Yein Kim1, Hyeong Jin Noh2, Doh Hun Shin1, and Seong Hwan Kim1,*
1Department of Microbiology, Dankook University, Cheonan 31116, Korea
2Forest Entomology and Pathology Division, National Institute of Forest Science, Seoul 02455, Korea
*Correspondence to piceae@dankook.ac.kr
Korean Journal of Mycology (Kor J Mycol) 2025 December, Volume 53, Issue 4, pages 237-243.
https://doi.org/10.4489/kjm.2025.53.4.1
Received on October 17, 2025, Revised on November 10, 2025, Accepted on November 10, 2025, Published on December 31, 2025.
Copyright © The Korean Society of Mycology.
This is an Open Access article which is freely available under the Creative Commons Attribution-Non-Commercial 4.0 International License (CC BY-NC) (https://creativecommons.org/licenses/by-nc/4.0/).
Jeju Island hosts a unique ecosystem that serves as a biodiversity hotspot for numerous species and rare organisms. During an investigation of fungal diversity in the hyphosphere soil of Pseudotulostoma japonicum in Tamna Valley and Armillaria mellea in the Gyorae Gotjawal Forest, we identified three species belonging to Ascomycota fungi that had not been previously recorded in Korea. To perform taxonomic study on these species, we observed morphology and analyzed the nucleotide sequences of the internal transcribed spacer regions, the large subunit of rDNA, the β-tubulin gene, and the translation elongation factor 1-α gene. Consequently, Mariannaea humicola, Nigrograna hydei, and Trichoderma pubescens were listed as newly recorded species in Korea.
Hyphosphere soil, Jeju Island, Mariannaea humicola, Nigrograna hydei, Trichoderma pubescens
Jeju Island in Korea, with its volcanic origin and unique temperate and subtropical climate, supports rich biodiversity, particularly within the rhizosphere where microorganisms thrive and engage in complex interactions [1]. This biodiversity is enhanced by high rainfall and diverse ecosystems, such as the Tamna Valley and Gyorae Gotjawal Forest, both of which are known for their fungal diversity [2,3]. These areas serve as prime locations for investigating fungal species associated with the rhizosphere. Gotjawal forests have unique volcanic soils that host diverse microbial communities shaped by environmental factors such as soil pH, organic matter content, and mineral composition [3,4]. Research in these areas has highlighted the distinctness of microbial communities in lava-formed soils and their ecological roles [3,5]. Furthermore, studies on endophytic fungi on Jeju Island have emphasized their role in nutrient cycling, environmental resilience, and their potential applications in agriculture and biotechnology [2,5]. This indicates the broad ecological significance of fungi on ecosystems of Jeju Island and underscores the importance of preserving these unique environments.
Mushrooms on Jeju Island have been surveyed in biodiversity study [1]. However, the hyphosphere soils of wild mushrooms have not been extensively studied. Recently, we identified 11 unrecorded fungal species from the hyphosphere soil of wild mushrooms in the Gotjawal Forest by exploring the biodiversity and species distribution of this ecologically and economically important taxon [6]. We further investigated the fungi in the hyphosphere soil of wild mushrooms grown in the Gyorae Gotjawal Forest and Tamna Valley. Four additional species of Ascomycota fungi that have not been previously reported in Korea were identified. In this study, we report the molecular and morphological identification and characterization of these species.
Sampling of mushroom hyphosphere soil on Jeju Island was performed for Pseudotulostoma japonicum in the Tamna Valley at 33.400035°N, 126.540828°E and for Armillaria mellea in Gyorae Gotjawal Forest at 33.28027°N, 126.26333°E on Jeju Island. Soils 10 cm deep beneath these two mushroom fruiting bodies were collected using a long seedling shovel, then placed in a zipper bag, and transported to the laboratory for analysis. Two grams of the collected wild mushroom hyphosphere soil sample and 20 mL of sterilized water were placed in a 50 mL plastic tube. The plastic tube was vortexed for 10 min to separate fungal spores or mycelia from the soil and allowed to float in sterilized water. The suspended solution was diluted to 10-3 using a step dilution method, and 100 μL of each diluted solution was spread on dichloran glycerol 18% (DG18; Becton, Dickinson and Company [BD], Franklin Lakes, NJ, USA) medium supplemented with the antibiotic chloramphenicol (50 µg/mL) using a smear plate method. After culturing the inoculated medium in a 25°C incubator for 7 d, the growing fungal colonies were confirmed by naked eye. Fungal colonies that appeared to have different morphological features were purified on potato dextrose agar (PDA; BD) and cultured in an incubator at 25°C for 7 d. Single-spore isolation was performed with sporulating isolates, and pure cultures were obtained and assigned Dankook University Culture Collection (DUCC) numbers. Representative strains among the assigned DUCC numbers were deposited at the National Institute of Biological Resources (NIBR), Incheon, Korea, and assigned the corresponding NIBRFGC accession numbers.
