Hyeok Park* , Sumi Kim, and Gye-Ryeong Bak
Highland Agriculture Research Institute, National Institute of Crop and Food Science, Pyeongchang 25342, Korea
*Correspondence to mycoph@korea.kr
Korean Journal of Mycology (Kor J Mycol) 2025 December, Volume 53, Issue 4, pages 237-243.
https://doi.org/10.4489/kjm.2025.53.4.1
Received on October 17, 2025, Revised on November 10, 2025, Accepted on November 10, 2025, Published on December 31, 2025.
Copyright © The Korean Society of Mycology.
This is an Open Access article which is freely available under the Creative Commons Attribution-Non-Commercial 4.0 International License (CC BY-NC) (https://creativecommons.org/licenses/by-nc/4.0/).
In this study, endophytic fungal strains were isolated from the leaves, roots, and stems of potato plants from spring-cultivated potato fields in Gangwon-do and Jeju-do, Korea. Morphological characteristics of the isolates were observed, and molecular analyses were conducted using the internal transcribed spacer, large subunit rDNA, and second-largest subunit of RNA polymerase II regions. Four endophytic fungal species, previously unrecorded in Korea and belonging to the families Chaetomiaceae and Didymellaceae, were identified. The morphological and phylogenetic analyses of these species have also been reported.
Collariella, Epicoccum, Neoascochyta, Ovatospora, Solanum tuberosum
Potato (Solanum tuberosum L.) is the second most-produced food crop in Korea [1], and springcultivated potatoes constitute approximately 63% of national production. Gangwon-do and Jeju-do are the leading provinces in spring-cultivated potato yield (kg/10 a) [2].
Fungal endophytes live symbiotically within plant tissues without causing disease. They protect the host plants from external pathogens by secreting secondary metabolites [3]. Several studies have reported endophytic fungi in diverse potato tissues [4–6], and some fungal species confer resistance against major potato diseases, such as late blight [5] or brown leaf spot [6]. Consequently, endophytic fungi have been investigated as potential biological control agents for reducing dependence on chemical pesticides. In this study, we have reported the morphological characteristics and phylogenetic analyses of four endophytic fungal species previously unrecorded in Korea, belonging to the families Chaetomiaceae and Didymellaceae, and isolated from spring-cultivated potatoes in Gangwon-do and Jeju-do.
Sampling was conducted in spring potato cultivation areas in Gangneung-si (Gangwon-do, 37°46′44.7″N, 128°55′34.8″E) and Jeju-si (Jeju-do, 33°28′43.9″N, 126°47′02.0″E) between May and June 2025. Healthy potato plants with no disease symptoms were collected, transported to the laboratory within 24 h, and rinsed with distilled water. Leaf and stem tissues were surface-sterilized in 1% NaOCl for 1 min and 70% ethanol for 2 min, excised into small pieces, and placed on potato dextrose agar (PDA) [7]. Root tissues were surface-sterilized in 3% NaOCl for 3 min and 70% ethanol for 2 min, treated with streptomycin (100 μg·mL-1) for 10 min, and placed on water agar (WA) [8].
When mycelia emerged from the tissue during cultivation at 25°C, they were sub-cultured on PDA. Morphological characteristics of the colonies, mycelia, and spores were observed (Table 1) on PDA, malt extract agar (MEA), and oatmeal agar (OA) by using an optical microscope (Nikon Eclipse E600, Nikon, Tokyo, Japan).
Table 1. Morphological characteristics of endophytic fungal strains isolated in this study
table
OA: oatmeal agar; PDA: potato dextrose agar; MEA: malt extract agar.
