Dae Ho Kim1 , Young-Nam Kwag1 , Hyung So Kim1 , Chang Sun Kim2* , and Jong Kyu Lee3*
1Forest Biodiversity Research Division, Korea National Arboretum, Pocheon, 11186, Korea
2Forest Biodiversity Conservation Division, Korea National Arboretum, Pocheon, 11186, Korea
3Department of Forest Environment Protection, College of Forest and Environmental Sciences, Kangwon National University, Chuncheon, 24341, Korea
*Correspondence to changsun84@korea.kr, jongklee@kangwon.ac.kr
Korean Journal of Mycology (Kor J Mycol) 2025 December, Volume 53, Issue 4, pages 237-243.
https://doi.org/10.4489/kjm.2025.53.4.1
Received on October 17, 2025, Revised on November 10, 2025, Accepted on November 10, 2025, Published on December 31, 2025.
Copyright © The Korean Society of Mycology.
This is an Open Access article which is freely available under the Creative Commons Attribution-Non-Commercial 4.0 International License (CC BY-NC) (https://creativecommons.org/licenses/by-nc/4.0/).
Recently added new species and taxonomic re-evaluations of previously recorded ones have warranted a revision of Scutellinia specimens deposited in Korea. By applying an integrative taxonomic framework to Scutellinia specimens curated at the Korea National Arboretum (KH), we recorded two species previously unreported in Korea, namely Scutellinia sinosetosa and S. subhirtella. Morphological assessment covered the macroscopic traits of apothecia as well as microscopic characteristics, including asci, paraphyses, marginal hairs, ascospore size, and ornamentation. Molecular evidence was obtained from internal transcribed spacer (ITS) and large subunit (LSU) rDNA sequences; phylogenetic analyses placed the Korean specimens in strongly supported clades alongside authenticated reference material, confirming their identities. S. sinosetosa is distinguished by small apothecia and low, reticulate ascospore ornamentation, whereas S. subhirtella is characterized by medium-sized, ellipsoid ascospores densely covered with discrete, fine verrucae. These findings represent the first confirmed records of both species in Korea, extend the known distribution of Scutellinia on the Korean Peninsula, and provide ITS/LSU barcode data to facilitate reliable molecular identification in future studies.
Pezizales, Scutellinia, Scutellinia sinosetosa, Scutellinia subhirtella
The genus Scutellinia (Pezizales, Pyronemataceae) is a group of saprophytic discomycetes that typically produces apothecia on decaying wood or moist soil. Members of this genus are morphologically characterized by disc- to cup-shaped fruiting bodies, dark-colored marginal hairs, and ornamented, ellipsoid ascospores [1]. Scutellinia species are distributed worldwide, with numerous reports originating from temperate regions of the Northern Hemisphere [1–3]. As of July 2025, approximately 150 valid species of the genus are listed in Index Fungorum (October 2025).
Many of these species were originally described based solely on morphological features. However, characteristics such as spore ornamentation, marginal hair structure, and coloration or size of the fruiting bodies can vary depending on specimen condition, environmental factors, and the observer’s interpretation, making species-level identification unreliable in many cases [1,4]. In response, molecular phylogenetic analyses using ribosomal DNA regions, such as internal transcribed spacer (ITS) and large subunit (LSU), have become increasingly important tools for accurate classification and identification of Scutellinia species [1,5,6].
Angelini et al. [1] conducted a broad phylogenetic analysis of numerous specimens and proposed nearly 50 putative phylogenetic species, noting that several morphologically identified taxa were actually polyphyletic. These findings, along with other recent studies, suggest that many previously reported species may not represent true taxonomic units, underscoring the need to reassess species concepts and perform more precise identifications [3,7].
Despite a growing body of sequencing data, more than half of the currently recognized Scutellinia species still lack ITS and LSU sequences in public databases, such as GenBank. Furthermore, several registered sequences were not linked to type specimens or voucher materials, and some were positioned unexpectedly in the phylogenetic trees [5,6]. These issues highlight the importance of careful selection and verification of comparative sequences, and they emphasize the need for a critical interpretation of phylogenetic results. Nevertheless, ITS and LSU remain the most widely used and essential molecular markers for species identification in Scutellinia. They have been consistently applied for the discovery and description of new species and taxa worldwide [1,8].