For detailed morphological analyses, the fungal plates were incubated at 25°C in the dark for 7–14 d on PDA, malt extract agar (MEA; MBcell, KisanBio, Seoul, Korea), Czapek yeast extract agar (CYA; MBcell, KisanBio), oatmeal agar (OA; BD), synthetic nutrient poor agar (SNA; MBcell, KisanBio), and corn meal agar (CMA; Difco, Detroit, MI, USA). Using colonies grown on PDA, microscopic features were observed under an optical microscope (BX53; Olympus, Tokyo, Japan) at 400× magnification. The lengths and thicknesses of the microstructures were measured 30 times each.
Genomic DNA was extracted directly from the mycelia of fungal isolates using a NaviBiotech Direct DNA Prep Kit (NaviBiotech Corp., Cheonan, Korea). Using the extracted DNA as a template, the sequences of the internal transcribed spacer (ITS) regions, the large subunit (LSU) of rDNA, the β-tubulin gene (TUB2), and the translation elongation factor 1-α gene (tef1-α) were amplified by PCR using the primers and reaction conditions given in Table 1. The amplified PCR products were verified using electrophoresis on a 1% (w/v) agarose gel. The PCR products were purified using a silica gel column and 80% ethanol solution. DNA sequencing was performed by Bionics Corp. (Seoul, Korea). The determined nucleotide sequences were edited using Chromas 2.6 program (https://chromas.software.informer.com/), and the edited sequences were searched for similar sequences using the Basic Local Alignment Search Tool (BLASTN) of the National Center for Biotechnology Information (https://www.ncbi.nlm.nih.gov/). The reference fungal nucleotide sequences of the ITS, LSU rDNA, TUB2, and tef1-α were downloaded from the GenBank DNA database of NCBI and given in Tables 2–4. For phylogenetic analysis, the sequences of the DUCC strains were aligned with the reference sequences using the CLUSTALW alignment tool in the MEGA 11.0 software program [7]. Maximum likelihood (ML) trees were constructed with aligned sequences using the Kimura 2-parameter model, and the reliability of phylogenetic tree branches was analyzed using 1,000 bootstrap replicates [8,9].
Table 1. Conditions of each primer set used for PCR
테일블
ITS: internal transcribed spacer, LSU rDNA: 28S ribosomal RNA gene; TUB2: β-tubulin gene; tef1-α: translation elongation factor 1-α gene.
Table 2. Sequence information on the related species used for the phylogenetic analysis of Nigrograna hydei NIBRFGC000510007
테일블
ITS: internal transcribed spacer, LSU rDNA: 28S ribosomal RNA gene.
Ttype strain. Strain identified in this study and its GenBank accession numbers are indicated in bold.
Table 3. Sequence information on the related species used for the phylogenetic analysis of Mariannaea humicola NIBRFGC000510020
테일블
ITS: internal transcribed spacer, LSU rDNA: 28S ribosomal RNA gene, TUB2: β-tubulin gene.
Ttype strain. Strain identified in this study and its GenBank accession numbers are indicated in bold.
Table 4. Sequence information on the related species used for the phylogenetic analysis of Trichoderma pubescens NIBRFGC000510021
테일블
ITS: internal transcribed spacer, tef1-α: translation elongation factor 1-α gene.
Ttype strain. Strain identified in this study and its GenBank accession numbers are indicated in bold.
Strains examined. Tamna Valley on Jeju Island, isolated from the mushroom hyphosphere soil of Pseudotulostoma japonicum in 2022, strain N. hydei DUCC 15293 (NIBRFGC000510007).
Phylogenetic analysis (Fig. 1)
The determined ITS sequence of strain NIBRFGC000510007 was 536 bp in size and showed 99.81% sequence similarity with that of the type strain N. hydei MFLU 18-2073 (GenBank accession OP741100). The LSU rDNA sequence of this strain was 908 bp and had a sequence similarity of 99.88% with that of the type strain (GenBank accession OP741109). The results of constructing an ML phylogenetic tree based on the 1,268 bp concatenated ITS and LSU rDNA sequences, including gaps, showed that the strain NIBRFGC000510007 formed an independent clade with N. hydei MFLU 18-2073 and N. hydei ELS12022.
