Min-Gyu Kim1, Seong-Keun Lim1, Geun-Tae Kim1, Jun-Woo Choi1, Hae-Dam Kim1, Che-Yeon Park1, Seo-Ryeong Lee1, Seung-Yeol Lee1,2*, and Hee-Young Jung1,2
1Department of Plant Medicine, Kyungpook National University, Daegu 41566, Korea
2Institute of Plant Medicine, Kyungpook National University, Daegu 41566, Korea
*Correspondence to leesy1123@knu.ac.kr
Korean Journal of Mycology (Kor J Mycol) 2025 December, Volume 53, Issue 4, pages 237-243.
https://doi.org/10.4489/kjm.2025.53.4.1
Received on October 17, 2025, Revised on November 10, 2025, Accepted on November 10, 2025, Published on December 31, 2025.
Copyright © The Korean Society of Mycology.
This is an Open Access article which is freely available under the Creative Commons Attribution-Non-Commercial 4.0 International License (CC BY-NC) (https://creativecommons.org/licenses/by-nc/4.0/).
Two fungal strains, designated as KNUF-21-062 and KNUF-23-215, were isolated from soil sample collected from Daegu provinces, Korea. Based on sequence analysis of the internal transcribed spacer (ITS) regions, the isolated strains were identified as members of the genera Penicillium and Talaromyces. For identification to species, each strain’s morphological characteristics were examined, and phylogenetic analyses were performed. To observe their morphological characteristics, the strains were cultured on potato dextrose agar, malt extract agar, czapek yeast extract agar, yeast extract sucrose agar, dichloran 18% glycerol agar and creatine sucrose agar. Strain KNUF-21-062 had biverticillate conidiophores, ampulliform phialides, and globose conidia, and strain KNUF-23-215 exhibited monoverticillate conidiophores, often solitary phialides, and subglobose conidia. Phylogenetic trees were constructed based on the ITS regions and the β-tubulin, calmodulin, and RNA polymerase II subunit genes using the maximum-likelihood method. Strains KNUF21-062 and KNUF-23-215 clustered with P. euglaucum CBS 323.71T and T. purpureus CBS 475.71T, respectively, which confirmed the identifications made based on morphology. Thus, morphological and phylogenetic analyses supported the taxonomic placement of isolated strains in Penicillium euglaucum and Talaromyces purpureus, two species previously unreported in Korea.
Morphology, Penicillium euglaucum, Phylogenetic analyses, Soil-derived fungi, Talaromyces purpureus
The order Eurotiales is one of the most abundant fungal groups, comprising five families and approximately 28 genera, including Penicillium and Talaromyces [1,2]. In 1809, the genus Penicillium first described by Link, who detailed their distinctive brush-like conidiophores [3]. Over the years, the taxonomy of Penicillium has undergone significant change, evolving from systems based on cultural characteristics to a classification system based on multi-locus phylogenies [1,2,4,5]. Traditionally, Talaromyces accommodated sexual forms of Penicillium species, but in the last decade, changes in nomenclature rules and advancements in phylogenetic analysis techniques have replaced the generic concept of these taxa [6,7].
When implemented, the current classification system, supported by four-gene phylogenies, reorganized Penicillium taxonomy [7]. Teleomorphs and anamorphs previously placed in Penicillium were regrouped based on DNA sequence data, and the implementation of nomenclatural priority and single name nomenclature led to the transfer of the majority of species from Penicillium subgenus Biverticillium to Talaromyces [7,8]. Taxonomy within these genera has been continuously revised with phylogenetic analyses, leading to the division of Penicillium into two subgenera, 32 sections, and 89 series and Talaromyces into eight sections [1,2,7–10]. Additionally, the availability of reliable and comprehensive sequence data has accelerated the discovery and description of new species in both genera [1,2]. Hence, the number of accepted species has grown rapidly, with 535 species of Penicillium and 203 species of Talaromyces currently recognized worldwide [2].
This study aimed to investigate the diversity of penicillium-like fungi in Korean soil. Both morphological and molecular characteristics were used to identify isolated strains, providing a comprehensive understanding of their taxonomy. These strains may be useful in future research and could potentially benefit human activities.
