Young-Ju Nam1,2, Seung-Yeol Lee2, Weon-Dae Cho1, and Wan-Gyu Kim1*
1Global Agro-Consulting Corporation, Suwon 16614, Korea
2Department of Plant Medicine, Kyungpook National University, Daegu 41566, Korea
*Correspondence to wgkim5121@naver.com
Korean Journal of Mycology (Kor J Mycol) 2025 December, Volume 53, Issue 4, pages 237-243.
https://doi.org/10.4489/kjm.2025.53.4.1
Received on October 17, 2025, Revised on November 10, 2025, Accepted on November 10, 2025, Published on December 31, 2025.
Copyright © The Korean Society of Mycology.
This is an Open Access article which is freely available under the Creative Commons Attribution-Non-Commercial 4.0 International License (CC BY-NC) (https://creativecommons.org/licenses/by-nc/4.0/).
In August 2021, stem rot symptoms sporadically occurred in potted plants of Euphorbia hypogaea growing in a greenhouse in Pyeongtaek, Korea. The leaves of diseased plants wilted, and the epidermal layer of the stems blackened from the soil line portion and rotted. The disease incidence in plants in the greenhouse ranged from 1 to 6%. We obtained 12 oomycete isolates from the stems of diseased plants. All isolates were identified as Phytophthora nicotianae based on an analysis of their morphological and phylogenetic characteristics (NADH dehydrogenase subunit 1 and beta-tubulin genes). Three isolates of P. nicotianae were tested for pathogenicity in E. hypogaea plants using artificial inoculation. The isolates induced stem rot symptoms in the inoculated plants, similar to those observed in the greenhouse where the disease first occurred. This is the first report that P. nicotianae causes Phytophthora stem rot in E. hypogaea.
Euphorbia hypogaea, Phytophthora nicotianae, Phytophthora stem rot
Euphorbia hypogaea Marloth, which belongs to the family Euphorbiaceae, is native to Central Cape Province, South Africa, and grows primarily in desert or dry shrubland biomes [1]. This plant was introduced to Korea many years ago, where it is cultivated in greenhouses as an ornamental succulent.
In August 2021, stem rot symptoms sporadically occurred in potted plants of E. hypogaea growing in a greenhouse in Pyeongtaek, Korea. The leaves of diseased plants wilted, and the epidermal layer of the stems blackened from the soil line portion and rotted (Fig. 1A and 1B). Three cultivation pedestals were observed in the greenhouse and 200 plants grown in pots for each cultivation pedestal were investigated for disease incidence. The disease incidence in plants in the greenhouse ranged from 1 to 6%.
Fungi were isolated from the diseased stems of E. hypogaea. The 3—5 mm long samples of lesions cut from the stems were immersed in 1% sodium hypochlorite for 1 min and then plated on 2% water agar (WA; FUJIFILM Wako Pure Chemical Corporation, Osaka, Japan). Single hyphal tips growing from the lesion samples were transferred to potato dextrose agar (PDA; BD Difco, Sparks, MD, USA) slants after incubating the plates at 25℃ for 2—3 days. Twelve oomycete isolates were obtained from the stems of diseased plants. Each isolate was cultured on PDA in 9-cm-diameter Petri dishes at 25°C in the dark for 10 days to investigate the cultural features. The colonies of the isolates displayed white mycelia with coralloid edges (Fig. 2A and 2B).
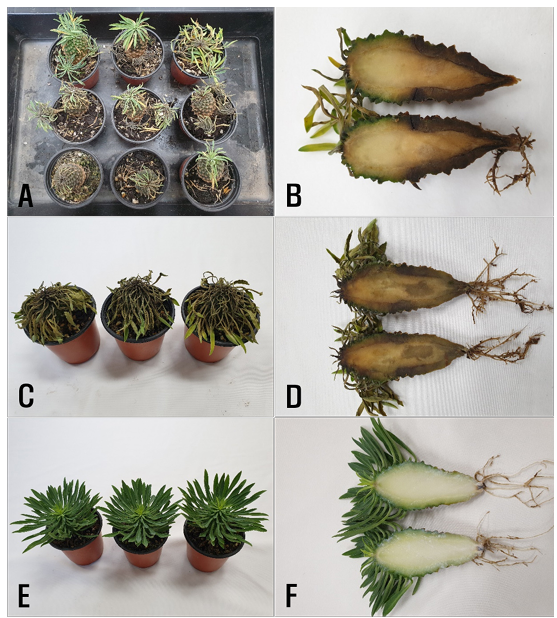

Fig. 1. Stem rot symptoms in Euphorbia hypogaea plants. A and B: Symptoms observed in the greenhouse. C and D: Symptoms induced by artificial inoculation tests with Phytophthora nicotianae isolates. E and F: Non-inoculated plants.