Genomic DNA was extracted from the mycelia by using the DiaStar™ Direct Multiplex PCR Kit (Solgent Co., Ltd., Daejeon, Korea), and PCR amplifications were performed. The internal transcribed spacer (ITS) region was amplified with ITS1F/ITS4 primers [9], the large subunit (LSU) rDNA with LR0R/LR16 primers [10], and the RNA polymerase II second-largest subunit (RPB2) region with fRPB2-5f/fRPB2-7cR primers [11]. PCR products were electrophoresed on a 1.5% agarose gel. Each DNA band was confirmed, and DNA sequencing was performed by Solgent Co., Ltd. (Daejeon, Korea). The resulting DNA sequences were compared against the NCBI BLAST database to determine the closest species matches, and alignments with related taxa were used to construct neighbor-joining phylogenetic trees in MEGA 7 [12]. All DNA sequences obtained in this study were registered in GenBank (accession numbers provided in the Results section), and the isolates were deposited in the Korean Agricultural Culture Collection (KACC), National Institute of Agricultural Sciences.
Collariella bostrychodes (Zopf) X. Wei Wang & Samson, Stud. Mycol. 84: 179 (2016) [MB#818862]
This strain was isolated from a potato root collected from Gangneung-si. On PDA incubated at 25°C for 7 days, colonies reached an average diameter of 37.1 mm; colonies were pinkish-white, flat, and exhibited an entire margin due to densely radiating mycelia (Fig. 1A). On MEA, the colonies grew slower (avg. 27.0 mm), displaying an undulate margin and sparser mycelia. The colony was light white or gray and flat on the plate (Fig. 1B). On OA, growth was the fastest (avg. 43.9 mm); the colonies were white at the margin, gray toward the center, and darkened as the ascomata matured (Fig. 1C). Asci were produced in a broomlike arrangement from the ascomata, containing lemon-shaped to ovoid ascospores with a pointed end; pigmentation ranged from light to dark brown at maturity (Fig. 1D & 1E). The size was (4.00–) 4.96 (–5.88) × (2.98–) 4.10 (–5.36) μm in diameter (n = 20).
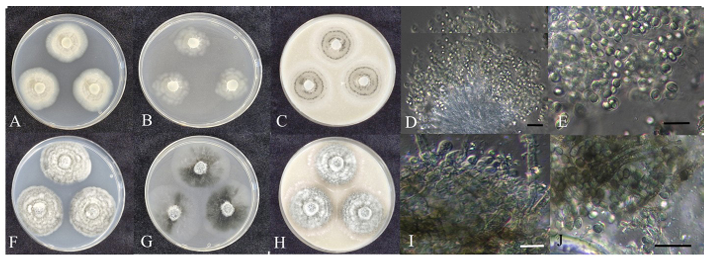
Fig. 1. Cultural characteristics of endophytic fungal strains belonging to Chaetomiaceae. Colonies of Collariella bostrychodes strain HARI25E108 grown for 7 days on PDA (A), MEA (B), and OA (C) and ascus and ascospores (D, E); Colonies of Ovatospora brasiliensis strain HARI25E097 grown for 7 days on PDA (F), MEA (G), and OA (H) and ascomata (I) and ascospores (J) (scale bars: D, I = 20 μm; E, J = 10 μm). PDA: potato dextrose agar; MEA: malt extract agar; OA: oatmeal agar.
Phylogenetic analysis. ITS sequence analysis showed 99.64% similarity to C. bostrychodes strain CBS 586.83 (MH861661.1), and the LSU sequence showed 97.07% similarity to C. bostrychodes strain CBS 586.83 (MH873374.1). In the concatenated ITS+LSU neighbor-joining tree, strain HARI25E108 formed a monophyletic clade with strains CBS 163.73 and CBS 586.83 (Fig. 2).
