Ardhianita Indi Nur Aida1,2 , In-Young Choi1,2* , Kyoung-Mo Koo3 , Ji-Hyun Park3* , Seong-Jin Lee4 , and Hyeon-Dong Shin5
1Department of Plant Medicine, Jeonbuk National University, Jeonju 54896, Korea
2Research Center for Plant Medicine, Jeonbuk National University, Jeonju 54896, Korea
3Department of Forest Resources, Kookmin University, Seoul 02707, Korea
4Plant Quarantine Technology Center, Department of Plant Quarantine, Animal and Plant Quarantine Agency, Gimcheon, 39660, Korea
5Division of Environmental Science and Ecological Engineering, Korea University, Seoul 02841, Korea
*Correspondence to iychoi@jbnu.ac.kr, jhpark10@kookmin.ac.kr
Korean Journal of Mycology (Kor J Mycol) 2025 December, Volume 53, Issue 4, pages 237-243.
https://doi.org/10.4489/kjm.2025.53.4.1
Received on October 17, 2025, Revised on November 10, 2025, Accepted on November 10, 2025, Published on December 31, 2025.
Copyright © The Korean Society of Mycology.
This is an Open Access article which is freely available under the Creative Commons Attribution-Non-Commercial 4.0 International License (CC BY-NC) (https://creativecommons.org/licenses/by-nc/4.0/).
This study provides a detailed morphological description of Pseudocercospora securinegae, a species that has long been overlooked, supplemented by comprehensive illustrations of its diagnostic features. The taxonomic value of this study is further supported by molecular phylogenetic analyses based on a multigene dataset, including the internal transcribed spacer (ITS) region, actin (actA), translation elongation factor 1-alpha (tef1), and DNAdirected RNA polymerase II second largest subunit (rpb2). Notably, a sequence of the rpb2 gene for this fungus is presented here for the first time.
Arching bushweed, Leaf spot, Multi-locus DNA dataset, Phytopathogen, Securinega suffruticosa
Pseudocercospora securinegae (Togashi & Katsuki) Deighton is a fungal pathogen that causes leaf spot disease. It was originally described as Cercospora securinegae Togashi & Katsuki from Securinega suffruticosa (Pall.) Rehder (currently Flueggea suffruticosa (Pall.) Baill.) in Japan [1]. Its current taxonomic placement within the genus Pseudocercospora was established by Deighton in 1976 [2].
In Korea, the occurrence of Ps. securinegae on F. suffruticosa was first observed during a survey of phytopathogenic fungi in 2003. A relevant specimen collected in Yangpyeong was deposited at the Westerdijk Fungal Biodiversity Institute, and the corresponding isolate was registered as CBS 131930 (Centraalbureau voor Schimmelcultures; CBS). Although this strain was later included in a taxonomic study of the genus Pseudocercospora by Crous et al. [3], its detailed morphological description was not provided. Our field observations show that the disease can be severe in Korea, with extensive leaf spots leading to premature defoliation of up to 80% before the expected leaf fall in autumn.
Subsequent comprehensive studies on cercosporoid fungi—i.e., Shin and Kim [4], Nakashima et al. [5], and Groenewald et al. [6]—overlooked this fungus. Moreover, nucleotide sequence data obtained from one aforementioned strain of Ps. securinegae are available in GenBank, which restricts conducting reliable molecular comparisons and phylogenetic placement.
Despite its early report, subsequent taxonomic studies have largely overlooked Ps. securinegae; hence, its morphological features have yet to be sufficiently characterized. To address this gap, this study provides a detailed morphological description supported by photomicrographs and supplemented with additional sequence data, including sequences of the DNA-directed RNA polymerase II second-largest subunit (rpb2) gene for the first time.
During our field forays in Korea, 14 samples of F. suffruticosa with leaf spot symptoms of Pseudocercospora infection were collected and preserved in the herbaria of Korea University (KUS; Seoul, Korea) and Jeonbuk National University (JBNU; Jeonju, Korea). Detailed information regarding the herbarium number, location, and collection dates are listed in Table 1.
Table 1. Korean samples of Pseudocercospora securinegae from Flueggea suffruticosa used in this study
table
CBS: Centraalbureau voor Schimmelcultures culture collection; KACC: Korean Agricultural Culture Collection, Rural Development Administration; ITS: internal transcribed spacer; actA: actin; tef1: translation elongation factor 1-alpha; rpb2: DNA-directed RNA polymerase II second-largest subunit.