Each isolate was cultured on V8-juice agar (V8A) in 9-cm-diameter Petri dishes for 7 days. Twenty milliliters of sterile distilled water was then added to the cultures and they were incubated at 24–26°C under f luorescent light for 6 days to produce sporangia. The morphological characteristics of 10–20 sporangia per isolate were investigated under a light microscope (Eclipse Ci-L, Nikon, Tokyo, Japan). Hyphae were nonseptate, and sporangia were globose or ovoid, papillate, persistent, and caducous with short pedicels (Fig. 2C—2E). Sporangia measured 37.6—58.1 × 28.9—36.2 μm (average 44.5 × 33.3 μm), and the papillae of sporangia measured 3.0—6.0 × 5.3—8.5 μm (average 4.0 × 7.1 μm). The morphological characteristics of the isolates were similar to those of Phytophthora nicotianae Breda de Haan described in a previous study [2].
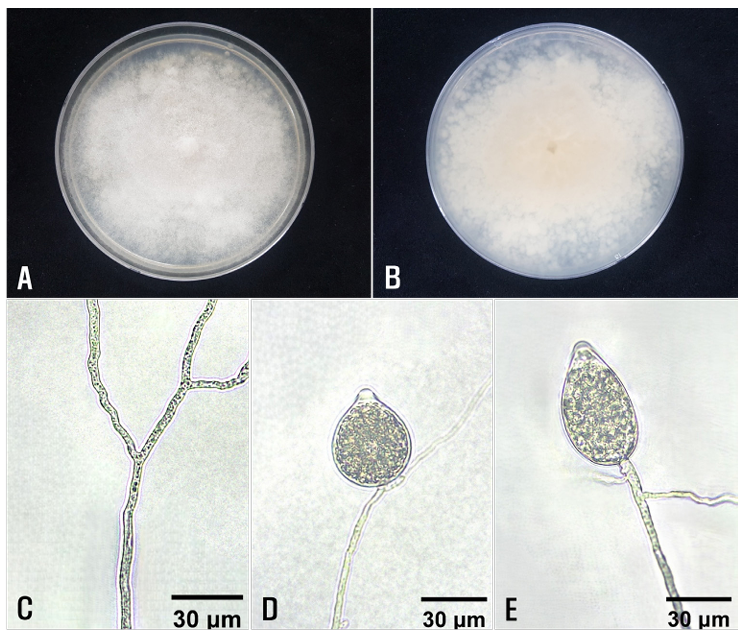
Fig. 2. Cultural and morphological features of Phytophthora sp. isolates from diseased Euphorbia hypogaea plants. Colony morphology on the surface (A) and reverse side (B) of potato dextrose agar. C: A non-septate hypha. D and E: Sporangiophores and sporangia.
Phylogenetic analysis was conducted to confirm the morphological identification of P. nicotianae isolates (EHPHY2101–EHPHY2112). Total genomic DNA was extracted from the isolates grown on PDA using a HiGene™ Genomic DNA Prep Kit (Biofact Co., Ltd., Daejeon, Korea) according to the manufacturer’s protocol. For molecular identification, the NADH dehydrogenase subunit 1 (NADH1) and beta-tubulin (TUB2) genes of the isolates were amplified using the primer pairs NADH1F/NADH1R and TubuF2/ TubuR1, respectively [3]. Amplified PCR products were purified using ExoSAP-IT (Thermo Fisher Scientific Inc., Waltham, MA, USA) and sequenced by Solgent Co., Ltd. (Daejeon, Korea). All obtained sequences were registered with the National Center for Biotechnology Information (NCBI) under accession numbers OR161944–OR161955 and OR161956–OR161967 for NADH1 and TUB2, respectively.
Phylogenetic analysis was performed using retrieved sequences registered in the NCBI database (Table 1). Partial sequences of NADH1 and TUB2 were analyzed for species identification, supported by the construction of phylogenetic trees using the neighbor-joining method, as described by the Kimura model [4,5]. The analysis was conducted using MEGA 11.0, with bootstrap values calculated from 1,000 replicates [6].
Table 1. List of species used in phylogenetic analyses along with their GenBank accession numbers
테일블
NADH1: NADH dehydrogenase subunit 1, TUB2: beta-tubulin.
*The present isolates: EHPHY2101–EHPHY2112.
The NADH1 (749 bp) and TUB2 (857 bp) gene sequences were obtained from all 12 isolates, and the sequences of both genes were identical across all isolates. BLAST results of the partial NADH1 gene sequences revealed high levels of similarity of 99.9% and 100% with Phytophthora nicotianae AR 238 and P582, respectively, compared to 97.5% similarity with Phytophthora clandestina IMI287317 and 97.3% with Phytophthora iranica CBS 374.72. High levels of similarity of 99.9% and 100% were observed with TUB2 gene sequences from P. nicotianae P582 and AR 238, respectively, compared with similarities of 95.1% with P. iranica CBS 374.72 and 94.4% with P. clandestina IMI287317. Phylogenetic trees were constructed based on the NADH1 and TUB2 sequences using the neighbor-joining method (Fig. 3). The 12 isolates clustered with P. nicotianae AR 238 and P582. Accordingly, the isolates were identified as P. nicotianae based on their morphological characteristics and phylogenetic analysis.
