Seung-Yoon Oh1* and Myung Soo Park2
1School of Advanced Biosciences, Changwon National University, Changwon 51140, Korea
2Department of Mushroom Science, Korea National University of Agriculture and Fisheries, Jeonju 54874, Korea
*Correspondence to syoh@changwon.ac.kr
Korean Journal of Mycology (Kor J Mycol) 2025 December, Volume 53, Issue 4, pages 237-243.
https://doi.org/10.4489/kjm.2025.53.4.1
Received on October 17, 2025, Revised on November 10, 2025, Accepted on November 10, 2025, Published on December 31, 2025.
Copyright © The Korean Society of Mycology.
This is an Open Access article which is freely available under the Creative Commons Attribution-Non-Commercial 4.0 International License (CC BY-NC) (https://creativecommons.org/licenses/by-nc/4.0/).
Spiders occupy diverse habitats and closely interact with environmental substrates through web construction, locomotion, and prey capture, facilitating frequent contact with fungal propagules. Despite their ecological relevance, fungal communities associated with spiders have received limited attention, and little is known about how habitat context influences spider-associated fungal assemblages. In this study, we investigated the diversity and composition of culturable fungi associated with the Joro spider, Trichonephila clavata, a common and ecologically important species in South Korea. Whether fungal communities differed between urban and forest habitats was evaluated. Spider samples were collected from three urban park sites and three mountain forest sites in Changwon, South Korea. A total of 67 fungal strains representing 28 species were recovered. The fungal community was largely composed of ascomycetous fungi, with Alternaria, Cladosporium, and Penicillium frequently isolated, whereas most species occurred at low frequencies. No significant differences were detected between the habitats in terms of alpha diversity indices or community composition. These results suggest limited habitat differentiation in fungal communities associated with T. clavata, with host-related factors playing a stronger role than the habitat context.
Arthropod, Diversity, Ecology, Fungi, Urban ecology
Spiders are widespread generalist predators that interact continuously with environmental substrates such as vegetation, soil particles, and airborne biological materials through web construction, prey capture, and movement across habitats [1,2]. Through these activities, spiders are persistently exposed to fungal propagules from the surrounding environment, rather than host-specific infection processes [3]. Culturebased studies have shown that living spiders commonly carry diverse fungi on their external surfaces, many of which correspond to environmentally ubiquitous saprotrophic taxa associated with soil, plant debris, and phyllosphere habitats [4,5]. These observations suggest that spider-associated fungal assemblages largely reflect passive environmental acquisition and include taxa involved in pathogenic interactions. Nevertheless, research on spider–fungus associations has historically emphasized conspicuous fungal pathogenicity, particularly entomopathogenic infections that are readily detected because of visible symptoms or host mortality [6–8]. Recent reviews have highlighted that this focus leads to a substantial underestimation of non-pathogenic and saprotrophic fungi associated with spiders, which are rarely targeted by traditional surveys [9]. Consequently, most environmentally derived fungi that are incidentally associated with spiders remain poorly documented, limiting our understanding of spiders as ecological interfaces between fungal communities and habitat conditions.
Habitat context is a key determinant of fungal community composition, as vegetation structure, organic matter availability, and microclimatic stability collectively shape the diversity and relative abundance of saprotrophic fungi [10]. Forest ecosystems generally support high fungal diversity owing to continuous litter input and stable moisture regimes [11]. In contrast, urban green spaces, such as city parks, are characterized by managed vegetation, habitat fragmentation, and frequent disturbances, which can reorganize fungal communities rather than uniformly reduce diversity [12,13]. These habitat-driven differences in environmental fungal assemblages are expected to influence fungi incidentally associated with resident arthropods, including spiders, which continuously interact with their surroundings [3,14].
The Joro spider, Trichonephila clavata, commonly known as the Mudang spider in South Korea, is a large and conspicuous species native to East Asia, widely distributed across the Korean Peninsula [15]. It occurs across a broad range of habitats from mountain forests to urban green spaces, making it suitable for habitat-comparative studies. Although T. clavata has been extensively studied in ecological and molecular contexts, particularly following its introduction to North America [16,17], the fungal diversity associated with this species has received little attention, and no systematic assessment has been conducted in its native range. In this study, the fungal diversity associated with T. clavata in Changwon, South Korea, was investigated focusing on culturable fungi isolated from spiders. The aim of the study was to characterize the taxonomic composition of fungi commonly isolated from T. clavata and assess whether fungal assemblages differed between mountain forests and urban park habitats. This study establishes baseline information for future investigations by providing basic descriptive data on spider-associated fungi across contrasting environments.
