Seong-Eun Kim1, Ju-Kyeong Eo2, and Ahn-Heum Eom1*
1Department of Biology Education, Korea National University of Education, Cheongju 28173, Korea
2National Ecosystem Survey Team, Division of Ecological Survey, National Institute of Ecology, Seocheon-gun 33657, Korea
*Correspondence to eomah@knue.ac.kr
Korean Journal of Mycology (Kor J Mycol) 2025 December, Volume 53, Issue 4, pages 237-243.
https://doi.org/10.4489/kjm.2025.53.4.1
Received on October 17, 2025, Revised on November 10, 2025, Accepted on November 10, 2025, Published on December 31, 2025.
Copyright © The Korean Society of Mycology.
This is an Open Access article which is freely available under the Creative Commons Attribution-Non-Commercial 4.0 International License (CC BY-NC) (https://creativecommons.org/licenses/by-nc/4.0/).
In this study, we characterized two endophytic fungal species, respectively isolated from the roots of Paeonia lactiflora and twigs of Gleditsia triacanthos in Korea, based on morphological and molecular characteristics. Following surface sterilization, fungal strains were isolated, cultured, and examined microscopically for morphological characteristics. Molecular identification was conducted using PCR amplification and sequencing of four loci: the internal transcribed spacer (ITS), large subunit (LSU) rDNA, β-tubulin (TUB), and RNA polymerase II second largest subunit (RPB2) regions. On the basis of these morphological and phylogenetic analyses, the isolates were identified as Pleiocarpon strelitziae and Neodidymelliopsis tinkyukuku, which are newly recorded species in Korea. The identification of these unrecorded species augments current knowledge regarding the fungal diversity in Korea and provides useful information for future studies on endophytic fungi and their ecological roles.
Endophyte, Fungi, Korea, Morphology, New record, Phylogeny
Endophytic fungi are important components of natural ecosystems that occur ubiquitously in almost all known plant species, encompassing a wide range of hosts growing in diverse habitats [1]. These fungi inhabit internal plant tissues without causing visible disease symptoms, often providing multiple benefits to their hosts, whilst simultaneously deriving advantages from these associations [2]. Moreover, bioactive metabolites produced by endophytic fungi have been reported to play significant roles in mediating plantfungal interactions [3].
Paeonia lactiflora Pall., a traditional medicinal and ornamental plant, has been established to harbor diverse endophytic fungi, some of which are known to produce the secondary metabolite paeoniflorin [4,5]. The genus Gleditsia (Fabaceae), comprises species of leguminous trees distributed in East Asia and North America, from which several bioactive compounds with therapeutic properties have been isolated [6,7].
During a survey of endophytic fungi associated with native plants in Korea, we obtained two isolates from the plants P. lactif lora and Gleditsia triacanthos. On the basis of an examination of morphological characteristics and molecular phylogenetic analyses, we identified these fungi as Pleiocarpon strelitziae and Neodidymelliopsis tinkyukuku, respectively. Pleiocarpon (Nectriaceae, Hypocreales) is a small genus comprising plant-associated or soil-borne fungi, some of which are phytopathogenic on Strelitzia [8]. Neodidymelliopsis (Didymellaceae, Pleosporales) includes species commonly occurring as endophytes or weak pathogens on a diverse range of herbaceous and woody hosts [9].
In this study, we describe the detailed morphological and phylogenetic characterization of these two endophytic fungi, which are newly recorded from Korea.
Samples were collected from the roots of Paeonia lactiflora in Yeongju, Gyeongsangbuk-do (36.88055°N, 128.53939°E), in July 2020, and from the twigs of Gleditsia triacanthos in Jobal-do, Yeosu-si, Jeollanamdo (34.632309°N, 127.56011°E), in May 2021. The samples were sealed in zipper bags and transported to the laboratory within 24 h. Upon arrival, the collected materials were thoroughly rinsed under running tap water, cut into 1.0–1.5 cm segments, and surface sterilized by immersing in 1% sodium hypochlorite (NaOCl) solution for 1 min, followed by washing with 70% ethanol for 30 s. The sterilized segments were subsequently placed on potato dextrose agar (PDA: Difco Laboratories, Detroit, MI, USA).