In Korea, studies on Scutellinia have been historically limited. However, two new species and eight new records were reported in morphological investigations by Han et al. [8] and Choi et al. [9], significantly expanding the known diversity of the genus in the region. Additional collections have since been reported in areas, such as Mudeungsan National Park [10]. Nonetheless, some previously reported species were later excluded based on morphological reassessments or their identification has been questioned [6,9], warranting a comprehensive taxonomic revision. Morphological and molecular examinations are essential for a more accurate assessment of species diversity [1,5,8].
Accordingly, we re-examined Scutellinia specimens deposited at the Korea National Arboretum (KH), during which we identified several species previously unrecorded in Korea. These taxa are reported herein.
This study used dried specimens of the genus Scutellinia deposited at the Forest Biological Specimen Center of the Korea National Arboretum (KH). To clarify the taxonomy of the unrecorded taxa, we examined both the morphological traits and DNA sequences of selected specimens (Table 1).
Table 1. GenBank accession numbers used for the phylogenetic analyses in this study, including origin and specimen number
table
ITS: internal transcribed spacer; LSU: large subunit ribosomal DNA.
Ttype strain.
The sequences from this study are bold.
Habitat conditions were recorded through photographs taken during the initial collection. The macroscopic and microscopic features were documented using a stereomicroscope (Leica Z16 APO; Leica, Wetzlar, Germany) and a compound light microscope (Leica DM 6000 FS; Leica). Microscopic preparations were mounted in distilled water, and 3% potassium hydroxide (KOH) to facilitate the observations and measurements of ascospores, asci, marginal hairs, and tissue structures.
Genomic DNA was isolated from the dried specimens using the DNeasy Plant Mini Kit (Qiagen, Hilden, Germany), following the manufacturer’s protocol. The ITS and LSU of ribosomal DNA regions were targeted for amplification. The ITS region was amplified using primers ITS5 and ITS4, whereas the LSU region was amplified using the primers LR0R and LR7 [11]. PCR reactions were prepared using either AccuPower® Taq PCR PreMix or AccuPower® Gold HotStart Taq PCR PreMix & Master Mix (Bioneer, Daejeon, Korea). The 20 μL PCR mixture included 1 μL each of the forward and reverse primers, 1 μL of DNA template, and nuclease-free water to the final volume. Each PreMix contained all necessary PCR reagents, including buffer, dNTPs, Taq polymerase, and tracking dye. Thermal cycling was conducted as follows: initial denaturation at 94°C for 5 min; 30 cycles of 94°C for 20 s, 56°C for 20 s, and 72°C for 30 s; and a final elongation at 72°C for 3 min. PCR products were stored at 4°C until sequencing.
The amplified products were purified and sent to Macrogen, Inc. (Seoul, Korea) for sequencing. Sequence reads were trimmed, assembled, and edited using Geneious Prime software. BLASTn searches (v. 2.13.0) were conducted against the GenBank database to determine species-level similarity. Sequences were aligned using MAFFT version 7 and manually adjusted as necessary. Phylogenetic relationships were inferred using maximum likelihood (ML) and Bayesian inference (BI). JModelTest v. 2.1.10 (https://github.com/ddarriba/jmodeltest2) was used to identify the best-fitting model of nucleotide substitution, which was GTR + I + G. BI analysis was performed over 1,000,000 generations, with the first 250,000 discarded as burn-in. Consensus trees were visualized using FigTree v. 1.4.4 (https://tree.bio.ed.ac. uk/software/figtree/).