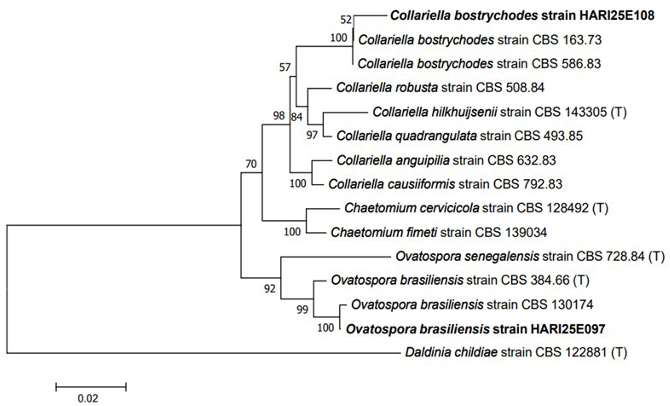
Fig. 2. Neighbor-joining phylogenetic tree of C. bostrychodes and O. brasiliensis on the basis of a concatenated alignment of internal transcribed spacer (ITS) and large subunit (LSU) rDNA sequences. Daldinia childiae was used as an outgroup. Numbers on branches indicate bootstrap values (1,000 replicates). Fungal strains isolated in this study are in bold. (T) indicates the type strains.
Specimen examined. Korea: Gangneung-si, Gangwon-do, 37°47′03.1″N, 128°55′25.6″E, June 29, 2025, isolated from healthy root of Solanum tuberosum, strain HARI25E108, KACC411096 (deposit No.), GenBank No. PX285310 (ITS), PX285311 (LSU).
Note. On OA, strain HARI25E108 was white, as described in the original description, but the color turned gray as the ascomata matured. The colonies were slightly larger than strain CBS 163.73 from the original description [13]. The asci darkened as the ascospores matured and had a liminiform shape, which was consistent with strain CBS 163.73; however, among the ascospores observed in this study, ovate shapes were rarely observed. These were thought to be immature ascospores. The asci formed a radial collar-like arrangement. Together with molecular phylogenetic evidence, these features support the identification of C. bostrychodes.
Ovatospora brasiliensis (Bat. & Pontual) X. Wei Wang & Samson, Stud. Mycol. 84: 207 (2016) [MB#818851]
This strain was isolated from a potato root collected from Gangneung-si. After 7 days at 25°C, colonies on PDA averaged 41.1 mm in diameter; colonies were initially dark beige with dark-gray concentric growth and a generally smooth margin due to dense hyphal growth and were slightly raised on the plate (Fig. 1F). On MEA, colonies averaged 48.6 mm and displayed black aerial mycelia over a translucent vegetative mycelium; colonies were flat on the plate and showed a smooth margin (Fig. 1G). On OA, average colony diameter was 48.0 mm; overall shape was similar to that on PDA, but the concentric rings were less distinct and coloration tended toward olivaceous gray (Fig. 1H). Numerous irregular black perithecia were present, interconnected by ascomatal hairs curved into whorls or coils. Obovate asci were observed within the perithecia (Fig. 1I), releasing fusiform to ovoid ascospores that were initially hyaline but became pigmented brown at maturity (Fig. 1J). Ascospore size was (4.94–) 5.65 (–6.42) × (2.90–) 3.89 (–4.86) μm in diameter (n = 20).
Phylogenetic analysis. ITS sequence analysis showed 99.62% similarity to O. brasilensis strain CBS 130714 (MH865522.1), and the LSU sequence showed 99.51% similarity to O. brasilensis strain CBS 130714 (MH876958.1). In the concatenated ITS+LSU neighbor-joining tree, strain HARI25E097 formed a monophyletic clade with strains CBS 163.73 and CBS 586.83 (Fig. 2).
Specimen examined. Korea: Gangneung-si, Gangwon-do, 37°46′44.7″N, 128°55′34.8″E, June 29, 2025, isolated from healthy root of Solanum tuberosum, strain HARI25E097, KACC411095 (deposit No.), GenBank No. PX285304 (ITS), PX285305 (LSU).
Note. On OA, the colony size and color, concentric growth pattern, and mainly ovate ascospores were consistent with those of O. brasiliensis strain CBS 140.50 from the original description [13]. The morphology of the ascospores was similar to that of the closely related species Ovatospora senegalensis, but the ascospores of CBS 728.84 (type strain) had a noticeably pointed tip [13]; thus, they were clearly distinct from those of strain HARI25E097 in this study. The presence of ascomatal hairs forming connections between the perithecia and obovate asci in our isolate was also consistent with strain CBS 140.50 [13], supporting its identification.