Fresh specimens were used for conducting morphological observations of the fungus using an Olympus BX50 microscope (Olympus, Tokyo, Japan). Stromata-bearing conidiophores and conidia were picked with a sterile needle, mounted in a drop of sterile water, and examined. Photomicrographs of taxonomic features were captured using a Zeiss AX10 microscope equipped with an AxioCam MRc5 digital camera (Carl Zeiss, Oberkochen, Germany). A total of 30 measurements were recorded for each diagnostic structure. To obtain pure conidial cultures, conidia were isolated from symptomatic tissues using sterile forceps and streaked onto 2% potato dextrose agar (PDA; Difco, France) supplemented with 200 mg/L streptomycin sulfate, followed by incubation at 25°C. After four days, the germinated conidia were transferred to a new PDA plate without antibiotics. A total of six isolates were obtained and deposited in the Korean Agricultural Culture Collection (KACC; Rural Development Administration, Korea). The accession numbers are listed in Table 1.
Observed morphological characteristics of the fungus include the following: Leaf spots were amphigenous, irregular to angular, 5–15 mm in diameter (or larger by coalescing discrete lesions), with greyish lesion with dark brown margin (Fig. 1A–D). Caespituli were amphigenous, albeit primarily hypophyllous, fasciculate to solitary (Fig. 1E). Mycelium was internal and external, with septate and branched hyphae. Stroma were well-developed, sub-globular, erumpent, light to dark brown, up to 30–60 µm in diameter (Fig. 1F). Conidiophores were numerous, in dense fascicles, smooth-walled, 2–4-septate, cylindrical to sub-cylindrical, straight, 14–37(–40) × (2.5–)3–5 µm. Conidiogenous cells were monoblastic, hyaline, unbranched, 3–6 µm, with inconspicuous scars (Fig. 1G). Conidia were solitary, narrowly obclavate to cylindrical, smooth, hyaline, straight to slightly curved, 6–11-septate, apex obtuse, base truncate, 25–85(–92) × (2.9–)3.3–5.4 µm, with an unthickened and not darkened hila (Fig. 1H–I). Fourweek-old fungal colonies grown on PDA at 25°C were slow-growing, 10 mm in diameter, olivaceous to greyish towards the center, with an erose or dentate margin, developed moderate aerial mycelium, and dark olivaceous to black on the reverse surface (Fig. 1J).

Fig. 1. Symptoms and morphological characteristics of Pseudocercospora securinegae on Flueggea suffruticosa. (A) Leaf spots with a greyish lesion on infected leaves. (B) Close-up view of individual leaf spots showing a well-defined greyish lesion. (C, D) Overall view of infected plants showing severe disease symptoms in early September; nearly all leaves had fallen, and only several new leaves remained (arrows). (E) Abundant greyish fungal fructification on the upper leaf surface. (F) Stromata. (G) Conidiophores and conidiogenous cells. (H, I) Conidia. (J) Four-week-old colonies of Ps. securinegae growing on a potato dextrose agar at 25°C.
Due to the absence of a detailed description of Ps. securinegae in previous studies, the morphological characteristics of the Korean isolates were compared with an original description of C. securinegae [1], from which the species was transferred. The Korean isolates correspond well with the original description of C. securinegae in stromata size (35–60 µm vs. 35–50 µm), conidiophore morphology and dimensions (1437(–40) × (2.5–)3–5 µm vs. 25–40 × 3–4 µm), and conidial shape and size (25–85(–92) × (2.9–)3.3–5.4 µm vs. 25–125 × 3–4 µm). Although our specimens exhibited slightly larger stromata and more septate conidia, these differences correspond to the expected intraspecific variation of Pseudocercospora. In addition, among Flueggea hosts, Ps. aberrans has been reported on F. virosa from China [7] and was therefore considered in our comparative assessment. The species shows clear distinguishing features, including smaller stromata (15–35 µm), very long conidiophores (up to 205 µm), and pale olivaceous, 1–3-septate conidia. These distinct characteristics clearly separate Ps. aberrans from Korean isolates.
To conduct molecular analysis, genomic DNA was extracted from six isolates cultured for two weeks on PDA using MaglistoTM 5M kits (Bioneer, Daejeon, Korea) following the manufacturer’s protocol. The primer pairs V9G/ITS4, ACT-512F/ACT-783R, EF1-728F/EF-986R, and P-RPB2-F/P-RPB2-R were used to amplify and sequence the nucleotide sequences of the internal transcribed spacer (ITS) region, actin (actA), translation elongation factor 1-alpha (tef1), and rpb2 genes, respectively [5,8,9]. The polymerase chain reaction products were purified and sequenced by a commercial sequencing company (Bionics, Seoul, Korea). The obtained forward and reverse sequences were assembled in MEGA 11 [10], and concatenated data were deposited in the NCBI database (Table 1).