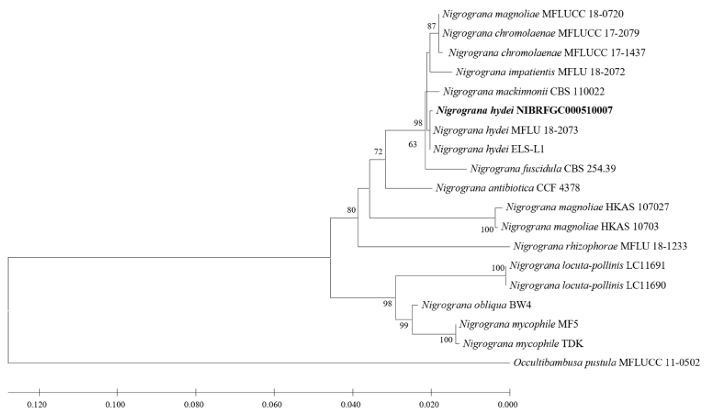
Fig. 1. Maximum likelihood (ML) phylogram constructed based on the concatenated ITS-LSU rDNA sequences. The number of nodes represents the reliability value (>60%) through 1,000 bootstrap replicates. Fungal strain isolated in this study is indicated in bold. Occultibambusa pustula MFLUCC 11-0502 is used as an outgroup. The reference sequences used to construct the ML phylogenetic tree are listed in Table 2.
Morphological characteristics (Fig. 2)
Description. On PDA, 15–16 mm diameter in 14 d at 25°C, aerial mycelia dense, floccose, raised at the center, circular, white at the margin, forming several concentric rings on the surface, white, reverse white at the margin, dark olive towards inside. On MEA, 22–23 mm diameter in 14 d at 25°C, aerial mycelia dense, floccose, raised at the center, circular, radial, white at the margin, forming several concentric rings on the surface, white, reverse light pinkish at the margin, dark brown towards inside. On CYA, 22–23 mm diameter in 14 d at 25°C, aerial mycelia dense, floccose, raised at the center, circular, white at the margin, forming several concentric rings on the surface, white, reverse white at the margin, dark olive towards inside. On OA, 32–33 mm diameter in 14 d at 25°C, aerial mycelia dense, floccose, raised at the center, circular, white at the margin, forming several concentric rings on the surface. Hyphae were hyaline, branched, septate, thin-walled, width 2–3 μm (mean ± SD: 2.5 ± 0.3 μm, n = 30), sterile. No asexual or sexual reproductive structures are observed.
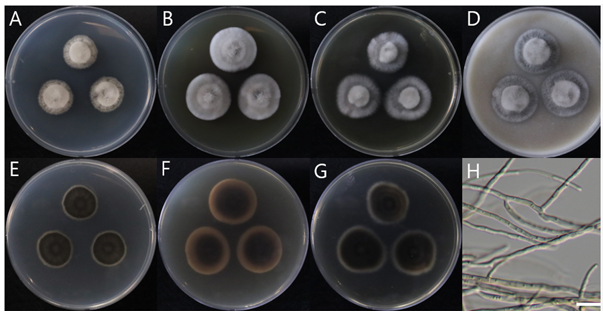
Fig. 2. Colony morphology and light microscopic image of Nigrograna hydei NIBRFGC000510007 grown on different media at 25°C for 14 d. A, E: PDA. B, F: MEA. C, G: CYA. D: OA. H: sterile mycelia. Obverse: A–D. Reverse: E–G. Scale bar = 10 μm.
Notes
The strain NIBRFGC000510007 did not exhibit sporulation on any of the test media (PDA and MEA).
It appears that host substrates, such as woody plants, are required for the sexual stage. The morphological characteristics of the type of strain N. hydei MFLU 18-2073 from Thailand were described using ascomata and ascospores produced in woody branches of an unknown species [14]. Its germinated ascospores formed colonies on PDA after 35 d at 25°C, reaching up to 33 mm. Its colony morphology includes white at the margin, forming several concentric rings on the surface, white, reverse white at the margin, circular, dense aerial mycelia, grey from above, brown to dark pigmented in reverse, similar to that of NIBRFGC000510007. However, no asexual morphs were observed in this strain. Attempts were made to observe spores on PDA, MEA, CYA, and OA, but no reproductive structures, including conidia or chlamydospores, were observed on any of these media. Interestingly, similar to the strain NIBRFGC000510007, the strain N. hydei ELS1-2022 isolated from healthy leaves of Elaeagnus angustifolia in India did not sporulate on growth media such as PDA and MEA [15]. The anamorphic characteristics of colony morphology described for strain N. hydei ELS1-2022 were also similar to those of strain NIBRFGC000510007 regarding aerial mycelia, floccose, dome-shaped (circular), slightly heaped at the center (raised at the center), and periphery turning white (white at the margin and forming several concentric rings on the surface). Sterile mycelia were also common features of strains NIBRFGC000510007 and N. hydei ELS1-2022. N. hydei ELS1-2022, intercalary chlamydospores in the mycelia were not observed in strain NIBRFGC000510007. In conclusion, based on the phylogenetic and morphological analysis results, we identified strain NIBRFGC000510007 as N. hydei.