Fungal mycelia growing within plant tissues were observed after incubation at 25°C under dark conditions for 7 d. The observed mycelia were subsequently sub-cultured on PDA medium to obtain pure isolates. To determine colony characteristics, including diameter, color, morphology, elevation, and texture, the pure cultures were examined macroscopically using an Axio Imager A1 light microscope (Carl Zeiss, Oberkochen, Germany). To induce sporulation, some isolates were co-cultured with sterilized pine needles on 2% water agar.
DNA was extracted from mycelia of the isolated strains using a HiGene Genomic DNA Prep Kit (BioFACT, Daejeon, Korea), following the manufacturer’s instructions. Polymerase chain reaction (PCR) was performed to amplify the internal transcribed spacer (ITS), large subunit rDNA (LSU), β-tubulin (TUB), RNA polymerase II second largest subunit (RPB2) regions. The PCR conditions for ITS, LSU, and TUB were as follows: an initial denaturation at 95°C for 2 min; followed by 35 cycles of denaturation at 95°C for 20s, annealing at 50°C (ITS [10]), 44°C (LSU [11]), or 55°C (TUB [12]) for 40s, and extension at 72°C for 1 min; with a final extension at 72°C for 10 min. The PCR program used for RPB2 [13] comprised an initial denaturation at 95°C for 5 min; followed by 35 cycles of denaturation at 95°C for 1 min, annealing at 52°C for 2 min, and extension at 72°C for 90s; with a final extension at 72°C for 10 min. The amplified PCR products were verified by agarose gel electrophoresis and subsequently subjected to Sanger sequencing performed by Solgent Co., Ltd (Daejeon, Korea). The DNA sequences obtained were analyzed using the Basic Local Alignment Search Tool (BLAST) provided by the National Center for Biotechnology Information (NCBI) to search for homologous sequences and to identify isolates at the species level. Multiple sequence alignments were performed using MEGA11 software [14], and phylogenetic trees were constructed using the maximum likelihood method. The robustness of the phylogenetic trees was assessed by performing 1,000 bootstrap replicates. Specimens of the newly identified and previously unrecorded fungal species have been deposited at the National Institute of Biological Resources (NIBR), and the DNA sequences used for phylogenetic analysis have been submitted to NCBI.
To determine their phylogenetic relationships, sequence analyses were performed for the two isolates KNUE 20A186 and KNUE 21E091. On the basis of analyses of the ITS, LSU, and TUB regions, the KNUE 20A186 strain was identified as Pleiocarpon strelitziae, which showed 100% (ITS), 99.6% (LSU), and 99.7% (TUB) sequence identity with strain CBS 142251. The maximum-likelihood tree based on the combined ITSLSU–TUB sequences placed the isolate in a well-supported clade (bootstrap support = 99%) with the type strain P. strelitziae CBS 142251, confirming its taxonomic identity (Fig. 1). On the basis of sequence analysis of the ITS and RPB2 regions, the KNUE 21E091 strain was identified as Neodidymelliopsis tinkyukuku, showing 99.6% (ITS) and 98.7% (RPB2) similarity to the type strain BRIP 69592. Within the maximumlikelihood phylogenetic trees, strain KNUE 20A186 clustered with P. strelitziae CBS 142251 (Fig. 1), whereas KNUE 21E091 formed a well-supported clade with N. tinkyukuku BRIP 69592 (Fig. 2), confirming their respective identifications. In the maximum-likelihood tree constructed from the concatenated ITS and RPB2 sequences, KNUE 21E091 grouped with N. tinkyukuku BRIP 69592 in a distinct and well-supported clade (bootstrap support = 92%) (Fig. 2).