To assess the genetic identity of Scutellinia specimens curated at the KH, we sequenced the rDNA ITS and LSU regions and queried them against NCBI BLAST. The ITS sequences of KA21-0226 and KA250537 showed the highest similarity to Scutellinia subhirtella MCKU 010829-01 (GenBank AY220824), with 99.66% (580/582 bp) and 100% (582/582) identity, respectively. Matches were also detected to the unidentified Scutellinia sp. ʻIN06’ OMDL iNat #221953006 (PX118746) at 97.73% (604/618) and 97.69% (591/605) as well as to S. subhirtella HMAS 287034 (OQ534227) at 97.68% (547/560) and 97.68% (548/561). LSU sequences were likewise closest to S. subhirtella HMAS 287034 (OQ534511) at 99.55% (879/883) and 99.44% (883/888), followed by S. subhirtella MYC2105-15 (OL832133) at 99.33% (817/821) and 99.21% (881/888). Specimen KA24-0296 showed an ITS sequence most similar to S. superba MCKU 010719-06 (AY220818) at 99.65% (572/574), with the following additional S. superba accessions from MCKU showing 98.78–99.30% identity. A further match to S. sinosetosa ZM 137 (OQ024064) was observed at 98.66% (589/597). The LSU sequence was closest to S. sinosetosa HKAS 124373 (OP291093) at 99.45% (899/904) and S. subhirtella KS-95-059A (C) (DQ220423) at 98.53% (870/883). Based on NCBI BLAST searches of the ITS and LSU sequences from the KH specimens, KA21-0226 and KA25-0537 were most similar to S. subhirtella, whereas KA24-0296 showed similarity to S. superba and S. sinosetosa.
A maximum-likelihood tree based on the concatenated ITS+LSU alignment, together with the reference accessions (Table 1), placed KA21-0226 and KA25-0537 in a single clade with S. subhirtella (including the type) with strong support (Fig. 1). KA24-0296 clustered in a well-supported clade with S. sinosetosa HKAS 124373, clearly separated from neighboring lineages (Fig. 1). These phylogenetic positions were similar to the BLAST results.
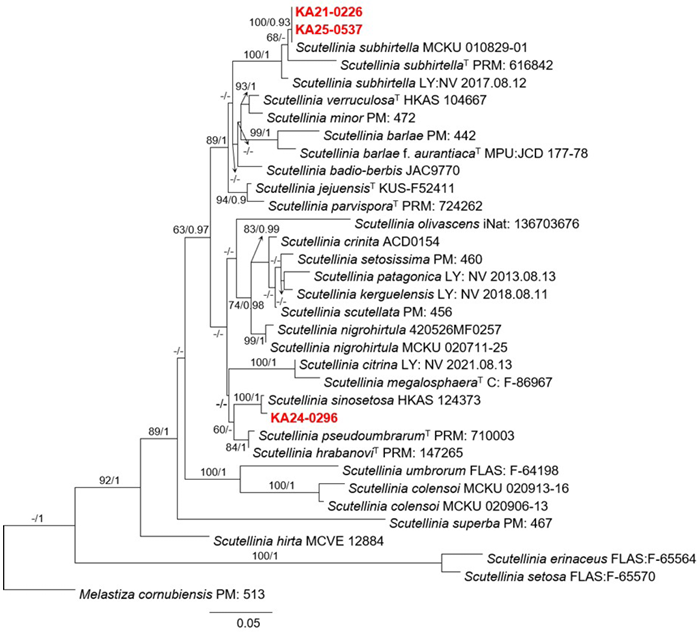
Fig. 1. RAxML phylogenetic tree inferred from a concatenated dataset of the internal transcribed spacer (ITS) and large subunit ribosomal DNA (LSU) regions. Maximum-likelihood bootstrap support (BS) values ≥ 50% and Bayesian posterior probabilities (BPP) ≥ 0.90 are shown at the nodes (BS/BPP). The newly described species in this study are indicated in red.
Scutellinia sinosetosa W.Y. Zhuang & Zheng Wang, Mycotaxon 69:352 (1998) [MB#447085] (Fig. 2)
Korean name: 동양털접시버섯
Description: Apothecia 3–7.9 mm in diameter, discoid, scattered, bright orange-yellow, margin with long and short hairs. Hairs 537–1,429 μm long, 29–55 μm thick, root-like branched at base, thick-walled, densely septate, yellow-brown. Asci cylindric, 193.2–269.38 × 14.5–20.25 μm, eight-spored, with uniseriate ascospores. Ascospores 18.8–23.3 × 11.73–15.79 μm, broadly ellipsoid to ellipsoid, hyaline, equilateral, rounded at the ends, verruculose. Paraphyses 233–290 μm long, 3.04–5.01 μm wide, filiform, slightly enlarged at apex, hyaline.