Epicoccum mackenziei Jayasiri, Camporesi & K.D. Hyde, Mycosphere 8 (8): 1093 (2017) [MB#552362]
The strain was isolated from a potato leaf collected from Gangneung-si. After 7 days on PDA at 25°C, colonies averaged 60.1 mm in diameter and were light yellow-brown, slightly raised with a smooth margin on the plate (Fig. 3A). On MEA, colonies averaged 63.6 mm, exhibited sparser mycelia than those on PDA; the colonies were light brown and flat on the plate. It showed an erose margin due to irregular radial mycelial growth (Fig. 3B). On OA, colonies averaged 62.7 mm, appeared bright white with dense central aerial mycelia giving an umbonate appearance in the center; the margin was irregular (Fig. 3C). Asexual chlamydospores were hyaline, ellipsoidal to obovate, and formed hyphae in chains of 2–4 (Fig. 3D & 3E). The size was (8.16–) 10.85 (–14.76) × (6.18–) 8.31 (–11.92) μm in diameter (n = 20).
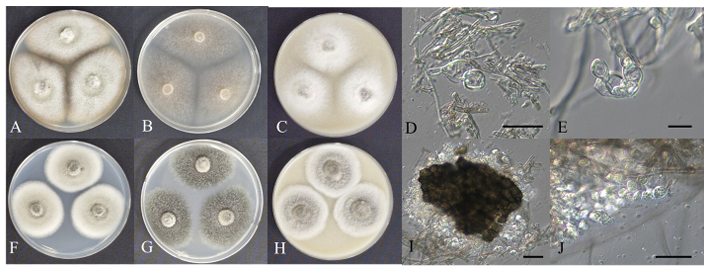
Fig. 3. Cultural characteristics of endophytic fungal strains belonging to Didymellaceae. Colonies of Epicoccum mackenziei HARI25D082 grown for 7 days on PDA (A), MEA (B), and OA (C) and chlamydospore (D, E); Colonies of Neoascochyta desmazieri HARI25E015 grown for 7 days on PDA (F), MEA (G), and OA (H) and pycnidial cell and pycnidia (I, J) (scale bars: D = 20 μm; E, I = 10 μm; J = 5 μm). PDA: potato dextrose agar; MEA: malt extract agar; OA: oatmeal agar.
Phylogenetic analysis. ITS, LSU, and RPB2 sequences showed 100% (strain KY45, OP550314.1), 99.62% (strain KY45, OP563622.1), and 98.88% (strain KY45, OP594302.1) similarity to E. mackenziei, respectively. In the concatenated ITS+LSU+RPB2 neighbor-joining tree, strain HARI25D082 formed a monophyletic clade with E. mackenziei strain KY45 [14] (Fig. 4).
Specimen examined. Korea: Gangneung-si, Gangwon-do, 37°50′02.5″N, 128°52′11.3″E, May 29, 2025, isolated from healthy leaf of Solanum tuberosum, strain HARI25D082, KACC411093 (deposit No.), GenBank No. PX285299 (ITS), PX285300 (LSU), PX289874 (RPB2).
Note. In the original description of E. mackenziei, ascospores were reported at the sexual stage, whereas asexual conidia were not observed and only chlamydospores were noted [15]. In this study, the sexual stage was not observed and only chlamydospores were confirmed, mirroring the original description. Phylogenetic analysis revealed that it was closely related to Epicoccum italicum (Fig. 4). However, while E. italicum clearly formed clavate-shaped conidiomata on MEA [16], no such morphology was observed in strain HARI25D082. Therefore, this strain was identified as E. mackenziei based on its chlamydospore morphology (spores in chain, size, and irregular hyaline shape) and molecular biology.