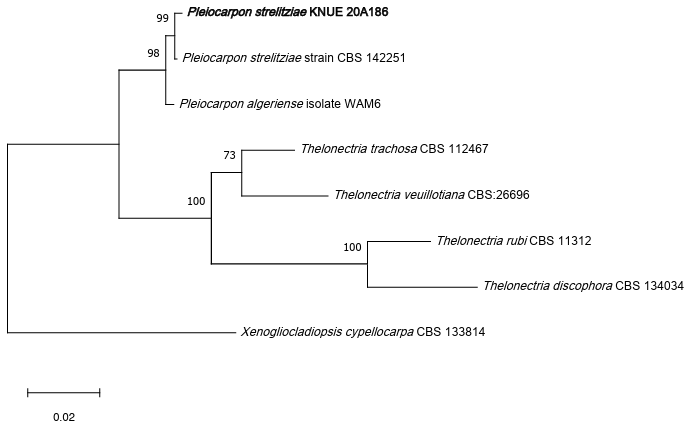
Fig. 1. A maximum likelihood phylogenetic tree (TN92+G) for Pleiocarpon strelitziae KNUE 20A186 based on a concatenated alignment of internal transcribed spacer (ITS), large subunit (LSU), and β-tubulin (TUB) sequences, with Xenogliocladiopsis cypellocarpa used as an outgroup. The numbers on branches indicate bootstrap values (1,000 replicates) greater than 50%. The fungal strain isolated in this study is shown in bold type.
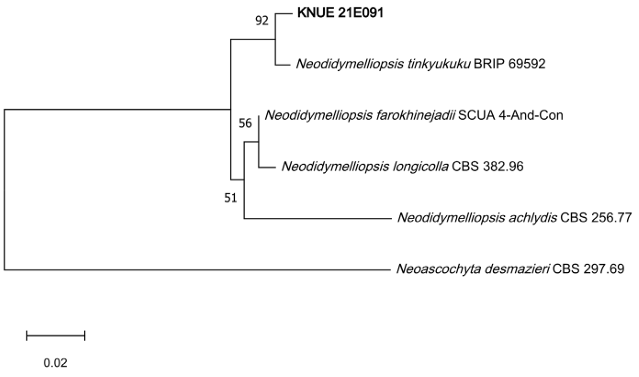
Fig. 2. A maximum likelihood phylogenetic tree (K2+G) for Neodidymelliopsis tinkyukuku KNUE 21E091 based on a concatenated alignment of internal transcribed spacer (ITS) and RNA polymerase II second largest subunit (RPB2) gene sequences, with Neoascochyta desmazieri used as an outgroup. The numbers on branches indicate bootstrap values (1,000 replicates) greater than 50%. The fungal strain isolated in this study is shown in bold type.
Pleiocarpon strelitziae L. Lombard & D. Aiello, IMA Fungus 8 (1): 73 (2017) [MB#820029] Fig. 3, Table 1
Morphological characteristics of KNUE 20A186: When grown in darkness for 7 days at 25°C, colonies on PDA reached 25–30 mm in diameter, consisting predominantly of light greenish-yellow aerial mycelia with a thin margin of white hyphae. The reverse surface was brilliant orange yellow. On malt extract agar (MEA) under the same conditions, colonies reached 34–39 mm in diameter, the central area of which was brilliant greenish yellow, gradually transitioning to light greenish-yellow aerial mycelia toward the margins. The reverse surface was similarly brilliant orange yellow. Colonies on both PDA and MEA were characterized by a cottony texture, growing in concentric rings. The conidiophores were erect, smoothwalled, and hyaline. Macro-conidia, which were boomerang-shaped and curved in a single direction, measured (33.9–) 40.5 (–48.2) × (4.5–) 5.7 (–7.4) μm (n = 20), were hyaline, septate with 1–5 septa, and scattered irregularly, whereas micro-conidia were lemon-shaped, smooth-walled, and measured (3.8–) 5.0 (–6.6) × (2.1–) 2.8 (–3.4) μm (n = 20).
Specimen examined: Yeongju-si, Gyeongsangbuk-do, Korea, 36.88055°N, 128.53939°E, July 7, 2020, isolated from roots of Paeonia lactiflora Pall., strain KNUE 20A186, NIBRFGC000508513, GenBank No. PX220004 (ITS), PX220007 (LSU), PX236115 (TUB).