Strains examined. Tamna Valley on Jeju Island, isolated from the mushroom hyphosphere soil of Pseudotulostoma japonicum in 2022, strain M. humicola DUCC 21466 (NIBRFGC000510020).
Phylogenetic analysis (Fig. 3)
A ML phylogenetic tree was constructed based on concatenated ITS, LSU rDNA, and TUB2 sequences. The size of the ITS region of strain NIBRFGC000510020 was 484 bp, displaying 100% sequence similarity with that of the type strain M. humicola CBS 740.95 (MW340821). The size of its LSU rDNA was 865 bp, showing 99.88% sequence similarity with that of the type strain (PP380805). The size of TUB2 was 318 bp, showing 100% sequence similarity to that of the type strain (KM232013). As a result of constructing an ML phylogenetic tree based on the 1,850 bp concatenated sequence including gaps, the strain NIBRFGC000510020 was clustered with M. humicola CBS 740.95, and CBS 102628 with sufficient node value and separated from M. aquaticola.
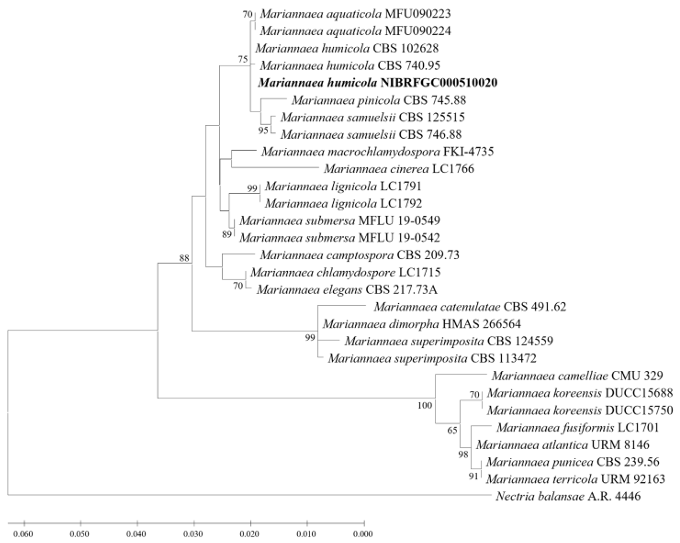
Fig. 3. Maximum likelihood (ML) phylogram constructed based on the concatenated ITS-LSU-TUB2 sequences. The number of nodes represents the reliability value (>60%) through 1,000 bootstrap replicates. Fungal strain isolated in this study is indicated in bold. Nectria balansae A.R. 4446 is used as an outgroup. The reference sequences used to construct the ML phylogenetic tree are listed in Table 3.
Morphological characteristics (Fig. 4)
Description. On PDA, 26–27 mm diameter in 7 d at 25°C, surface dirty white in the center becoming tan to light yellow brown towards the margins with dirty white, irregularly distributed tuffs of fascicles, aerial mycelium abundant, forming several concentric rings on the surface, which are reverse dark brown in color. On MEA, 18–19 mm diameter in 7 d at 25°C, surface white in the center, irregularly distributed to fascicles, aerial mycelium abundant, reverse dark reddish pigment. On CYA, 26–27 mm diameter in 7 d at 25°C, surface white in the center becoming light yellow towards the margins with white, irregularly distributed to fascicles, aerial mycelium abundant, forming several concentric rings on the surface, and a reverse yellowto-light yellowish pigment fascicles, aerial mycelium abundant, forming several concentric rings on the surface, which are reverse dark brown in color. On OA, 27–28 mm diameter in 7 d at 25°C, surface dirty white in the center becoming tan to light yellow towards the margins with dirty white, irregularly distributed tuffs of fascicles, reverse dark brown. Conidiophores arising from the agar surface from aerial hyphae or fascicles, 10.63–23.25 × 2.12–3.63 μm (mean ± SD: 16.28 ± 3.73 × 2.74 ± 0.39 μm, n = 30), hyaline, branched verticillately at 2–3 levels, with a terminal whorl of 1–8 phialides, and 1–2 lower nodes of 1–4 phialides, septate. Phialide hyaline, thin-walled, subulate, 7.81–13.67 × 3.04–3.88 μm (mean ± SD: 11.20 ± 1.83 × 3.61 ± 0.28 μm, n = 30) at the broadest part with periclinal thickening. Conidia fusiform to ellipsoidal to obovoid, hyaline, smooth, 3.80–6.15 × 2.18–3.88 μm (mean ± SD: 5.23 ± 0.54 × 3.07 ± 0.41 μm, n = 30), with a distinct hilum at both or at one end. Chlamydospores seen. No ascomatal state was observed.