Neoascochyta desmazieri (Cavara) Qian Chen & L. Cai, Stud. Mycol. 82: 198 (2015) [MB#814142]
This strain was isolated from a potato stem collected from Jeju-si. After 7 days at 25°C, colonies on PDA averaged 49.5 mm in diameter and were grayish green with a smooth margin. The colonies were slightly raised on the plate (Fig. 3F). On MEA, colonies averaged 57.2 mm and were predominantly dark olivaceous with a lighter gray center; the margin was undulate, and the colony was raised (Fig. 3G). On OA, colonies averaged 56.7 mm, appearing white overall with light gray-green central aerial mycelia; the margin was smooth, and the colony was convex (Fig. 3H). Dark brown, oval pycnidia containing dozens to hundreds of pycniospores released upon pycnidial opening (Fig. 3I). Pycniospores were hyaline, colorless, and cylindrical (Fig. 3J); size was (1.84–) 2.17 (–2.67) × (3.09–) 4.16 (–5.96) μm in diameter (n = 20).
Phylogenetic analysis. The ITS sequences showed 99.59% similarity to N. desmazieri strain CBS 297.69 (NR_136130), the LSU sequence showed 100% similarity to N. desmazieri strain CBS 297.69 (NG_069315), and the RPB2 sequence showed 99.71% similarity to N. desmazieri strain CBS 297.69 (NG_069315). In the concatenated ITS+LSU+RPB2 neighbor-joining tree, strain HARI25E015 formed a monophyletic clade with N. desmazieri strain CBS 247.79 and CBS 297.69 (Fig. 4).
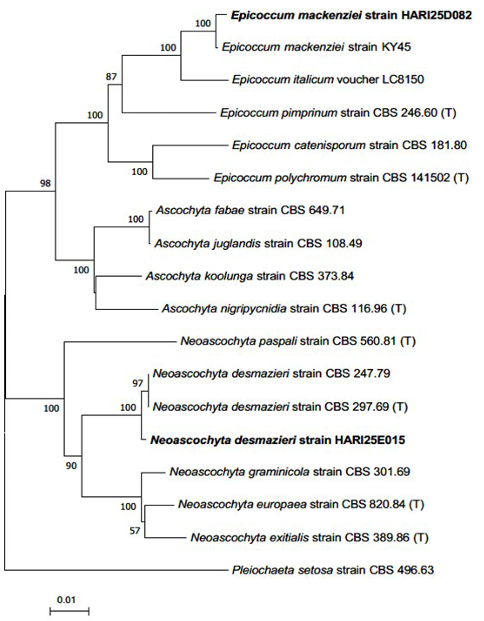
Fig. 4. Neighbor-joining phylogenetic tree of E. mackenziei and N. desmazieri on the basis of a concatenated alignment of internal transcribed spacer (ITS), large subunit (LSU) rDNA, and RNA polymerase II second-largest subunit (RPB2) DNA sequences. Pleiochaeta setosa was used as an outgroup. Numbers on branches indicate bootstrap values (1,000 replicates). Fungal strains isolated in this study are in bold. (T) indicates the type strains.
Specimen examined. Korea: Jeju-si, Jeju-do, 33°28′43.9″N, 126°47′02.0″E, May 19, 2025, isolated from healthy stem of Solanum tuberosum, strain HARI25E015, KACC411094 (deposit No.), GenBank No. PX285301 (ITS), PX285303 (LSU), PX289875 (RPB2).
Note. Phylogenetic analysis revealed a clear distinction between Neoascochyta and Ascochyta clades (Fig. 4). Although conidia were not observed in our material, the presence of ellipsoidal pycnidia and pycniospores, and molecular data support the identification of N. desmazieri [17].
Species belonging to the family Chaetomiaceae are commonly found in diverse environments, such as soil, indoor habitats, plant saprophytes, and endophytes [18,19]. These species, owing to their ability to produce unique enzymes and secondary metabolites, are widely used in biotechnology and agriculture [20–22]. They are typified by pigmented ascospores in fasciculate, evanescent asci [23,24], and, in many species, only the sexual stage is commonly observed [25,26].