Specimen examined: Korea, Gyeonggi province, Pochun city, Gwangneung forest, 37°45’45″N, 127°09’25″E, elevation 183 m, on soil, June 11, 2024 (KA24-0296).
Remark: Scutellinia sinosetosa is morphologically similar to S. setosa, S. crinita, and S. olivascens in having ellipsoid, warted ascospores and long marginal hairs [12]. However, it can be distinguished from S. setosa by its longer marginal hairs (700–1200 µm vs. 600–900 µm) with less frequent branching and more densely warted spore surfaces. Compared to S. crinita, S. sinosetosa has thicker marginal hairs and rougher, more extensively ornamented spores. It differs from S. olivascens by its thicker, longer marginal hairs, and more coarsely warted spores, while S. olivascens exhibits finer ornamentation and paler, narrower hairs [3,8,12,13].
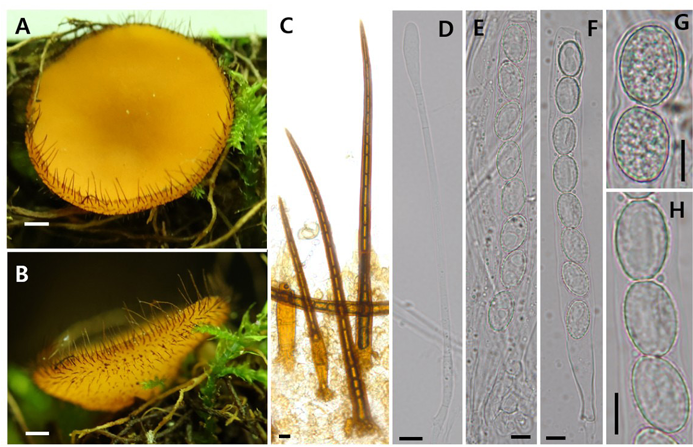

Fig. 2. Scutellinia sinosetosa KA24-0296. (A, B) Apothecia; (C) Marginal hairs; (D) Paraphyses; (E, F) Ascus; (G, H) Ascospores. Scale bars = 1 mm (A, B), 10 μm (C–H).
Scutellinia subhirtella Svrček, Česká Mykol. 25(2): 85 (1971) [MB#323350] (Fig. 3)
Korean name: 잔털접시버섯
Description: Apothecia 1.3–7.9 mm in diameter, scattered but occasionally gregarious, patellarioid to disciform, orange to orange-red, margin shortly but not densely hairy, with yellow-brown hairs. Hairs 251–443 μm long, 16–26 μm thick, root-like branched at base, thick-walled, densely septate. Asci cylindric, 197.59–280.59 × 14.45–17.98 μm, eight-spored, with uniseriate ascospores. Ascospores 17.73–22.68 × 10.97–14.72 μm, broadly ellipsoid to ellipsoid, rarely slightly narrowly ellipsoid, hyaline, equilateral, rounded at the ends, densely verrucose. Paraphyses 258–290 μm long, 3.06–4.17 μm wide, filiform, slightly enlarged at apex, hyaline.
Specimen examined: Korea, Gangwon province, Pyongchang-gun, Jinbu-myeon, Dongsan-ri, 37°44’20″N, 128°35’33″E, on decayed wood, elevation 789 m, May 30, 2021 (KA21-0226); Korea, Gangwon province, Inje-gun, Girin-myeon, Gombaeryeong, 38°02’23″N, 128°28’5″E, elevation 757 m, on decayed wood, July 02, 2025 (KA25-0537).