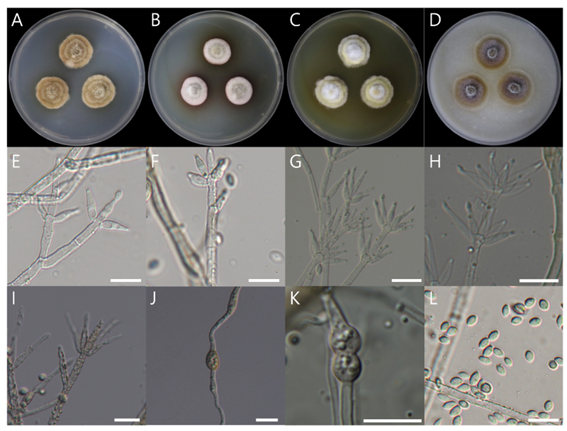
Fig. 4. Colony morphology and light microscopic images of Mariannaea humicola NIBRFGC000510020 grown on different media at 25°C for 7 d. A: PDA. B: MEA. C: CYA. D: OA. E–I: conidiophores with verticillate phialides. J–K: chlamydospores. L: conidia. Scale bar = 10 μm.
Notes
The strain NIBRFGC000510020 shows similar morphological characteristics of cultural and mycelial structures compared to those of the type strain M. humicola CBS 740.95 [16]. In the original description of the type strain, conidia are described as fusiform, ellipsoidal, or obovoid, hyaline, and smooth, with sizes ranging from 4.0–6.0 × 2.0–3.0 μm (mean ± SD: 5.0 ± 0.6 × 3.0 ± 0.3 μm, n = 30) which are similarly seen in the conidia of strain NIBRFGC000510020. The arising conidiophores from the agar surface from aerial hyphae or fascicles also like those of the strain NIBRFGC000510020, regarding size (80–100 μm long, axis 3–7 μm wide) and morphology of phialides which formed verticillately at 2–3 levels, with a terminal whorl of 1–5 phialides, and 1–2 lower nodes of 1–3 phialides. The ascomatal state was not observed in either strain. Unlike M. humicola CBS 740.95, chlamydospores were observed in the mycelium of NIBRFGC000510020. In conclusion, based on the phylogenetic and morphological analysis results, we identified strain NIBRFGC000510020 as M. humicola.
Strains examined. Gyorae Gotjawal Forest on Jeju Island, isolated from the mushroom hyphosphere soil of Armillaria mellea in 2022, strain T. pubescens DUCC21469 (NIBRFGC000510021).
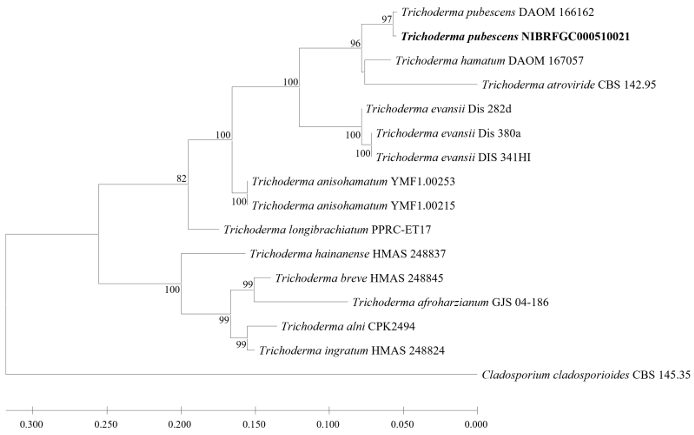
Fig. 5. Maximum likelihood (ML) phylogram constructed based on the concatenated ITS-tef1-α sequences. The number of nodes represents the reliability value (>70%) through 1,000 bootstrap replicates. Fungal strain isolated in this study is indicated in bold. Cladosporium cladosporioides CBS 145.35 is used as an outgroup. The reference sequences used to construct the ML phylogenetic tree are listed in Table 4.
Phylogenetic analysis (Fig. 5)
A ML phylogenetic tree was constructed based on the concatenated ITS and tef1-α sequences of Trichoderma spp., including those of the strain NIBRFGC000510021. The size of the ITS region of strain NIBRFGC000510021 was 364 bp, displaying 100% sequence similarity to that of the type strain Trichoderma pubescens DAOM 166162 (MN562094). The size of tef1-α was 573 bp, showing 99.18% sequence similarity to that of the type strain (LN897315). As a result of constructing an ML phylogenetic tree based on the 1,587 bp concatenated sequence including gaps, the strain NIBRFGC000510021 was grouped together with T. pubescens DAOM 166162 with a high node value. The most closely related species is T. hamatum.