Didymellaceae is one of the largest families within Dothideomycetes and includes the genera Ascochyta, Didymella, Epicoccum, and Phoma [27]. Species that belong to the family Didymellaceae have traditionally been recognized as plant pathogens that affect a broad host range, causing leaf and stem diseases [16]; however, some species have recently been reported as endophytes [4,28].
Collariella bostrychodes is the type species of the genus Collariella. Previously placed in Chaetomium, C. bostrychodes was transferred to Collariella on the basis of its ascomatal morphology; the genus name refers to the collar-like arrangement of developing asci [13]. Although originally described as an indoor airborne fungus [13], C. bostrychodes has since been reported as an endophyte in woolly plectranthus (Coleus forskohlii) [29] and Mongolian pine (Pinus sylvestris var. mongolica) [30]. The antagonistic activity of this species against Rigidoporus microporus, the causative agent of white root disease in rubber trees (Ficus elastica), has also been reported [31].
Ovatospora brasiliensis, the type species of Ovatospora, was formerly included in Chaetomium and later segregated on the basis of its ovate ascospore morphology [13]. Although originally isolated as a saprophyte from jute cloth [13], O. brasiliensis has been recovered as an endophyte from turmeric (Curcuma caesia) and has been implicated in the bioconversion of curcumin into calebin-A [32]. There has also been a report of the strong anti-inflammatory activity of O. brasiliensis isolated as an endophyte from moss [33].
Epicoccum mackenziei was first reported as a saprophytic fungus from dead branches of restharrow (Ononis spinosa) [15]; it has also been isolated from tea tree leaves (Camellia sinensis) as both a pathogen [14] and an endophyte [34].
Neoascochyta desmazieri was originally assigned to Ascochyta but was transferred to Neoascochyta on the basis of molecular evidence [17]. This species was first described as a Poaceae pathogen (e.g., Lolium perenne) [33]; however, this species was isolated from healthy potato stems, so we considered it an endophyte of potatoes.
In this study, endophytic fungal strains were isolated from healthy potato plant tissues collected from spring potato cultivation areas in Korea, and four endophytic fungal species previously unrecorded in Korea were identified. Several of the identified species have been reported to produce secondary metabolites and exhibit anti-fungal or anti-inflammatory activities, suggesting their potential utility in biological disease control. Given concerns regarding pesticide residues and the development of fungicide resistance, evaluation of endophytic fungi as biological control agents is increasingly important. The strains described here warrant further study to evaluate their antagonistic activity against potato pathogens and their suitability for development as biocontrol agents.
No potential conflict of interest relevant to this article was reported.
This study was supported by the Rural Development Administration (grant number: PJ017574032025).
1. Lee GB, Choi JG, Kwon DH, Yi JY, Lee HT, Jin YI. Effect of spring potato cultivation period on growth, yield and processing quality of autumn potato cultivars. Korean J Plant Res 2024;37:149-60. https://doi.org/10.7732/kjpr.2024.37.2.149
[DOI]
2. Korean Statistical Information Service. Potatoes production (Field) [Internet]. Daejeon: KOSIS; 2023 [cited 2025 Oct 13]. Available from: https://kosis.kr
3. Gao FK, Dai CC, Liu XZ. Mechanisms of fungal endophytes in plant protection against pathogens. Afr J Microbiol Res 2010;4:1346-51.