Remark: Scutellinia subhirtella is morphologically comparable to S. setosa, S. umbrorum, and S. olivascens, sharing ellipsoid, warted ascospores and marginal hairs. However, it can be readily distinguished from S. setosa by its shorter, thick-walled marginal hairs with bifurcate bases and more uniformly ornamented ascospores. Compared to S. umbrorum, S. subhirtella has less reticulate ornamentation and shorter, broader hairs. It differs from S. olivascens by possessing more conspicuous, evenly distributed warts on the spore surface and generally darker, thicker marginal hairs. These morphological differences support the recognition of S. subhirtella as a distinct species [8,13–15]. This species resembles S. patagonica Gamundí in spore size and general apothecial appearance. However, S. subhirtella is distinguished by the presence of subcylindrical, thin-walled, hyaline hairs intermixed with marginal hairs, which are absent in S. patagonica. In addition, the ascospores of S. subhirtella (18–22 × 12–14 µm) are slightly smaller and more densely verrucose than those of S. patagonica (18.5–23.5 × 14.5–21 µm), and the ornamentation is typically more evenly distributed [13–15].
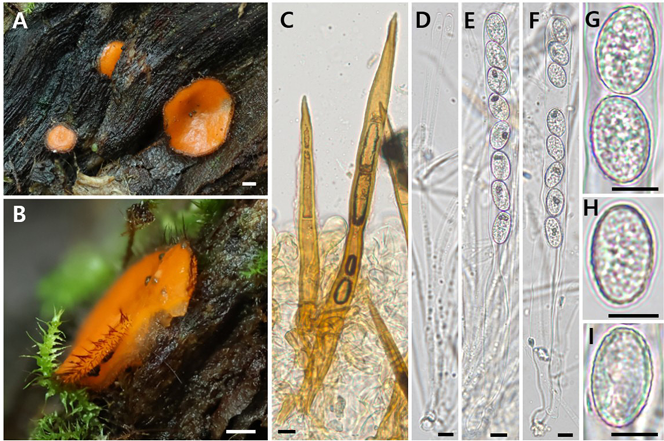

Fig. 3. Scutellinia subhirtella KA25-0537. (A, B) Apothecia; (C) Marginal hairs; (D) Paraphyses; (E, F) Ascus; (G-I) Ascospores. Scale bars = 1 mm (A, B), 10 μm (C–I).
Using an integrative taxonomic approach to KH vouchers, we confirmed two species previously unrecorded in Korea, Scutellinia sinosetosa and S. subhirtella. BLAST searches of the ITS and LSU sequences and a concatenated (ITS+LSU) ML phylogeny yielded similar outcomes: KA21-0226 and KA25-0537 were consistently supported as S. subhirtella, whereas KA24-0296 was supported as S. sinosetosa. Morphologically, S. sinosetosa is characterized by long, thick-walled marginal hairs and densely, coarsely ornamented ascospores, while S. subhirtella shows short, thick-walled marginal hairs that are often bifurcate at the base and ellipsoid ascospores (18–22 × 12–14 μm) with uniformly distributed fine verrucae.
In NCBI BLAST searches, Scutellinia sinosetosa KA24-0296 showed the highest similarity to MCKU accessions labeled S. superba, but these sequences lacked publicly verifiable linked morphology; therefore, a species-level assignment based on the labels alone is not possible. Consistent with this, Angelini et al. [1] withheld species-level confirmation, and Carbone and Rubini [2] treated the same clade as S. sinosetosa. Morphologically, S. sinosetosa has long, thick-walled marginal hairs and spores with dense, coarse warts and only weakly reticulate elements, whereas S. superba has been described with relatively shorter hairs and a more evident reticulate component with less densely developed warts [12,15]. Considering the diagnostic morphology of our vouchers, together with their phylogenetic position, the Korean material is best interpreted as S. sinosetosa.
In NCBI BLAST searches and ITS+LSU phylogenetic analyses, the Korean spcimens KA21-0226 and KA25-0537 showed the highest similarity to, and clustered with, reference sequences of S. subhirtella. Morphologically, they agree with S. subhirtella in having short, stout, thick-walled marginal hairs often bifurcate at the base, ellipsoid ascospores with dense fine verrucae lacking a true reticulum, and filiform paraphyses with orange pigment and slightly clavate apices [8,13–15]. Therefore, we regarded the Korean specimens as S. subhirtella.