Morphological characteristics (Fig. 6)
Description. On PDA, 89–90 mm diameter in 7 d at 25°C, floccose, circular, sometimes concentric, white, aerial mycelium limited, surface appearing downy, conidiation in numerous, white at first gradually turning bright green shades, yellowish, and exudate is not observed. On SNA, 89–90 mm diameter in 7 d at 25°C, floccose, circular, white in the center, sparse hyphae, aerial mycelium limited, and exudate is not observed. On CMA, 79–80 mm diameter in 7 d at 25°C, flat, circular, radial, white, aerial mycelium limited, and exudate is not observed. Hyphae were hyaline, branched, septate, thin-walled, 1–5 μm (mean ± SD: 3.05 ± 1.30 μm, n = 30). Conidiophore were hyaline, thin walled, coarse, developed at the branch of the hyphae, 8.42–11.44 × 2.30–4.60 μm (mean ± SD: 9.80 ± 1.28 × 3.14 ± 0.94 μm, n = 30), branched sterile conidiophore apices, usually at right angles with branches reflexed slightly towards the apex. Phialides were hyaline, thin walled, ampulliform to lageniform, 1–3 developed at the end or middle of the conidiophore, 6.10–16.09 × 2.11–3.93 μm (mean ± SD: 10.18 ± 3.03 × 3.18 ± 0.52 μm, n = 30). Conidia had smooth surface, without septum, dilute green, ellipsoidal, 3.55–5.31 × 2.43–3.51 μm (mean ± SD: 4.44 ± 0.39 × 2.96 ± 0.27 μm, n = 30), both ends broadly rounded, aggregated in a persistent mucilage. No sexual state was observed.
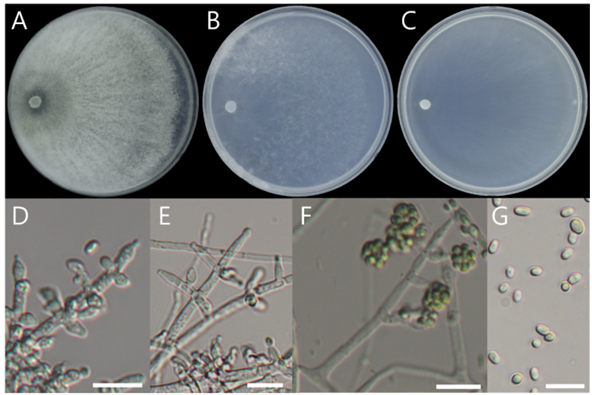
Fig. 6. Colony morphology and light microscopic images of Trichoderma pubescens NIBRFGC000510021 grown on different media at 25°C for 7 d. A: PDA. B: SNA. C: CMA. D–F: conidiophores and phialides. G: conidia. Scale bar = 10 μm.
Notes
The strain NIBRFGC000510021 showed similar morphological characteristics of cultural and mycelial structures compared to those of the type strain T. pubescens DAOM 166162 (T) [17]. T. hamatum which is a morphologically similar species and phylogenetically closely related species (Fig. 5) to T. pubescens has compact, bluish-green, hemispherical conidiogenous pustules and ellipsoidal conidia of moderate size [17]. T. pubescens can be distinguished from T. hamatum by its branching conidiophore apices, which give the surface of the conidiogenous pustules a cottony appearance. Thus, we identified the strain NIBRFGC000510021 as T. pubescens.
Nigrograna belongs to the family Nigrogranaceae of the order Pleosporales and is found in various habitats, including terrestrial, estuarine, and marine environments [18]. Recently, this genus included Nigrograna mackinnonii as the agent responsible for most cases of black-grain eumycetoma in Latin America [19]. N. hydei was initially reported as a saprophyte in Thailand [14] and was later identified as an endophyte of Elaeagnus angustifolia, a plant species that grows in the arid and cold regions of India [15]. Importantly, a recent report on deep cutaneous fungal infection indicated that N. hydei can infect both immunocompromised and immunocompetent individuals [20]. This study reports the first isolation of N. hydei from the hyphosphere soil under A. mellea in Korea, suggesting a wide range of habitats for this species. The ecological role of the isolated strain in the hyphosphere environment of mushrooms remains unknown; thus, further studies are required.