The fungal strains used in this study were isolated from soil samples collected in Yongsu-dong, Donggu, Daegu (35°59’33.9″N, 128°41’12.7″E), Korea and Guji-myeon, Dalseong-gun, Daegu (35°41’42.9″N, 128°22’24.2″E), Korea, using the serial dilution technique. Soil samples (1 g) was immersed in 9 mL of distilled water and vortexed to thoroughly mix them. The mixture was serially diluted from 10−1 to 10−5, and 100 μL aliquots from each dilution were spread onto potato dextrose agar (PDA; Difco, Detroit, MI, USA) plates, which were then incubated at 25°C for 7 days. Colonies displaying signs of germination were then subcultured to new PDA plates, with incubation continuing at 25°C. Fungal strains KNUF-21062 and KNUF-23-215 were chosen for molecular analyses and cultural and morphological evaluations. Both strains have been deposited at the National Institute of Biological Resources (NIBR) as metabolically inactive cultures, KNUF-21-062 under accession no. NIBRFGC000509192 and KNUF-23-215 under NIBRFGC000510716.
The isolates were cultured on various agar media to observe the cultural characteristics of the two strains. Strains KNUF-21-062 and KNUF-23-215 were cultured on PDA, Czapek yeast extract agar (CYA; MB Cell, KisanBio, Seoul, Korea), malt extract agar (MEA; Difco, Detroit, MI, USA), dichloran 18% glycerol agar (DG18; 31.5 g of dichloran glycerol agar base, 220 g of glycerol, 1 mL of trace element stock solution, 0.05 g of chloramphenicol, and 20 g of agar in distilled water to 1,000 mL), yeast extract sucrose agar (YES; 4 g of yeast extract, 20 g of sucrose, 1 g of KH2 PO4 , 0.5 g of MgSO4 , and 15 g of agar in distilled water to 1,000 mL), and creatine sucrose agar (CREA; 30 g of sucrose, 3 g of creatine, 1.6 g of K3 PO4 ·7H2 O, 0.5 g of MgSO4 ·7H2 O, 0.5 g of KCl, 0.01 g of FeSO4 ·7H2 O, 1 mL of trace elements stock solution, 0.05 g of bromocresol purple, and 20 g of agar in distilled water to 1,000 mL); strain KNUF-23-215 was also cultured on oatmeal agar (OA; Difco, Detroit, MI, USA) [10]. All cultures were incubated at 25°C for 7 days in darkness [10]. And then cultural characteristics, including the diameter and coloration of the colony, and morphological characteristics, including conidiophores, stipes, metulae, phialides, and conidia were observed. Morphological characteristics were observed using a light microscope (BX-50; Olympus, Tokyo, Japan).
For each strain, total genomic DNA was extracted from growing colony on PDA using the HiGene™ Genomic DNA Prep Kit (Biofact, Daejeon, Korea) following the manufacturer’s instructions. By using the primer pairs ITS1F/ITS4, Bt2a/Bt2b, CMD5/CMD6, and RPB2-5f/RPB2-7cR, the internal transcribed spacer (ITS) regions, β-tubulin (BenA), calmodulin (CaM), and RNA polymerase II subunit (RPB2) were amplified, respectively [11–14]. Amplification was confirmed by electrophoresis using HP Agarose (Biopure, Cambridge, MA, USA) 1.0% gels. Purification of amplified products were conducted by using ExosapIT (Thermo Fisher Scientific, Waltham, MA, USA) and sequencing services were provided by Bioneer (Daejeon, Korea).
For the phylogenetic analyses, outgroup species were selected based on analyses in previous studies, and additional related species were added based on BLAST (Basic Local Alignment Search Tool) results [1,9]. All sequences were obtained from the National Center for Biotechnology Information (NCBI) database (Table 1). Maximum-likelihood (ML) trees were constructed using the general time reversible model in MEGA11 based on the concatenated alignments of the four loci, and bootstrap values were generated based on 1,000 replications [15,16].
Table 1. GenBank accession numbers of fungal strains used for phylogenetic analyses in this study
테일블
ITS: internal transcribed spacer regions; BenA: β-tubulin; CaM: calmodulin; RPB2: RNA polymerase II subunit.
TType strain. The strains isolated in this study are indicated in bold.
Colony diameters (mm) after 7 days at 25°C: 22–24 on PDA, 21–23 on MEA, 15–17 on CYA, 22–24 on YES, 17–19 on DG18, and 9–10 on CREA (Fig. 1A).