This study was conducted in Changwon (South Korea) and focused on two distinct habitat types: urban parks and mountain forests. Three sites were selected for each habitat type, resulting in six sampling sites. The urban park sites were chosen from densely populated areas with moderate human activity, including recreational green spaces characterized by mowed lawns, ornamental plants, and paved paths: Gaeumjeong-dong Wetland Park (35.231°N, 128.694°E), Seok-dong 2 Park (35.212°N, 128.705°E), and Yongho-dong Yongji Park (35.222°N, 128.681°E). In contrast, mountain forest sites were selected from relatively undisturbed natural woodlands with dense canopy cover, leaf litter, and native vegetation, including broadleaf and coniferous trees, on Mt. Bulmo (35.231°N, 128.748°E), Mt. Cheonjabong (35.224° N, 128.742°E), and Mt. Jeongbyeong (35.244°N, 128.701°E). These forest sites had minimal human intervention and high humidity and soil organic matter content. Sampling was performed during the peak activity period of T. clavata (August–September, 2020) to ensure sufficient spider abundance. Spiders were collected from each of the six sites using sterile forceps and plastic containers (50 mL Falcon tubes) to minimize contamination. At each site, 5–8 individuals were captured from the orb webs. Collected spiders were immediately transported to the laboratory in a refrigerator at 4°C.
Before fungal isolation, the spiders were gently washed three times with sterile distilled water to remove external debris and potential transient contaminants from the exoskeleton. Washed spiders were then placed individually in a sterile 50 mL tube and refrigerated at 4°C for 3 days under starvation conditions. This starvation period allowed for complete digestion and excretion of the ingested prey as feces, reducing the risk of isolating gut-derived fungi unrelated to the spider-associated mycobiota. Following starvation, spiders were euthanized by light freezing at −20°C for 5 min, then dissected with sterile scalpels. Each spider was separated into two parts: the cephalothorax and legs. The dissected parts were immediately used for fungal isolation to prevent post-mortem microbial overgrowth. Fungal isolation was performed on potato dextrose agar (PDA; Difco, BD Biosciences, Franklin Lakes, NJ, USA). For each spider part (cephalothorax or legs), the tissue was placed on the surface of the medium (90 mm Petri dishes) and incubated at 25°C in the dark for 2 weeks, with daily monitoring for emerging mycelia. Emerging fungal colonies were subcultured onto fresh 60 mm PDA for pure isolation.
DNA extraction and PCR amplification were performed as previously described [18]. Briefly, genomic DNA was extracted using the AccuPrep Genomic DNA Extraction Kit (Bioneer, Daejeon, Korea) and PCR amplification was performed using an AccuPower PCR PreMix (Bioneer) and purified using Expin PCR Purification Kit (GeneAll Biotechnology, Seoul, Korea). The thermal cycling conditions consisted of an initial denaturation at 95°C for 5 min, followed by 35 cycles of denaturation at 94°C for 30 sec, annealing at 55°C for 30 sec, and extension at 72°C for 40 sec, with a final extension at 72°C for 5 min. PCR amplification was verified using 1% agarose gel electrophoresis before downstream sequencing. All fungal strains were characterized by amplification of the internal transcribed spacer (ITS) region of the nuclear ribosomal DNA using ITS1F and ITS4 [19,20]. After confirming the genus, each sample was further amplified with locusspecific primers for refined identification: beta-tubulin (BenA) using bt2a and bt2b [21] and translation elongation factor 1-alpha (EF1a) using EF1-728F and EF1-905R [22]. PCR conditions for BenA and EF1a were similar to those for ITS, with annealing temperatures adjusted to 55°C and 58°C, respectively. Purified amplicons were sequenced by Macrogen Inc. (Seoul, Korea) using an ABI 3730xl sequencer.
The raw sequences were assembled and edited using MEGA5 [23]. Initial identification was performed using BLASTn against the NCBI GenBank database. For each identified genus, sequences from the type or authentic strains of related species were downloaded from GenBank. Multiple sequence alignments were generated using MAFFT v7.526 with default settings [24]. Phylogenetic trees were constructed in the MEGA5 software using the neighbor-joining method with the Kimura 2-parameter model, 1,000 bootstrap replicates for node support, and pairwise deletion of gaps. The final species assignments were based on monophyletic clustering with the reference strains. Duplicate species within each sample were removed to create unique species lists for each spider and site for downstream analyses. All sequences generated in this study were deposited in the NCBI GenBank under the accession numbers PX705746–PX705757, PX754293–PX754329, and PX754724–PX754741.