Mariannaea belongs to the family Nectriaceae of the order Hypocreales. Most Mariannaea species are terrestrial, with some newly discovered species found in freshwater habitats, suggesting ecological diversity [21]. M. humicola has been reported in decaying wood, in rhizosphere soil under Araucaria angustifolia, and marine environments [16,22]. M. humicola NIBRFGC000510020, isolated from the hyphosphere soil under A. mellea in the Gyorae Gotjawal Forest, represents a new ecological record for this species. This strain expanded the known habitat range of M. humicola and highlighted its adaptability to diverse ecological niches. Furthermore, M. humicola produces bioactive secondary metabolites with antifungal properties, suggesting its potential as a natural antifungal or biological control agents with applications in agriculture and pharmaceuticals [22].
Trichoderma belongs to the family Hypocreaceae of the order Hypocreales [23]. Trichoderma spp. are predominant over wide geographic regions in all climatic zones and are significant decomposers of woody and herbaceous materials [24]. They are also found in marine environment [25]. T. pubescens was first isolated in 1992 and has been reported in diverse habitats, including hardwood forest soils, mangrove soils, and tomatoes [18,26,27]. T. pubescens has demonstrated potential as an effective biocontrol agent, particularly for mitigating the effects of Rhizoctonia solani, a major cause of root rot in tomato plants [26]. Furthermore, from an environmental engineering perspective, its ability to degrade organic pollutants highlights its potential role in soil ecosystem restoration and sustainable agricultural management [27]. T. pubescens isolation from the hyphosphere soil under A. mellea in the Tamna Valley is the first report of this species in Korea, highlighting its ecological importance and adaptability to unique mushroom habitats on Jeju Island.
The authors declare no potential conflicts of interest.
This work was supported by a grant from the National Institute of Biological Resources (NIBR) funded by the Ministry of Environment (MOE) of the Republic of Korea (NIBR202203204). The Department of Microbiology was supported by the Research Focused-Department Promotion and Interdisciplinary Convergence Research Project as part of the Support Program for University Development for Dankook University in 2025.
1. Ko PY, Seok SJ, Lee HB, Ko HS, Jeun YC. Species diversity of spontaneous mushrooms on Jeju Island. Kor J Mycol 2014;42:104-32. https://doi.org/10.4489/KJM.2014.42.2.104
[DOI]
2. Yang JH, Oh SY, Kim W, Woo JJ, Kim H, Hur JS. Effect of isolation conditions on diversity of endolichenic fungal communities from a foliose lichen, Parmotrema tinctorum. J Fungi 2021;7:335. https://doi.org/10.3390/jof7050335
[DOI]
3. Woo JJ, Lücking R, Oh SY, Jeun YC, Hur JS. Two new foliicolous species of Strigula (Strigulaceae, Strigulales) in Korea offer insight in phorophyte-dependent variation of thallus morphology. Phytotaxa 2020;443:1-12. https://doi.org/10.11646/phytotaxa.443.1.1
[DOI]
4. Oh SY, Yang JH, Woo JJ, Oh SO, Hur JS. Diversity and distribution patterns of endolichenic fungi in Jeju Island, South Korea. Sustainability 2020;12:3769. https://doi.org/10.3390/su12093769
[DOI]
5. Lee TK, Han I, Kim MS, Seong HJ, Kim JS, Sul WJ. Characterization of a nifH-harboring bacterial community in the soil-limited Gotjawal forest. Front Microbiol 2019;10:1858. https://doi.org/10.3389/fmicb.2019.01858
[DOI]
6. Noh HJ, Kim YI, Lee DH, Ko PY, Park HS, Lee KH, Kim SH. Eleven previously unrecorded fungal species isolated from hyphosphere soil supporting wild mushrooms in Jeju Island. J Mushrooms 2023;21:228-40. https://doi.org/10.14480/JM.2023.21.4.228
[DOI]
7. Thompson JD, Higgins DG, Gibson TJ. CLUSTAL W: Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 1994;22:4673-80. https://doi.org/10.1093/nar/22.22.4673
[DOI]
8. Kumar S, Stecher G, Li M, Knyaz C, Tamura K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 2018;35:1547-9. https://doi.org/10.1093/molbev/msy096
[DOI]
9. Kimura MA. Simple method for estimating evolutionary rate of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 1980;16:111-120. https://doi.org/10.1007/BF01731581
[DOI]
10. White TJ, Bruns T, Lee S, Taylor J. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, editors. PCR protocols: A guide to methods and applications. New York: Academic Press; 1990. p. 315-22.
[DOI]
11. Cubeta MA, Echandi E, Abernethy T, Vilgalys R. Characterization of anastomosis groups of binucleate Rhizoctonia species using restriction analysis of an amplified ribosomal RNA gene. Phytopathology 1991;81:1395-400.