After 7 days of incubation at 25°C, growth patterns differed across the tested media. On PDA: colonies umbonate, sulcate; margin entire, with a velutinous texture; mycelium white; exudates absent, and soluble pigments light yellow; reverse light yellow (Fig. 1A). On MEA: colonies flat, sulcate; margin entire, with a velvety texture; mycelium white; exudates clear droplets in center, and soluble pigments light yellow; reverse light yellow (Fig. 1A). On CYA: colonies umbonate, sulcate; margin entire, with a velutinous texture; mycelium white; exudates absent, and soluble pigments light yellow; reverse yellow brown (Fig. 1A). On YES: colonies umbonate, sulcate; margin entire, with a floccose texture; mycelium white; exudates absent, and soluble pigments light yellow; reverse yellow (Fig. 1A). On DG18: colonies umbonate, sulcate; margin entire, with a floccose texture; mycelium white; exudates absent, and soluble pigments light yellow; reverse yellow (Fig. 1A). On CREA: colonies sparse, compact, with limited radial expansion; acid production absent (Fig. 1A). When compared the cultural characteristics of strain KNUF-21-062 with Penicillium euglaucum CBS 323.71T, strain KNUF-21-062 growth slower than CBS 323.71T in CYA (15–17 vs. 23–29 mm) and DG18 (17–19 vs. 23–29 mm) (Fig. 1A) [17]. However, both produces yellow soluble pigments in CYA and YES media which are identical to P. euglaucum CBS 323.71T (Fig. 1A) [17].
Morphological characteristics: conidiophores biverticillate (Fig. 1B–D), with smooth walled stipes, 8.5–54.5 × 1.5–2.8 µm (n = 30) (Fig. 1B–D); metulae 11.3–18.8 × 1.5–3.2 µm (n = 50) (Fig. 1C and D); phialides ampulliform, 6.8–9.2 × 2.1–3.3 µm (n = 50) (Fig. 1B–D); conidia globose and finely roughened, 2.1–2.5 × 2.2–2.7 (n = 50) μm (Fig. 1E and F). The morphological characteristics of strain KNUF-21-062 were identical to those previously reported for Penicillium euglaucum CBS 323.71T (Table 2).
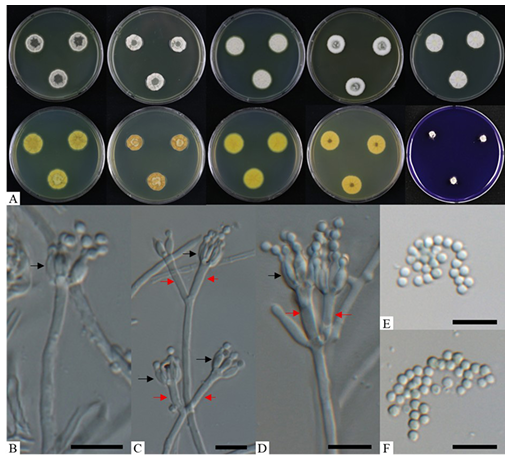
Fig. 1. Morphological characteristics of Penicillium euglaucum (KNUF-21-062). A: colonies after 7 days at 25°C, from left to right (top row) PDA obverse, CYA obverse, YES obverse, DG18 obverse, and MEA observe; (bottom row) left to right PDA reverse, CYA reverse, YES reverse, DG18 reverse, and CREA obverse; B–D: biverticillate conidiophores with metulae (red arrows) and phialides (black arrows); E, F: conidia. Scale bars = 10 µm.
Table 2. Morphological characteristics of Penicillium euglaucum KNUF-21-062 and Penicillium euglaucum CBS 323.71T
테일블
CYA: Czapek yeast extract agar; MEA: malt extract agar; YES: yeast extract sucrose agar; DG18: dichloran 18% glycerol agar; CREA: creatine sucrose agar.
aFungal strain in this study; bSource of descriptions [17]; TType strain.
The phylogenetic analysis for strain KNUF-21-062 was based on the ITS regions and the BenA, CaM, and RPB2. The sequences obtained from strain KNUF-21-062 were 597 bp for the ITS regions and 430, 632, and 1064 bp for BenA, CaM, and RPB2, respectively. Based on the BLASTn results, the strain KNUF21-062 belongs to the genus Penicillium and is closely related to P. euglaucum CBS 323.71T (sequence similarity for ITS = 99.8%, BenA = 99.3%, CaM = 98.7%, RPB2 = 98.7%) which is classified in subgenus Penicillium section Citrina series Euglauca. ML phylogenetic tree of section Citrina were constructed based on the concatenated sequences of four loci. The strain KNUF-21-062 formed a same cluster with P. euglaucum CBS 323.71T in ML phylogenetic tree identifying KNUF-21-062 as P. euglaucum (Fig. 2). Thus, the comparison of morphological characteristics and phylogenetic analysis, the strain KNUF-21-062 was identified as P. euglaucum.
