J, Tačić A, Dodevska M, Milenković M, Sisto F, Masia C, Farronato G, Nikolić V, Kundaković T. Cytotoxic and antimicrobial activities of Cantharellus cibarius Fr. (Cantarellaceae). J Med Food 2017;20:790-6. https://doi.org/10.1089/jmf.2016.0176
[DOI]
5. Zhao D, Ding X, Hou Y, Hou W, Liu L, Xu T, Yang D. Structural characterization, immune regulation and antioxidant activity of a new heteropolysaccharide from Cantharellus cibarius Fr. Int J Mol Med 2018;41:2744-54. https://doi.org/10.3892/ijmm.2018.3450
[DOI]
6. Kozarski M, Klaus A, Vunduk J, Zizak Z, Niksic M, Jakovljevic D, Vrvic MM, Van Griensven LJ. Nutraceutical properties of the methanolic extract of edible mushroom Cantharellus cibarius (Fries): Primary mechanisms. Food Funct 2015;6:1875-86. https://doi.org/10.1039/C5FO00312A
[DOI]
7. Luo J, Ganesan K, Xu B. Unlocking the power: New insights into the anti-aging properties of mushrooms. J Fungi 2024;10:215. https://doi.org/10.3390/jof10030215
[DOI]
8. Khalili M, Ebrahimzadeh MA, Omrani F, Karami M. Antihypoxic activities of the golden Chanterelle Mushroom, Cantharellus cibarius (higher Basidiomycetes). Int J Med Mushrooms 2014;16:339-44. https://doi.org/10.1615/IntJMedMushrooms.v16.i4.40
[DOI]
9. Lemieszek MK, Nunes FM, Rzeski W. Branched mannans from the mushroom Cantharellus cibarius enhance the anticancer activity of natural killer cells against human cancers of lung and colon. Food Funct 2019;10:5816-26. https://doi.org/10.1039/C9FO00510B
[DOI]
10. Chen X, Xu B. Insights into chemical components, health-promoting effects, and processing impact of golden chanterelle mushroom Cantharellus cibarius. Food Funct 2024;15:7696-732. https://doi.org/10.1039/D4FO00891J
[DOI]
11. Lemieszek MK, Marques PS, Ribeiro M, Ferreira D, Marques G, Chaves R, Pożarowski P, Nunes FM, Rzeski W. Mushroom small RNAs as potential anticancer agents: A closer look at Cantharellus cibarius proapoptotic and antiproliferative effects in colon cancer cells. Food Funct 2019;10:2739-51. https://doi.org/10.1039/C8FO02378F
[DOI]
12. Lee YJ, Kim M, Hong H, Yeon JE, Kim C, Chi WJ, Kim SY. Whitening effect of Cantharellus applanatus extract on B16F10 melanoma cells. J Appl Biol Chem 2024;67:141-6. https://doi.org/10.3839/jabc.2024.019
[DOI]
13. Seok SJ, Lim YW, Kim CM, Ka KH, Lee JS, Han SK, Kim SO, Hur JS, Hyun IH, Hong SG, et al. List of mushrooms in Korea. Seoul: The Korean Society of Mycology; 2013.