4. Park H, Kim J, Kim S. Unreported endophytic fungi isolated from potato (Solanum tuberosum L.) leaves in Korea: Didymella sinensis and Sagenomella oligospora. Kor J Mycol 2024;52:257-63. https://doi.org/10.4489/kjm.520404
[DOI]
5. El-Hasan A, Ngatia G, Link TI, Voegele RT. Isolation, identification, and biocontrol potential of root fungal endophytes associated with solanaceous plants against potato late blight (Phytophthora infestans). Plants 2022;11:1605. https://doi.org/10.3390/plants11121605
[DOI]
6. Zhang L, Xu W, Zhao Z, Long Y, Fan R. Biocontrol potential and growth-promoting effect of endophytic fungus Talaromyces muroii SD1-4 against potato leaf spot disease caused by Alternaria alternata. BMC Microbiol 2024;24:255. https://doi.org/10.1186/s12866-024-03411-4
[DOI]
7. Park H, Jung CR, Eom AH. Species diversity and antifungal activity of endophytic fungi isolated from Angelica gigas Nakai. Kor J Mycol 2021;49:497-505. https://doi.org/10.4489/KJM.20210048
[DOI]
8. Kim JH, Kim DY, Park H, Cho JH, Eom AH. Neocosmospora rubicola, an unrecorded endophytic fungus isolated from roots of Glycyrrhiza uralensis in Korea. Kor J Mycol 2017;45:63-7. https://doi.org/10.4489/KJM.20170007
[DOI]
9. White TJ, Bruns TD, Lee SB, Taylor JW. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, editors. PCR protocols: A guide to methods and applications. New York: Academic Press; 1990. p. 315-22.
https://doi.org/10.1016/B978-0-12-372180-8.50042-1
[DOI]
10. Vilgalys R, Hester M. Rapid genetic identification and mapping of enzymatically amplified ribosomal DNA from several Cryptococcus species. J Bacteriol 1990;172:4238-46. https://doi.org/10.1128/jb.172.8.4238-4246.1990
[DOI]
11. Liu YJ, Whelen S, Hall BD. Phylogenetic relationships among ascomycetes: Evidence from an RNA polymerse II subunit. Mol Biol Evol 1999;16:1799-808. https://doi.org/10.1093/oxfordjournals.molbev.a026092
[DOI]
12. Kumar S, Stecher G, Tamura K. MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 2016;33:1870-4. https://doi.org/10.1093/molbev/msw054
[DOI]
13. Wang XW, Houbraken J, Groenewald JZ, Meijer M, Andersen B, Nielsen KF, Crous PW, Samson RA. Diversity and taxonomy of Chaetomium and chaetomium-like fungi from indoor environments. Stud Mycol 2016;84:145-224. https://doi.org/10.1016/j.simyco.2016.11.005
[DOI]
14. Zhang J, Yin QX, Jiang SL, Li DX, Tang Q, Zhang Y, Ma YL, Li PY, Wang Y, Wang DL, et al. First report of Epicoccum mackenziei causing dark brown spot of tea leaf in Guizhou province, southwestern China. Plant Dis 2023;107:2247. https://doi.org/10.1094/PDIS-10-22-2372-PDN
[DOI]
15. Jayasiri SC, Hyde KD, Jones EBG, Jeewon R, Ariyawansa HA, Bhat JD, Camporesi E, Kang JC. Taxonomy and multigene phylogenetic evaluation of novel species in Boeremia and Epicoccum with new records of Ascochyta and Didymella (Didymellaceae). Mycosphere 2017;8:1080-101. https://doi.org/10.5943/mycosphere/8/8/9
[DOI]
16. Chen Q, Hou LW, Duan WJ, Crous PW, Cai L. Didymellaceae revisited. Stud Mycol 2017;87:105-59. https://doi.org/10.1016/j.simyco.2017.06.002
[DOI]
17. Chen Q, Jiang JR, Zhang GZ, Cai L, Crous PW. Resolving the Phoma enigma. Stud Mycol 2015;82:137-217. https://doi.org/10.1016/j.simyco.2015.10.003
[DOI]
18. Betancourt O, Zaror L, Senn C. Isolation of filamentous fungi from haircoat cats without skin lesions in Temuco, Chile. Revista Cientifica, Facultad de Ciencias Veterinarias, Universidad del Zulia 2013;23:380-7.