Colony diameters (mm) after 7 days at 25°C: 24–25 on PDA, 23–25 on MEA, 14–15 on CYA, 17–19 on YES, 12–13 on DG18, 18–20 on OA, and 4–6 on CREA (Fig. 3A).
After incubation at 25°C for 7 days, colony growth patterns varied across the tested media. On PDA: colonies flat; margin entire, with a yeast-like slimy texture; mycelium white and pinkish red; exudates and soluble pigments absent; reverse light white and pinkish red (Fig. 3A). On MEA: colonies flat; margin entire, with a yeast-like slimy texture; mycelium white; exudates and soluble pigments absent; reverse brownish orange (Fig. 3A). On CYA: colonies flat; margin entire, with a yeast-like slimy texture; mycelium white and pinkish white; exudates and soluble pigments absent; reverse light orange and grayish yellow (Fig. 3A).
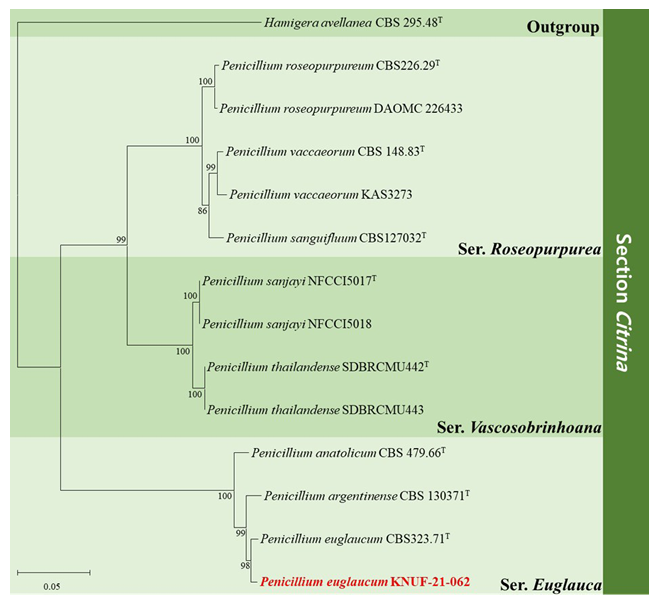
Fig. 2. Maximum-likelihood phylogenetic tree of Penicillium section Citrina, including series Roseopurpurea, series Vascosobrinhoana, and series Euglauca based on dataset of combining the ITS regions, BenA, CaM and RPB2 sequences. Hamigera avellanea CBS 295.48T was used as the outgroup. The numbers above/below the branches are bootstrap values, including only those greater than 80% (percentage of 1,000 replications). The Penicillium species isolated in this study is highlighted in bold and red. The scale bar represents 0.05 substitutions per nucleotide position.
On YES: colonies flat; margin entire, with a yeast-like slimy texture; mycelium white and red; exudates absent, and soluble pigments red; reverse red and brownish red (Fig. 3A). On DG18: colonies flat; margin entire, with a yeast-like slimy texture; mycelium white and pinkish red; exudates and soluble pigments absent; reverse light orange and pinkish red (Fig. 3A). On OA: colonies flat; margin entire, with a yeast-like slimy texture; mycelium white and pinkish red; exudates and soluble pigments absent (Fig. 3A). On CREA: colonies rarely grew; acid production absent (Fig. 3A). When compared the cultural characteristics of strain KNUF-23-215 with Talaromyces purpureus CBS 475.71T, strain KNUF-23-215 growth faster than CBS 475.71T in CYA (14–15 vs. 2–4 mm), MEA (23–24 vs. 15–16 mm), YES (17–19 vs. 7–8 mm) and DG18 (12–13 vs. 1–2 mm) (Fig. 3A) [9]. However, the color of mycelium and characteristics from all media were identical to T. purpureus CBS 475.71T [9].