[DOI]
12. Vilgalys R, Hester M. Rapid genetic identification and mapping of enzymatically amplified ribosomal DNA from several Cryptococcus species. J Bacteriol 1990;172:4238-46. https://doi.org/10.1128/jb.172.8.4238-4246.1990
[DOI]
13. Evidente A, Ricciardiello G, Andolfi A, Sabatini MA, Ganassi S, Altomare C, Favilla M, Melck D. Citrantifidiene and citrantifidiol: Bioactive metabolites produced by Trichoderma citrinoviride with potential antifeedant activity toward aphids. J Agric Food Chem 2008;56:3569-73. https://doi.org/10.1021/jf073541h
[DOI]
14. Zhang JF, Liu JK, Thambugala KM, Yang J, Meng ZH, Liu ZY. Two new species and a new record of Nigrograna (Nigrogranaceae, Pleosporales) from China and Thailand. Mycol Progress 2020;19:1365-75. https://doi.org/10.1007/s11557-020-01633-0
[DOI]
15. Digra S, Nonzom S. New record of novel endophyte Nigrograna hydei from the Northern Himalayas, India. Czech Mycol 2023;75:105-15. https://doi.org/10.33585/cmy.75201
[DOI]
16. Lombard L, van der Merwe NA, Groenewald JZ, Crous PW. Generic concepts in Nectriaceae. Stud Mycol 2015;80:189-245. https://doi.org/10.1016/j.simyco.2014.12.002
[DOI]
17. Bissett J. A revision of the genus Trichoderma. III. Section Pachybasium. Can J Bot 1991;69:2373-417. https://doi.org/10.1139/b91-298
[DOI]
18. Kolařík M. New taxonomic combinations in endophytic representatives of the genus Nigrograna. Czech Mycol 2018;70:123-6. https://doi.org/10.33585/cmy.70202
[DOI]
19. Ahmed SA, González GM, Tirado-Sánchez A, Moreno-López LM, de Hoog S, Bonifaz A. Nigrograna mackinnonii, not Trematosphaeria grisea (syn., Madurella grisea), is the main agent of black grain eumycetoma in Latin America. J Clin Microbiol 2018;56:e01723-17. https://doi.org/10.1128/JCM.01723-17
[DOI]
20. Wang SF, Liu JY, Bao FF, Liu YX, Yu CP, Wu M, Zhang FR. First report of deep cutaneous fungal infection caused by Nigrograna hydei. Lancet Infect Dis 2024;24:e659. https://doi.org/10.1016/S1473-3099(24)00422-5
[DOI]
21. Hu DM, Wang M, Cai L. Phylogenetic assessment and taxonomic revision of Mariannaea. Mycol Progress 2017;16:271-83. https://doi.org/10.1007/s11557-016-1252-2
[DOI]
22. Botta L, Saladino R, Barghini P, Fenice M, Pasqualetti M. Production and identification of two antifungal terpenoids from the Posidonia oceanica epiphytic Ascomycota Mariannaea humicola IG100. Microb Cell Fact 2020;19:184. https://doi.org/10.1186/s12934-020-01445-7
[DOI]
23. Zare R, Gams W. More white verticillium-like anamorphs with erect conidiophores. Mycol Progress 2016;15:993-1030. https://doi.org/10.1007/s11557-016-1214-8
[DOI]
24. Hoyos-Carvajal L, Orduz S, Bissett J. Genetic and metabolic biodiversity of Trichoderma from Colombia and adjacent neotropic regions. Fungal Genet Biol 2009;46:615-31. https://doi.org/10.1016/j.fgb.2009.04.006
[DOI]
25. Saravanakumar K, Vivek R, Boopathy NS, Yaqian L, Kathiresan K, Chen J. Anticancer potential of bioactive 16-methylheptadecanoic acid methyl ester derived from marine Trichoderma. J Appl Biomed 2015;13:199-212. https://doi.org/10.1016/j.jab.2015.04.001
[DOI]
26. Narendran R, Kathiresan K, Sathishkumar RS, Kayalvizhi K, Sundaramanickam A. Bioremoval of toxic substances in synthetic wastewater using Trichoderma pubescens (NPK2), isolated from mangrove soil. Biocatal Agric Biotechnol 2019;19:101100. https://doi.org/10.1016/j.bcab.2019.101100
[DOI]
27. Behiry S, Soliman SA, Massoud MA, Abdelbary M, Kordy AM, Abdelkhalek A, Heflish A. Trichoderma pubescens elicit induced systemic resistance in tomato challenged by Rhizoctonia solani. J Fungi 2023;9:167. https://doi.org/10.3390/jof9020167
[DOI]
28. Glass NL, Donaldson GC. Development of primer sets designed for use with the PCR to amplify conserved genes from filamentous ascomycetes. Appl Environ Microbiol 1995;61:1323-30. https://doi.org/10.1128/aem.61.4.1323-1330.1995
[DOI]