19. Wang XW, Lombard L, Groenewald JZ, Li J, Videira SIR, Samson RA, Liu XZ, Crous PW. Phylogenetic reassessment of the Chaetomium globosum species complex. Persoonia 2016;36:83-133. https://doi.org/10.3767/003158516X689657
[DOI]
20. Wang XW, Han PJ, Bai FY, Luo A, Bensch K, Meijer M, Kraak B, Han DY, Sun BD, Crous PW, et al. Taxonomy, phylogeny and identification of Chaetomiaceae with emphasis on thermophilic species. Stud Mycol 2022;101:121-243. https://doi.org/10.3114/sim.2022.101.03
[DOI]
21. Yang CH, Lin MJ, Su HJ, Ko WH. Multiple resistance-activating substances produced by Humicola phialophoroides isolated from soil for control of Phytophthora blight of pepper. Bot Stud 2014;55:40. https://doi.org/10.1186/1999-3110-55-40
[DOI]
22. van den Brink J, Samson RA, Hagen F, Boekhout T, de Vries RP. Phylogeny of the industrial relevant, thermophilic genera Myceliophthora and Corynascus. Fungal Divers 2012;52:197-207. https://doi.org/10.1007/s13225-011-0107-z
[DOI]
23. Hawksworth DL. A revision of genus Ascotricha Berk. Mycol Pap 1971;126:1-28.
24. Whiteside WC. Morphological studies in the Chaetomiaceae. I. Mycologia 1961;53:512-23.
https://doi.org/10.1080/00275514.1961.12017979
[DOI]
25. Von Arx JA, Dreyfuss M, Müller E. A revaluation of Chaetomium and the Chaetomiaceae. Persoonia 1984;12:169-79.
26. von Arx JA, Guarro J, Figueras MJ. The ascomycete genus Chaetonium. Nova Hedwigia Beih 1986;84:1-162.
27. de Gruyter J, Aveskamp MM, Woudenberg JHC, Verkley GJM, Groenewald JZ, Crous PW. Molecular phylogeny of Phoma and allied anamorph genera: Towards a reclassification of the Phoma complex. Mycol Res 2009;113:508-19. https://doi.org/10.1016/j.mycres.2009.01.002
[DOI]
28. Tan Y, Wang YD, Li Q, Xing XK, Niu SB, Sun BD, Chen L, Pan RL, Ding G. Undescribed diphenyl ethers betaethrins A-I from a desert plant endophytic strain of the fungus Phoma betae A.B. Frank (Didymellaceae). Phytochemistry 2022;201:113264. https://doi.org/10.1016/j.phytochem.2022.113264
[DOI]
29. Crasta GL, Anandrao RK. Seasonal diversity & spaciotemporal distribution of fungal endophytes associated with the medicinal plant Coleus forskohlii Briq. Plant Sci Today 2024;11:223-33. https://doi.org/10.14719/pst.2729
[DOI]
30. Ren N, Wang L, You C. Diversity, community structure, and antagonism of endophytic fungi from asymptomatic and symptomatic Mongolian pine trees. J Fungi 2024;10:212. https://doi.org/10.3390/jof10030212
[DOI]
31. Soytong K, Kahonokmedhakul S, Song J, Tongon R. Chaetomium application in agriculture. In: Ahmad F, Sultan M, editors. Technology in agriculture. London: IntechOpen; 2021. https://doi.org/10.5772/intechopen.99402
[DOI]
32. Majeed A, Majeed M, Thajuddin N, Arumugam S, Ali F, Beede K, Adams SJ, Gnanamani M. Bioconversion of curcumin into calebin-A by the endophytic fungus Ovatospora brasiliensis EPE-10 MTCC 25236 associated with Curcuma caesia. AMB Express 2019;9:79. https://doi.org/10.1186/s13568-019-0802-9
[DOI]
33. Yang J, Sun Y, Li M, Yu Q. Anti-inflammatory potential of extracellular polysaccharide from the moss endophyte Ovatospora brasiliensis during pathogen infection. Microorganisms 2025;13:2037. https://doi.org/10.3390/microorganisms13092037
[DOI]
34. Wang Y, Tu Y, Chen X, Jiang H, Ren H, Lu Q, Wei C, Lv W. Didymellaceae species associated with tea plant (Camellia sinensis) in China. MycoKeys 2024;105:217-51. https://doi.org/10.3897/mycokeys.105.119536
[DOI]