Morphological characteristics: conidiophores monoverticillate, with smooth walled stipes, 9.118.3 × 1.9–2.3 µm (n = 30), often with a solitary phialide (Fig. 3B–E); phialides ampulliform, 6.010.1 × 2.3–3.3 µm (n = 30) (Fig. 3B–E); conidia rough, subglobose, 3.3–3.6 × 2.8–3.4 µm (n = 50) (Fig. 3F). Strain KNUF-23-215’s cultural and morphological characteristics were similar to those previously reported for T. purpureus (Table 3).
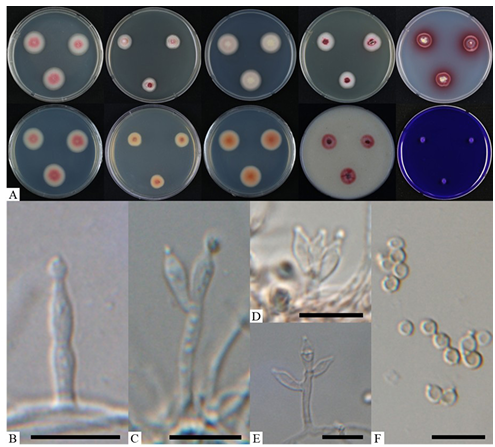
Fig. 3. Morphological characteristics of Talaromyces purpures (KNUF-23-215). A: colonies after 7 days at 25°C, from left to right (top row) PDA obverse, CYA obverse, MEA obverse, DG18 obverse, and YES observe; (bottom row) left to right PDA reverse, CYA reverse, MEA reverse, OA obverse, and CREA obverse; B: a conidiophore with solitary phialide; C–E: monoverticillate conidiophores; F: conidia. Scale bars = 10 µm.
Table 3. Morphological characteristics of Talaromyces purpureus KNUF-23-215 and Talaromyces purpureus CBS 475.71T
table
CYA: Czapek yeast extract agar; MEA: malt extract agar; YES: yeast extract sucrose agar; DG18: dichloran 18% glycerol agar; OA: oatmeal agar; CREA: creatine sucrose agar.
aFungal strain in this study; bSource of descriptions [9]; TType strain.
The phylogenetic analysis of strain KNUF-23-215 were conducted through ITS regions, BenA and RPB2 sequences. The sequence obtained from KNUF-23-215 were 560 bp for ITS regions, 418 bp and 1034 bp for BenA and RPB2, respectively. Based on the BLASTn results, the strain KNUF-23-215 belongs to the genus Talaromyces and is closely related to T. purpureus CBS 475.71T (sequence similarity for ITS = 100%, BenA = 100%, RPB2 = 99.5%) which is classified in section Purpurea. ML phylogenetic tree of section Purpurei were constructed based on the concatenated three loci, identifying that the strain KNUF23-215 as T. purpureus which formed monophyletic group with T. purpureus CBS 475.71T (Fig. 4). Thus, the comparison of morphological characteristics and phylogenetic analysis, the strain KNUF-23-215 was identified as T. purpureus.
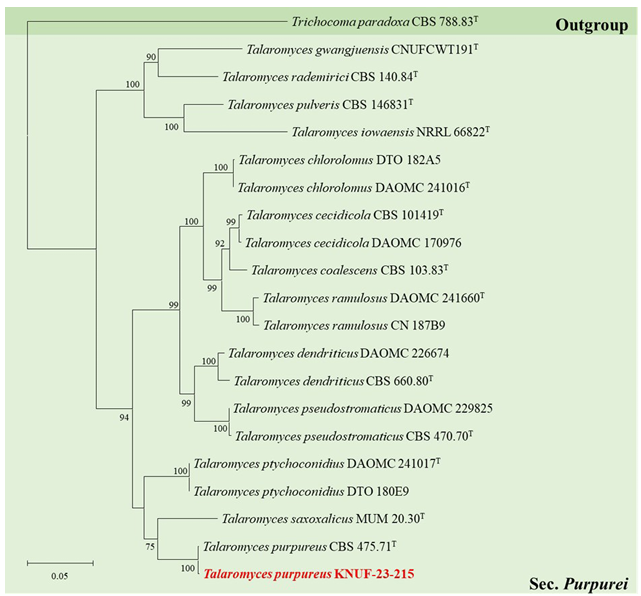
Fig. 4. Maximum-likelihood phylogenetic tree of Talaromyces section Purpurei based on a combined dataset of ITS regions, BenA and RPB2 sequences. Trichocoma paradoxa CBS 788.83T was used as an outgroup. The numbers above/below the branches are bootstrap values, including only those greater than 70% (percentage of 1,000 replications). The Talaromyces species isolated in this study is highlighted in bold and red. The scale bar represents 0.05 substitutions per nucleotide position.