The dataset of each gene was created and aligned separately, and each alignment was combined into a single multigene dataset of ITS + actA + tef1 + rpb2 using the SequenceMatrix software [11]. Trochophora simplex (CBS 214744) was selected as an outgroup [5]. A phylogenetic tree was constructed using three different methods: maximum parsimony (MP), maximum likelihood (ML), and Bayesian inference (BI) analyses. For the MP analysis, a heuristic search with the tree-bisection algorithm was used in PAUP* 4.0a [12]. The final dataset contained 34 taxa with a total of 1,853 characters, of which 225 (12.14%) were variable and parsimony-uninformative, and 541 (29.2%) were informative for the parsimony analysis. The ML analysis was conducted using raxmlGUI 2.0.13, with the GTR substitution model and a GAMMA distribution [13]. The reliability of the trees was tested using 1,000 bootstrap replications. To perform the BI analysis, the optimal nucleotide substitution model was first determined using MrModelTest version 2.4 [14], which identified GTR + I + G as the best model. This model was then used for the BI analysis in MrBayes version 3.2.7 [15]. The analysis utilized a Markov Chain Monte Carlo approach with four chains, running for 1,000,000 generations and sampling every 1,000 generations. Bayesian consensus trees were generated after a burn-in period of 25%. The resulting tree was visualized using FigTree version 1.4.4 [16]. Bootstrap support values greater than 70% from the MP and ML analyses, along with posterior probabilities of 0.90 or higher from the BI analysis, were added to the corresponding branches and nodes of the tree to indicate the robustness of the branches. The six newly obtained isolates clustered together with Ps. securinegae (CBS 131930) and formed a distinct clade, separated from other Pseudocercospora species. This branch was strongly supported by high bootstrap values and posterior probabilities from the MP, ML, and BI analyses (100/100/1) (Fig. 2).
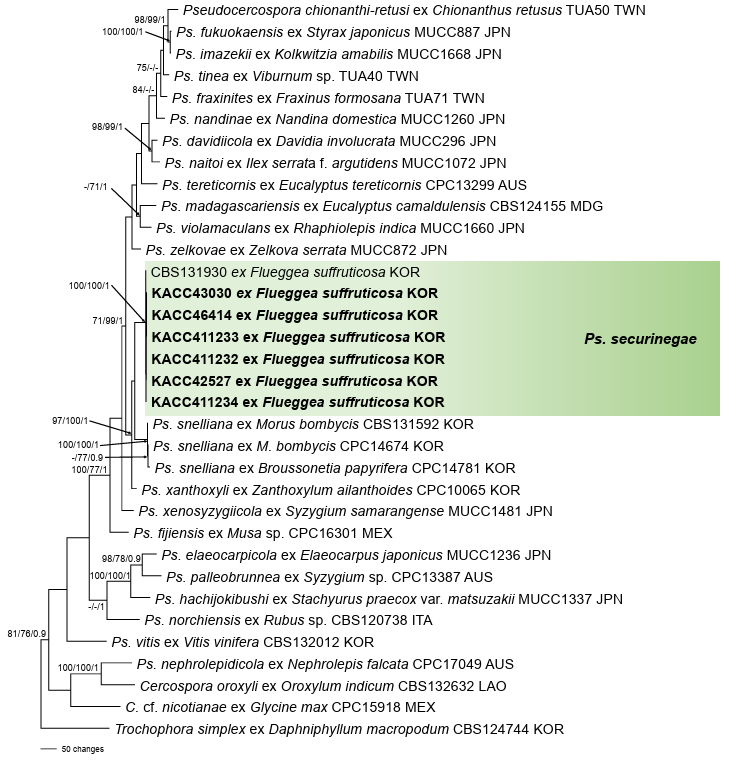
Fig. 2. A maximum parsimony tree of Pseudocercospora securinegae constructed using a multigene dataset of ITS + actA + tef1 + rpb2. Isolates obtained in this study are shown in bold. Bootstrap values (>70%) obtained from maximum parsimony and maximum likelihood as well as posterior probabilities (>90%) from Bayesian inference are shown on the respective related branches. ITS: internal transcribed spacer; actA: actin; tef1: translation elongation factor 1-alpha; rpb2: DNA-directed RNA polymerase II second-largest subunit.