Notes: Pleiocarpon strelitziae was first described from a botanical garden in eastern Sicily, Italy, as a pathogen causing dry basal stem rot on Strelitzia reginae Ait. Although the sexual morph has not been observed, the asexual morph has been reported to be Cylindrocarpon-like. The macro-conidia have been described as almost cylindrical, hyaline, straight to curved, and with 0–5 septa [8], which is consistent with the characteristics observed in P. strelitziae KNUE 20A186.
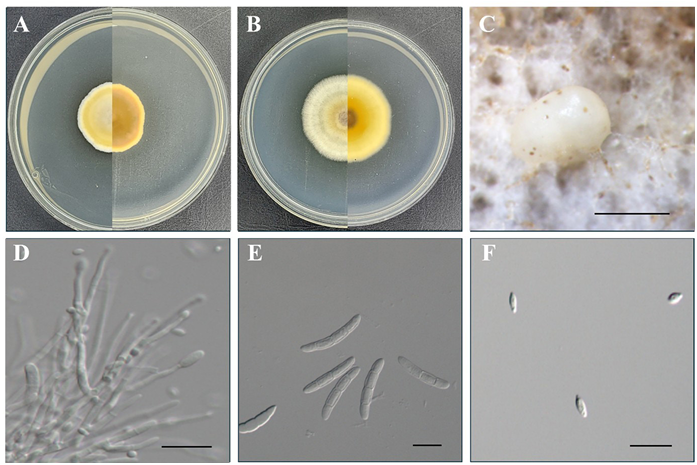
Fig. 3. Morphological characteristics of Pleiocarpon strelitziae KNUE 20A186. Colonies grown for 7 days on potato dextrose agar (A) and malt extract agar (B). Conidiomata (C). Conidiophore (D). Macroconidia (E). Micro-conidia (F). Scale bars: 1 mm for C, and 20 μm for D, E, and F.
Table 1. Morphological characteristics of Pleiocarpon strelitziae KNUE 20A186
table
PDA: potato dextrose agar; MEA: malt extract agar.
Neodidymelliopsis tinkyukuku E.C. Keirnan, M.H. Laurence, R.G. Shivas & Y.P. Tan, MycoKeys 78: 11 (2021) [MB#833692] Fig. 4, Table 2
Morphological characteristics of KNUE 21E091: When grown on PDA at 25°C for 7 days, colonies reached 46–49 mm in diameter, characterized by a white margin with a dark brown center and lacking pigmentation. Mycelium expanded radially at a consistent rate, forming circular colonies of uniform density that grew parallel to the agar surface, resulting in an even, velvety texture comprising densely packed short hyphae. The reverse surface has a morphology similar to that of the upper surface. On MEA, colonies reached 50–51 mm in diameter, with an opaque white margin and a yellowish-brown center, maintaining the same growth pattern, texture, and reverse morphology as observed on PDA. Conidia were cylindrical, hyaline with light yellow, measuring (4.37–) 5.06 (–5.96) × (2.29–) 2.73 (–3.12) μm (n = 20).
Specimen examined: Jobaldo island, Yeosu-si, Jeollanam-do, Korea, 34.632309°N, 127.56011°E, May 6, 2021, isolated from twigs of Gleditsia triacanthos, strain KNUE 21E091, NIBRFGC000509079, GenBank No. PX459707 (ITS), LC902017(RPB2).
Notes: N. tinkyukuku was first reported from the leaves of Hardenbergia violacea (Schneev.) Stearn collected in Clare, South Australia. Fine structural features provide a more reliable basis of determining similarity than external colony morphology [15].
In this study, the two species P. strelitziae and N. tinkyukuku, which were previously unrecorded in Korea, were isolated from the roots of Paeonia lactiflora and twigs of Gleditsia triacanthos, respectively, and were characterized based on morphological observations and molecular phylogenetic analyses. N. tinkyukuku was originally reported as a pathogenic fungus causing disease on the leguminous plant H. violacea, although in the present study, it was isolated from surface-sterilized tissues from a different leguminous host, indicating that this fungus can also establish associations as an endophytic symbiont. Such cases, in which endophytic fungi have both symbiotic and pathogenic behaviors, tend to be relatively common given that the interactions between endophytes and their host plants range along a continuum from mutualism to pathogenicity [16]. For example, depending on environmental and host conditions, Colletotrichum gloeosporioides and Fusarium oxysporum are known to function as either endophytes or pathogens, and Chaetomium globosum can also occur as an endophyte, saprophyte, or pathogen [17–19]. Such examples accordingly illustrate the ecological diversity and functional versatility of plant endophytic fungi.