Taxonomic research on the order Eurotiales has expanded globally following the adoption of the “one fungus, one name” principle established by the International Code of Nomenclature for algae, fungi and plants (ICBN) [1,2]. Current taxonomic research on Eurotiales in Korea focuses on discovering and characterizing novel and previously unreported species of Penicillium and Talaromyces from diverse habitats, contributing to the reporting of 155 Penicillium species and 37 Talaromyces species now reported in the region [18–21]. Throughout this study, two unrecorded Penicillium and Talaromyces species isolated in Korea were identified as P. euglaucum (subgenus Aspergilloides, section Citrina, series Euglauca) and T. purpureus (section Purpurei), respectively. Penicillium series Euglauca comprises three species reported globally, none of which have been recorded in Korea [1,21]. Similarly, Talaromyces section Purpurei encompasses 15 species worldwide, of which only two species (T. cecidicola and T. gwangjuensis) have been reported from Korea [1,2,21].
Penicillium and Talaromyces species are notable for their extrolite production [1,2,9]. For instance, members of Penicillium series Euglauca, P. anatolicum and P. argentinense, produce curvularin and dehydrocurvularin which exhibit phytotoxic and antifungal activities capable of suppressing soilborne pathogens [1,22]. In addition, P. euglaucum produces terrein, a secondary metabolite with phytotoxic activity that affects plant growth and induces lesions on the fruit surface [1,23]. Similarly, T. coalescens and T. dendriticus members of Talaromyces section Purpurei produce penicillides which exhibit antibacterial, antifungal, anti-inflammatory, and cytotoxic activities [9,24]. Species belonging to this section are also known to produce diverse extrolites, including apiculides, pentacecilides, and vermiculin [9].
In conclusion, Penicillium and Talaromyces species exhibit distinctive characteristics, and the discovery of the unreported species P. euglaucum and T. purpureus in Korea highlights the importance of further research into their ecological and biological roles. While this study focused on these two strain’s morphological characteristics and taxonomy, more extensive investigations are needed to fully understand the bioactive compounds and pathogenicity profiles of Penicillium and Talaromyces species, as such research may identify beneficial applications.
The authors declare no conflict of interest.
This research was supported by a grant from the National Institute of Biological Resources (NIBR), funded by the Ministry of Environment (MOE) of the Republic of Korea (NIBR202102107 and NIBR202304104).
1. Houbraken J, Kocsubé S, Visagie CM, Yilmaz N, Wang XC, Meijer M, Kraak B, Hubka V, Bensch K, Samson RA, et al. Classification of Aspergillus, Penicillium, Talaromyces and related genera (Eurotiales): An overview of families, genera, subgenera, sections, series and species. Stud Mycol 2020;95:5-169. https://doi.org/10.1016/j.simyco.2020.05.002
[DOI]
2. Visagie CM, Yilmaz N, Kocsubé S, Frisvad JC, Hubka V, Samson RA, Houbraken J. A review of recently introduced Aspergillus, Penicillium, Talaromyces and other Eurotiales species. Stud Mycol 2024;107:1-66. https://doi.org/10.3114/sim.2024.107.01
[DOI]
3. Link L. Observationes in ordines plantarum naturales. Dissertatio 1. Mag Ges Naturf Freunde Berlin 1809;3:3-42.
4. Thom C. The Penicillia. Baltimore (MD): Willams & Wilkins Company; 1949.
5. Pitt JI. The genus Penicillium and its teleomorphic states Eupenicillium and Talaromyces. London: Academic Press Inc. Ltd.; 1979.