The resulting sequences were compared with reference sequences in GenBank using the BLASTn search. The ITS region sequences of all six isolates were identical and shared 99.56% similarity with Ps. fuligena (GU214675) and Ps. chengtuensis (GU214672). For the actA gene, the sequences from all six isolates were also identical and showed 100% identity with Ps. securinegae (GU320479). For the tef1 gene, three isolates (KACC411234, KACC42527, and KACC411232) were identical and exhibited 99.68% similarity with Ps. securinegae (GU384487). The remaining three isolates (KACC411233, KACC43030, and KACC46414) differed by a single nucleotide substitution (adenine to thymine), resulting in 99.36% similarity with the same reference sequence. Due to the unavailability of Ps. securinegae rpb2 sequence data in GenBank, one isolate (KACC411234) showed 98.06% similarity with Ps. halleriae (PP404492), whereas the remaining five isolates differed by one nucleotide (cytosine to thymine) and showed 97.89% similarity with Ps. halleriae (PP404492).
F. suffruticosa (Euphorbiaceae) is a shrub or small tree species native to Russia and East Asia, including China, Japan, Korea, and Mongolia [17]. In Korea, it commonly grows across the country along forest edges, valleys, and mountainous areas, and its young sprouts are edible, having been widely used as an indigenous vegetable [18–20]. Historically, F. suffruticosa has also been used in traditional medicine in East Asia, primarily for the treatment of infantile paralysis, neurasthenia, and facial paralysis [21]. Its chemical constituents include a variety of bioactive compounds, such as alkaloids, terpenoids, flavonoids, and phenols. Among these, Securinega alkaloids are particularly important natural products because of their pharmaceutical potential [22]. Leaf spot disease caused by Ps. securinegae can significantly affect plant growth, as premature defoliation reduces the photosynthetic capacity of infected plants, which, in turn, weakens the overall vigor and decreases the amount of harvestable plant material. Premature defoliation may also influence the accumulation of key bioactive compounds that are important for the medicinal use of this species.
Although plants belonging to the genus Flueggea are widely used in traditional medicine, little is known about the phytopathogenic fungi associated with this genus [23]. To date, only a few fungi have been reported on F. suffruticosa, including Erysiphe securinegae (powdery mildew) in China, Japan, Korea, and Russia; Nothoravenelia japonica (rust) in China and Korea; and Ps. securinegae (leaf spot) in China, Japan, and Korea [3,24–26]. The distribution of F. suffruticosa, which is native to East Asia, aligns with the reports of Ps. securinegae, which has been documented only in China, Japan, and Korea.
This study provides the first comprehensive description of the morphological characteristics of Ps. securinegae from Korean isolates and contributes additional molecular data for the species—rpb2 gene sequences in particular, which were previously unavailable for this species. Given the medicinal importance of F. suffruticosa and its widespread distribution across the Korean Peninsula, our findings provide valuable insights into the identity and biology of this fungal pathogen.
No conflict of interest was reported by the authors.
This research was supported by a grant of the Global-Learning and Academic Research Institute for Master’s and PhD students and Postdocs (LAMP) Program of the National Research Foundation of Korea (NRF) (No. RS-2024-00443714) funded by the Ministry of Education. Support was also received by the R&D Program for Resolving Current Issues in Forest Disaster and Damage (Project No. RS-202400403034) funded by the Korea Forest Service (Korea Forestry Promotion Institute).
1. Chupp C. A monograph of the fungus genus Cercospora. New York: Ithaca; 1954.
2. Deighton FC. Studies on Cercospora and allied genera. VI. Pseudocercospora Speg., Pantospora Cif. and Cercoseptoria Petr. Mycol Pap 1976;140:1-168.
3. Crous PW, Braun U, Hunter GC, Wingfield MJ, Verkley GJM, Shin HD, Nakashima C, Groenewald JZ. Phylogenetic lineages in Pseudocercospora. Stud Mycol 2013;75:37-114. https://doi.org/10.3114/sim0005
[DOI]
4. Shin HD, Kim JD. Cercospora and allied genera from Korea. Suwon: National Institute of Agricultural Science and Technology; 2001.