Taken together, the concordance of morphological and molecular evidence enhances the confidence in our determinations. This study provides the first Korean records of S. sinosetosa and S. subhirtella, extends the known distribution of Scutellinia on the Korean Peninsula, and improves reproducibility and routine identification by linking KH-curated vouchers to publicly available ITS and LSU sequences. In Scutellinia, morphological characteristics alone are insufficient for identification, and an integrative taxonomic approach coupled with molecular markers is essential.
No potential conflict of interest was reported by the authors.
This study was supported by the Korea National Arboretum (Project No. KNA 1-1-28-24-1).
1. Angelini P, Bistocchi G, Arcangeli A, Venanzoni R, Rubini A. Molecular phylogenetic assessment of the genus Scutellinia (Pezizales, Pyronemataceae). Plant Biosyst 2023;157:18898. https://doi.org/10.1080/11263504.2022.2098869
[DOI]
2. Carbone M, Rubini A. Scutellinia blumenaviensis (Pezizales, Pyronemataceae) collected on the Caribbean Island of Guadeloupe (France). Ascomycete.org 2024;16:137-48. https://doi.org/10.25664/art-0392
[DOI]
3. Choi YJ, Shin HD, Han JG, Pfister DH. Scutellinia (Pezizales) in Korea, with a new species and eight new records. Nova Hedwigia 2013;97:457-76. https://doi.org/10.1127/0029-5035/2013/0120
[DOI]
4. Denison WC. Some species of the genus Scutellinia. Mycologia 1959;51:605-35.
https://doi.org/10.1080/00275514.1959.12024845
[DOI]
5. Lindemann U, Fellmann B, Friebes G, Van Vooren N. Scutellinia albopilosa (Pezizales, Pyronemataceae) – a new species from southern Germany. Ascomycete.org 2024;16:175-84. https://doi.org/10.25664/art-0397
[DOI]
6. Han JG, Choi YJ, Pfister DH, Shin HD. Scutellinia jejuensis (Pezizales), a new species from Korea. Mycotaxon 2010;112:47-53. https://doi.org/10.5248/112.47
[DOI]
7. Jeannerot F. European key to the genus Scutellinia. Ascomycete.org 2019;11:297-308. https://doi.org/10.25664/art-0287
[DOI]
8. Schumacher T. The genus Scutellinia (Pyronemataceae). Copenhagen: Council for Nordic Publications in Botany; 1990.
9. Wang YZ. The genera Scutellinia and Geneosperma (Discomycetes, Pezizales) in Taiwan. Bull Natl Mus Nat Sci 1998;11:119-28.
10. Zeng M, Gentekaki E, Zeng XY, Tian Q, Zhao Q, Hyde KD. Evolutionary relationships and allied species of Pyronemataceae, with segregation of the novel family Pyropyxidaceae. Mycosphere 2022;13:207-80. https://doi.org/10.5943/mycosphere/si/1f/7
[DOI]
11. Zhuang WY. The genus Scutellinia (Pyronemataceae) from China, with a key to the known species of the country. Mycosystema 2013;32:429-47.
12. Zhuang WY, Wang Z. Discomycetes of tropical China. II. Collections from Yunnan. Mycotaxon 1998;69:339-58.
https://doi.org/10.5962/p.415549
[DOI]
13. Kullman B. A revision of the genus Scutellinia (Pezizales) in Soviet Union. Scripta Mycol 1982;10:1-158.
14. Svrček M. Tschechoslowakische arten der Discomycetengattung Scutellinia (Cooke) Lamb. emend. Le Gal (Pezizales) I. Česká Mykol 1971;25:77-87.
https://doi.org/10.33585/cmy.25203
[DOI]
15. Yao YJ, Spooner BM. Notes on British species of Scutellinia. Mycol Res 1996;100:859-65. https://doi.org/10.1016/S0953-7562(96)80035-9
[DOI]