6. Benjamin CR. Ascocarps of Aspergillus and Penicillium. Mycologia 1955;47:669-87. https://doi.org/10.2307/3755578
[DOI]
7. Houbraken J, Samson RA. Phylogeny of Penicillium and the segregation of Trichocomaceae into three families. Stud Mycol 2011;70:1-51. https://doi.org/10.3114/sim.2011.70.01
[DOI]
8. Samson RA, Yilmaz N, Houbraken J, Spierenburg H, Seifert KA, Peterson SW, Varga J, Frisvad JC. Phylogeny and nomenclature of the genus Talaromyces and taxa accommodated in Penicillium subgenus Biverticillium. Stud Mycol 2011;70:159-83. https://doi.org/10.3114/sim.2011.70.04
[DOI]
9. Yilmaz N, Visagie CM, Houbraken J, Frisvad JC, Samson RA. Polyphasic taxonomy of the genus Talaromyces. Stud Mycol 2014;78:175-341. https://doi.org/10.1016/j.simyco.2014.08.001
[DOI]
10. Visagie CM, Houbraken J, Frisvad JC, Hong SB, Klaassen CHW, Perrone G, Seifert KA, Varga J, Yaguchi T, Samson RA. Identification and nomenclature of the genus Penicillium. Stud Mycol 2014;78:343-71. https://doi.org/10.1016/j.simyco.2014.09.001
[DOI]
11. White TJ, Bruns T, Lee S, Taylor J. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, editors. PCR protocols: A guide to methods and applications. San Diego: Academic Press; 1990. p. 315-22. https://doi.org/10.1016/B978-0-12-372180-8.50042-1
[DOI]
12. Glass NL, Donaldson GC. Development of primer sets designed for use with the PCR to amplify conserved genes from filamentous ascomycetes. Appl Environ Microbiol 1995;61:1323-30. https://doi.org/10.1128/aem.61.4.1323-1330.1995
[DOI]
13. Hong SB, Cho HS, Shin HD, Frisvad JC, Samson RA. Novel Neosartorya species isolated from soil in Korea. Int J Syst Evol Microbiol 2006;56:477-86. https://doi.org/10.1099/ijs.0.63980-0
[DOI]
14. Liu YJ, Whelen S, Hall BD. Phylogenetic relationships among Ascomycetes: Evidence from an RNA polymerase II subunit. Mol Biol Evol 1999;16:1799-808. https://doi.org/10.1093/oxfordjournals.molbev.a026092
[DOI]
15. Felsenstein J. Evolutionary trees from DNA sequences: A maximum likelihood approach. J Mol Evol 1981;17:368-76. https://doi.org/10.1007/BF01734359
[DOI]
16. Tamura K, Stecher G, Kumar S. MEGA11: Molecular evolutionary genetics analysis version 11. Mol Biol Evol 2021;38:3022-7. https://doi.org/10.1093/molbev/msab120
[DOI]
17. Houbraken J, Frisvad JC, Samson RA. Taxonomy of Penicillium section Citrina. Stud Mycol 2011;70:53-138. https://doi.org/10.3114/sim.2011.70.02
[DOI]
18. Nguyen TTT, Kang KH, Kim DH, Kim SJ, Mun HY, Cheon W, Lee HB. Additions to the knowledge of the fungal order Eurotiales in Korea: Eight undescribed species. Mycobiology 2023;51:417-35. https://doi.org/10.1080/12298093.2023.2290759
[DOI]
19. Kim MG, Lim SK, Back CG, Oh Y, Cheon W, Mun HY, Lee SY, Jung HY. Isolation and characterization of four unreported Penicillium species isolated from the freshwater environments in Korea. Mycobiology 2025;53:269-79. https://doi.org/10.1080/12298093.2025.2473141
[DOI]
20. Kim MG, Lim SK, Choi JW, Goh J, Lee CS, Mun HY, Lee SY, Jung HY. Identification and characterization of five previously unrecorded Penicillium species of subgenus Aspergilloides isolated in Korea. Mycobiology 2025;53:731-46. https://doi.org/10.1080/12298093.2025.2550814
[DOI]
21. National Institute of Biological Resources (NIBR). National list species of Korea [Internet]. Incheon: NIBR; 2024 [cited 2025 Sep 20]. Available from: https://species.nibr.go.kr
22. Meepagala KM, Johnson RD, Duke SO. Curvularin and dehydrocurvularin as phytotoxic constituents from Curvularia intermedia infecting Pandanus amaryllifolius. J Agric Chem Environ 2016;5:12-22. http://dx.doi.org/10.4236/jacen.2016.51002
[DOI]
23. Zaehle C, Gressler M, Shelest E, Geib E, Hertweck C, Brock M. Terrein biosynthesis in Aspergillus terreus and its impact on phytotoxicity. Chem Biol 2014;21:719-31. https://doi.org/10.1016/j.chembiol.2014.03.010
[DOI]
24. Salvatore MM, Nicoletti R, Fiorito F, Andolfi A. Penicillides from Penicillium and Talaromyces: Chemical structures, occurrence and bioactivities. Molecules 2024;29:3888. https://doi.org/10.3390/molecules29163888
[DOI]