14. Korea National Arboretum. National list of standard mushrooms [Internet]. Pocheon (Gyeonggi-do): Korea National Arboretum; n.d. [cited 2025 Jun 25]. Available from: http://www.nature.go.kr/kfni
15. Antonín V, Hofstetter V, Ryoo R, Ka KH, Buyck B. New Cantharellus species from the Republic of Korea. Mycol Progress 2017;16:753-9. https://doi.org/10.1007/s11557-017-1312-2
[DOI]
16. Buyck B, Antonín V, Chakraborty D, Baghela A, Das K, Hofstetter V. Cantharellus sect. Amethystini in Asia. Mycol Progress 2018;17:917-24. https://doi.org/10.1007/s11557-018-1403-8
[DOI]
17. Buyck B, Hofstetter V, Ryoo R, Ka KH, Antonín V. New Cantharellus species from South Korea. MycoKeys 2020;76:31-47. https://doi.org/10.3897/mycokeys.76.58179
[DOI]
18. Hyde KD, Suwannarach N, Jayawardena RS, Manawasinghe IS, Liao CF, Doilom M, Cai L, Zhao P, Buyck B, Phukhamsakda C, et al. Mycosphere notes 325-344 – Novel species and records of fungal taxa from around the world. Mycosphere 2021;12:1101-56. https://doi.org/10.5943/mycosphere/12/1/14
[DOI]
19. Wang XH, Hofstetter V, Cao SQ, Liu PG, Buyck B. Finding correct names for economically important chanterelles (Cantharellus, Hydnaceae, Cantharellales) in southwestern China: A plea for third party annotation of sequences in GenBank. Mycosphere 2023;14:153-94. https://doi.org/10.5943/mycosphere/14/1/3
[DOI]
20. Vilgalys R, Hester M. Rapid genetic identification and mapping of enzymatically amplified ribosomal DNA from several Cryptococcus species. J Bacteriol 1990;172:4238-46. https://doi.org/10.1128/jb.172.8.4238-4246.1990
[DOI]
21. White TJ, Bruns TD, Lee SB, Taylor JW. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ, editors. PCR protocols: A guide to methods and applications. New York: Academic Press; 1990. pp. 315-22.
[DOI]
22. Gardes M, Bruns TD. ITS primers with enhanced specificity for basidiomycetes-application to the identification of mycorrhizae and rusts. Mol Ecol 1993;2:113-8. https://doi.org/10.1111/j.1365-294X.1993.tb00005.x
[DOI]
23. Buyck B, Kauff F, Eyssartier G, Couloux A, Hofstetter V. A multilocus phylogeny for worldwide Cantharellus (Cantharellales, Agaricomycetidae). Fungal Divers 2014;64:101-21. https://doi.org/10.1007/s13225-013-0272-3
[DOI]
24. Kumar S, Stecher G, Suleski M, Sanderford M, Sharma S, Tamura K. MEGA12: Molecular evolutionary genetic analysis version 12 for adaptive and green computing. Mol Biol Evol 2024;41:msae263. https://doi.org/10.1093/molbev/msae263
[DOI]
25. Ka KH, Hur TC, Park H, Kim HS, Bak WC, Yoon KH. Production and transplanting of ectomycorrhizal pine seedlings using the old fairy ring of Tricholoma matsutake. J Kor Soc For Sci 2006;95:636-42.
26. Ka KH, Hur TC, Park H, Kim HS, Bak WC. Mycelial growth and fairy-ring formation of Tricholoma matsutake from matsutake-infected pine trees. Kor J Mycol 2010;38:16-20.
[DOI]
27. Ka KH, Kim HS, Jeon SM, Ryoo R, Jang Y, Wang EJ, Jeong YS. Determination of the minimum size of seedlings with matsutake mycelia that can survive in the field for matsutake-infected pine tree production. Kor J Mycol 2017;45:188-95. https://doi.org/10.4489/KJM.20170023
[DOI]
28. Ka KH, Kim HS, Lee JW, Jang Y, Ryoo R. Confirmation of fruiting bodies of Tricholoma matsutake occurring from matsutake-infected pine trees using microsatellite markers. Kor J Mycol 2021;49:469-76.
[DOI]
29. Danell E, Camacho F. Successful cultivation of the golden chanterelle. Nature 1997;385:303. https://doi.org/10.1038/385303a0
[DOI]
30. Ogawa W, Takeda Y, Endo N, Yamashita S, Takayama T, Fukuda M, Yamada A. Repeated fruiting of Japanese golden chanterelle in pot culture with host seedlings. Mycorrhiza 2019;29:519-30. https://doi.org/10.1007/s00572-019-00908-z
[DOI]