Although research on endophytic fungi in Korea has gradually been increasing in recent years, further taxonomic and ecological studies are required to gain a more comprehensive understanding of their diversity [20]. Consequently, the discovery of previously unrecorded fungi in Korea highlights the necessity of continuous investigation of indigenous microorganisms, the study of which has potentially important implications for plant health, ecosystem functioning, and potential biotechnological applications.
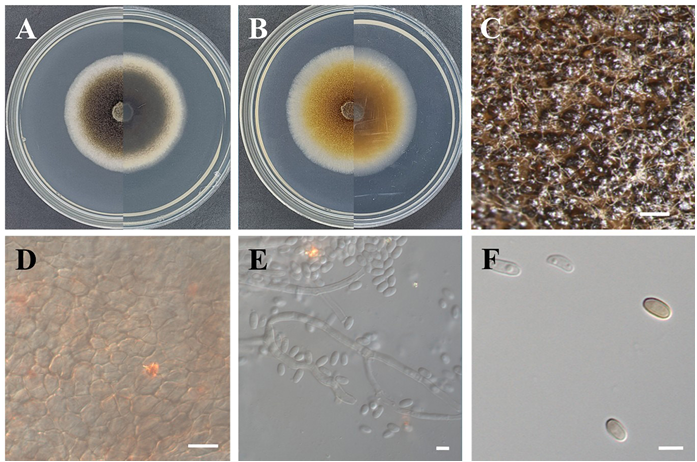
Fig. 4. Morphological characteristics of Neodidymelliopsis tinkyukuku KNUE 21E091. Colonies grown for 7 days on potato dextrose agar (A) and malt extract agar (B). Conidiomata (C). Pycnidial wall (D). Conidiophore (E). Conidia (F). Scale bars: 0.5 mm for C, 10 μm for D, 5 μm for E, and 10 μm for F.
Table 2. Morphological characteristics of Neodidymelliopsis tinkyukuku KNUE 21E091
table
PDA: potato dextrose agar; MEA: malt extract agar; OA: oatmeal agar.
The authors declare that they have no competing interests.
This work was supported by a grant from the National Institute of Biological Resources (NIBR202502103) and National Institute of Ecology (NIE-A-2025-01), funded by the Ministry of Environment (MOE) of the Republic of Korea.
1. Sun X, Guo LD. Endophytic fungal diversity: review of traditional and molecular techniques. Mycology 2012;3:65-76. https://doi.org/10.1080/21501203.2012.656724
[DOI]
2. Baron NC, Rigobelo EC. Endophytic fungi: A tool for plant growth promotion and sustainable agriculture. Mycology 2022;13:39-55. https://doi.org/10.1080/21501203.2021.1945699
[DOI]
3. Schulz B, Boyle C, Draeger S, Römmert AK, Krohn K. Endophytic fungi: A source of novel biologically active secondary metabolites. Mycol Res 2002;106:996-1004. https://doi.org/10.1017/S0953756202006342
[DOI]
4. He DY, Dai SM. Anti-inflammatory and immunomodulatory effects of paeonia lactiflora Pall., a traditional Chinese herbal medicine. Front Pharmacol 2011;2:10. https://doi.org/10.3389/fphar.2011.00010
[DOI]
5. Cheng X, Wei Z, Pu S, Xiang M, Yan A, Zhang Y, Wang X. Diversity of endophytic fungi of Paeonia lactiflora Pallas and screening for fungal paeoniflorin producers. FEMS Microbiol Lett 2018;365:fny263. https://doi.org/10.1093/femsle/fny263
[DOI]
6. Ashraf H, Moussa AY, Eldahshan OA, Singab ANB. Genus Gleditsia: A phytochemical and biological review (2015-2020). J Biol Act Prod Nat 2022;12:1-23. https://doi.org/10.1080/22311866.2021.2013943
[DOI]
7. Zhang JP, Tian XH, Yang YX, Liu QX, Wang Q, Chen LP, Li HL, Zhang WD. Gleditsia species: An ethnomedical, phytochemical and pharmacological review. J Ethnopharmacol 2016;178:155-71. https://doi.org/10.1016/j.jep.2015.11.044
[DOI]