5. Nakashima C, Motohashi K, Chen CY, Groenewald JZ, Crous PW. Species diversity of Pseudocercospora from Far East Asia. Mycol Prog 2016;15:1093-117. https://doi.org/10.1007/s11557-016-1231-7
[DOI]
6. Guo YL, Hsieh WH. The genus Pseudocercospora in China. Beijing: International Academic Publishers; 1995.
7. Groenewald JZ, Chen YY, Zhang Y, Roux J, Shin HD, Shivas RG, Summerell BA, Braun U, Alfenas AC, Ujat AH, et al. Species diversity in Pseudocercospora. Fungal Syst Evol 2024;13:29-89. https://doi.org/10.3114/fuse.2024.13.03
[DOI]
8. Videira SIR, Groenewald JZ, Nakashima C, Braun U, Barreto RW, de Wit PJGM, Crous PW. Mycosphaerellaceae – Chaos or clarity? Stud Mycol 2017;87:257-421. https://doi.org/10.1016/j.simyco.2017.09.003
[DOI]
9. Choi IY, Abasova L, Park JH, Shin HD. Revisiting Pseudocercospora puerariicola associated with leaf spot on Pueraria montana var. lobata with morphological and molecular-phylogenetic data in Korea. J Plant Pathol 2024;107:675-80. https://doi.org/10.1007/s42161-024-01749-2
[DOI]
10. Tamura K, Stecher G, Kumar S. MEGA11: Molecular evolutionary genetics analysis version 11. Mol Biol Evol 2021;38:3022-7. https://doi.org/10.1093/molbev/msab120
[DOI]
11. Vaidya G, Lohman DJ, Meier R. SequenceMatrix: Concatenation software for the fast assembly of multi-gene datasets with character set and codon information. Cladistics 2011;27:171-80. https://doi.org/10.1111/j.1096-0031.2010.00329.x
[DOI]
12. Swofford DL. PAUP: Phylogenetic analyses using parsimony (and other methods) 4.0b10. Sunderland: Sinauer Associates; 2002.
13. Silvestro D, Michalak I. RaxmlGUI: A graphical front-end for RAxML. Org Divers Evol 2012;12:335-7. https://doi.org/10.1007/s13127-011-0056-0
[DOI]
14. Nylander JAA. MrModeltest 2.0. Program distributed by the author. Uppsala: Uppsala University; 2004.
15. Ronquist F, Teslenko M, van der Mark P, Ayres DL, Darling A, Höhna S, Larget B, Liu L, Suchard MA, Huelsenbeck JP. MrBayes 3.2: Efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol 2012;61:539-42. https://doi.org/10.1093/sysbio/sys029
[DOI]
16. Rambaut A. Fig Tree ver. 1.3.1 [Internet]. Edinburgh: University of Edinburgh; 2009 [cited 2025 Jul 16]. Available from http://tree.bio.ed.ac.uk/software/figtree
17. POWO. Plants of the world online [Internet]. London: Royal Botanic Gardens, Kew; 2025 [cited 2025 Jul 16]. Available from: https://powo.science.kew.org/
18. Park KR, Park CW. Flueggea suffruticosa (Pall.) Baill. In: Flora of Korea Editorial Committee, editor. Flora of Korea – Vol. 5b Rosidae: Elaeagnaceae to Sapindaceae. Incheon: The National Institute of Biological Resources; 2015.
19. Chung TH. Korean Flora. Vol. 1. Woody plants. Seoul: Sinji-sa; 1957.
20. eFloras. Flora of China [Internet]. Cambridge, MA: Harvard University Herbaria; 2025 [cited 2025 Sep 16]. Available from: http://www.efloras.org
21. Peng YL, Zeng N, Yao QY, Peng CY, Sheng WB, Li B, Wang W. A review of the medicinal uses, phytochemistry and pharmacology of genus Flueggea. Curr Chin Sci 2023;3:213-41. https://doi.org/10.2174/2210298102666220920110106
[DOI]
22. Lee C, Han S. Discovery, structure revision, synthesis, and application of all known and even unknown securingine alkaloids. Nat Prod Rep 2025;42:1251-64. https://doi.org/10.1039/D5NP00025D
[DOI]
23. Farr DF, Rossman AY. Fungal databases, U.S. National Fungus Collections [Internet]. Washington DC: Agricultural Research Service, U.S. Department of Agriculture; 2025 [cited 2025 Jul 16]. Available from: https://fungi.ars.usda.gov/
24. Braun U, Cook RTA. Taxonomic manual of the Erysiphales (powdery mildews). CBS Biodiversity Series No. 11. Utrecht: CBS-KNAW Fungal Biodiversity Centre; 2012.
25. Ji JX, Li Z, Li Y, Kakishima M. Life cycle of Nothoravenelia japonica and its phylogenetic position in Pucciniales, with special reference to the genus Phakopsora. Mycol Prog 2019;18:855-64. https://doi.org/10.1007/s11557-019-01496-0
[DOI]
26. Kim CJ. A provisional List of Uredinales of Korea. Kor J Microbiol 1963;1:51-64.