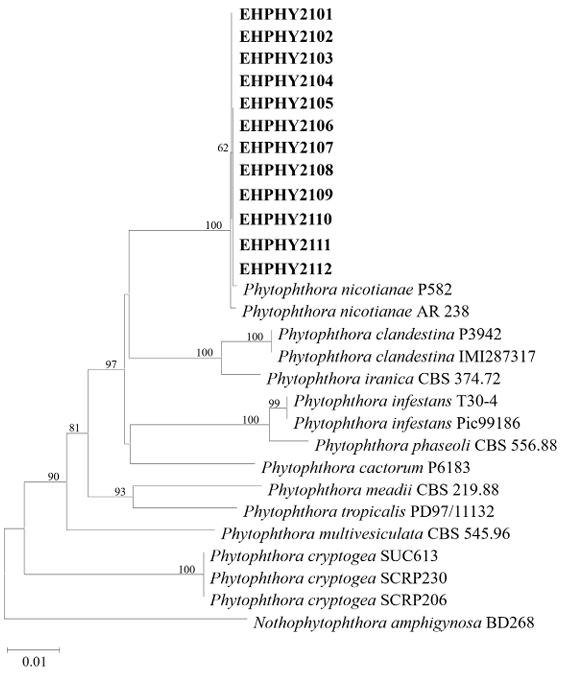

Fig. 3. Neighbor-joining phylogenetic tree based on a combined dataset of partial sequences of NADH dehydrogenase subunit 1 and beta-tubulin genes showing the phylogenetic position of the isolates (EHPHY2101‒EHPHY2112) from Euphorbia hypogaea among Phytophthora species. Bootstrap values with more than 60% (percentage of 1,000 replicates) are shown at branching points. The tree was rooted using Nothophytophthora amphigynosa BD268 as an outgroup. The bar at 0.01 represents substitutions per nucleotide position.
No conflict of interest was reported by the authors.
This study was supported by a research grant (PJ01450701) from the Rural Development Administration, Korea.
1. Plants of the World Online. Euphorbia hypogaea Marloth [Internet]. London: Royal Botanic Gardens, Kew; 2025 [cited 2025 August 22]. Available from https://powo.science.kew.org/.
2. Abad ZG, Burgess TI, Bourret T, Bensch K, Cacciola SO, Scanu B, Mathew R, Kasiborski B, Srivastava S, Kageyama K, et al. Phytophthora: taxonomic and phylogenetic revision of the genus. Stud Mycol 2023;106:259-348. https://doi.org/10.3114/sim.2023.106.05
[DOI]
3. Kroon LPNM, Bakker FT, van den Bosch GBM, Bonants PJM, Flier WG. Phylogenetic analysis of Phytophthora species based on mitochondrial and nuclear DNA sequences. Fungal Genet Biol 2004;41:766-82. https://doi.org/10.1016/j.fgb.2004.03.007
[DOI]
4. Saitou N, Nei M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 1987;4:406-25. https://doi.org/10.1093/oxfordjournals.molbev.a040454
[DOI]
5. Kimura M. A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 1980;16:111-20. https://doi.org/10.1007/BF01731581
[DOI]
6. Tamura K, Stecher G, Kumar S. MEGA11: Molecular evolutionary genetics analysis version 11. Mol Biol Evol 2021;38:3022-7. https://doi.org/10.1093/molbev/msab120
[DOI]
7. Panabieres F, Ali GS, Allagui BM, Dalio RJD, Gudmestad NC, Kuhn ML, Guha Roy S, Schena L, Zampounis A. Phytophthora nicotianae diseases worldwide: New knowledge of a long-recognised pathogen. Phytopathol Mediterr 2016;55:20-40. https://doi.org/10.14601/Phytopathol_Mediterr-16423
[DOI]
8. Korean Society of Plant Pathology. List of Plant Diseases in Korea [Internet]. Seoul: Korean Society of Plant Pathology; 2025 [cited 2025 August 22]. Available from http://genebank.rda.go.kr/kplantdisease.do.
9. Yakubu I, Song JH, Lee YJ, Son MA, Han SH, Kong HG. First report of soft rot induced by Dickeya dadantii on Euphorbia hypogaea in Korea. Res Plant Dis 2024;30:95-8. https://doi.org/10.5423/RPD.2024.30.1.95
[DOI]