8. Aiello D, Polizzi G, Crous PW, Lombard L. Pleiocarpon gen. nov. and a new species of Ilyonectria causing basal rot of Strelitzia reginae in Italy. IMA Fungus 2017;8:65-76. https://doi.org/10.5598/imafungus.2017.08.01.05
[DOI]
9. Qian Z, Zhang S, Manawasinghe IS, Sun D, Song J, Xu B. Taxonomic revision of Neodidymelliopsis with N. cynanchi sp. nov., associated with Cynanchum sibiricum leaf spot in Xinjiang, China. N Z J Bot 2024;62:151-64. https://doi.org/10.1080/0028825X.2023.2267807
[DOI]
10. Gardes M, Bruns TD. ITS primers with enhanced specificity for basidiomycetes-application to the identification of mycorrhizae and rusts. Mol Ecol 1993;2:113-8. https://doi.org/10.1111/j.1365-294X.1993.tb00005.x
[DOI]
11. Moncalvo JM, Lutzoni FM, Rehner SA, Johnson J, Vilgalys R. Phylogenetic relationships of agaric fungi based on nuclear large subunit ribosomal DNA sequences. Syst Biol 2000;49:278-305. https://doi.org/10.1093/sysbio/49.2.278
[DOI]
12. Glass NL, Donaldson GC. Development of primer sets designed for use with the PCR to amplify conserved genes from filamentous ascomycetes. Appl Environ Microbiol 1995;61:1323-30. https://doi.org/10.1128/aem.61.4.1323-1330.1995
[DOI]
13. Liu YJ, Whelen S, Hall BD. Phylogenetic relationships among ascomycetes: Evidence from an RNA polymerse II subunit. Mol Biol Evol 1999;16:1799-808. https://doi.org/10.1093/oxfordjournals.molbev.a026092
[DOI]
14. Tamura K, Stecher G, Kumar S. MEGA11: Molecular evolutionary genetics analysis version 11. Mol Biol Evol 2021;38:3022-7. https://doi.org/10.1093/molbev/msab120
[DOI]
15. Keirnan EC, Tan YP, Laurence MH, Mertin AA, Liew ECY, Summerell BA, Shivas RG. Cryptic diversity found in Didymellaceae from Australian native legumes. MycoKeys 2021;78:1-20. https://doi.org/10.3897/mycokeys.78.60063
[DOI]
16. Hardoim PR, van Overbeek LS, Berg G, Pirttilä AM, Compant S, Campisano A, Döring M, Sessitsch A. The hidden world within plants: Ecological and evolutionary considerations for defining functioning of microbial endophytes. Microbiol Mol Biol Rev 2015;79:293-320. https://doi.org/10.1128/MMBR.00050-14
[DOI]
17. Edel-Hermann V, Lecomte C. Current status of Fusarium oxysporum formae speciales and races. Phytopathology 2019;109:512-30. https://doi.org/10.1094/PHYTO-08-18-0320-RVW
[DOI]
18. Photita W, Taylor PWJ, Ford R, Hyde KD, Lumyong S. Morphological and molecular characterization of Colletotrichum species from herbaceous plants in Thailand. Fungal Divers 2005;18:117-33.
19. Arnold AE. Understanding the diversity of foliar endophytic fungi: Progress, challenges, and frontiers. Fungal Biol Rev 2007;21:51-66. https://doi.org/10.1016/j.fbr.2007.05.003
[DOI]
20. Eo JK, Choi JW, Eom AH. Diversity, distribution, and host plant of endophytic fungi: A focus on Korea. Mycobiology 2022;50:399-407. https://doi.org/10.1080/12298093.2022.2154044
[DOI]