Fungal diversity and community structure were analyzed using R v4.5.1 [25] with vegan and ggplot2 packages [26,27]. Alpha diversity indices were calculated for each site: number of species, Chao1 estimator for unobserved species, Shannon–Wiener diversity index, and Pielou’s equitability. Differences in alpha diversity between urban parks and mountain forests were tested using unpaired t-tests, with normality checked via Shapiro–Wilk tests and variance homogeneity by Levene’s test. For beta diversity, community dissimilarities were computed using Bray–Curtis dissimilarities. Nonmetric multidimensional scaling (NMDS) ordination was performed with stress values of < 0.2, indicating a reliable representation. A permutational multivariate analysis of variance (PERMANOVA) was conducted using adonis2 to test for significant differences among habitats.
A total of 67 fungal strains were successfully isolated and cultured from the cephalothoraxes and legs of 38 T. clavata spiders collected from six sampling sites in Changwon, South Korea. These strains were identified using molecular marker regions, yielding 28 species belonging to three phyla, six classes, 11 orders, 13 families, and 18 genera (Fig. 1).
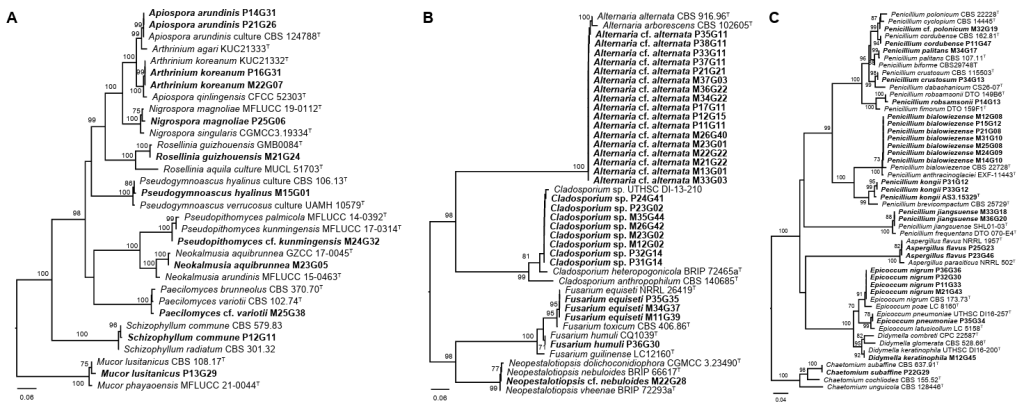
Fig. 1. Phylogenetic trees based on the neighbor-joining method with 1,000 bootstrap replicates for the (A) ITS, (B) EF1a, and (C) BenA regions. Bootstrap support values ≥ 70% are shown on the branches. Superscript T indicates sequences derived from ex-type strains. The scale bar indicates the number of substitutions per site. Sequences generated in this study are shown in bold. ITS: internal transcribed spacer; EF1a: elongation factor 1-alpha; BenA: beta-tubulin.
Most strains (65 of 67) belonged to the phylum Ascomycota, highlighting the dominance of Ascomycetes in the fungal communities associated with this spider species. At the class level, Dothideomycetes was the most abundant, comprising 33 strains (49.3%), followed by Eurotiomycetes with 19 strains (27.0%), and Sordariomycetes with 12 strains (17.9%). At the order level, Pleosporales dominated with 25 strains (37.3%), reflecting its common association with environmental molds, followed by Eurotiales with 19 strains (28.4%), and Cladosporiales with 8 strains (11.9%). Familylevel analysis revealed Aspergillaceae as the most abundant, with 19 strains (28.4%), closely followed by Alternariaceae with 17 strains (25.4%) and Cladosporiaceae with 8 strains (11.9%). At the genus level, Alternaria was the most dominant with 17 strains (25.4%), followed by Penicillium (16 strains, 23.9%), Cladosporium (8 strains, 11.9%), Epicoccum (5 strains, 7.5%), Apiospora (4 strains, 6.0%), and Fusarium (4 strains, 6.0%). Species identification was performed using phylogenetic analysis, and 28 species were identified. The dominant species was Alternaria cf. alternata (17 strains), followed by Cladosporium sp. (8 strains), and Penicillium bialowiezense (7 strains) (Fig. 2). Other notable species included Epicoccum nigrum (4 strains), Fusarium equiseti (3 strains), Apiospora arundinis (2 strains), Apiospora koreana (2 strains), Aspergillus flavus (2 strains), Penicillium jiangsuense (2 strains), and Penicillium kongii (2 strains). Of the 28 species, 21 species (75.0%) were found in only one sample. Among these 21 species, 18 were isolated as single strains.
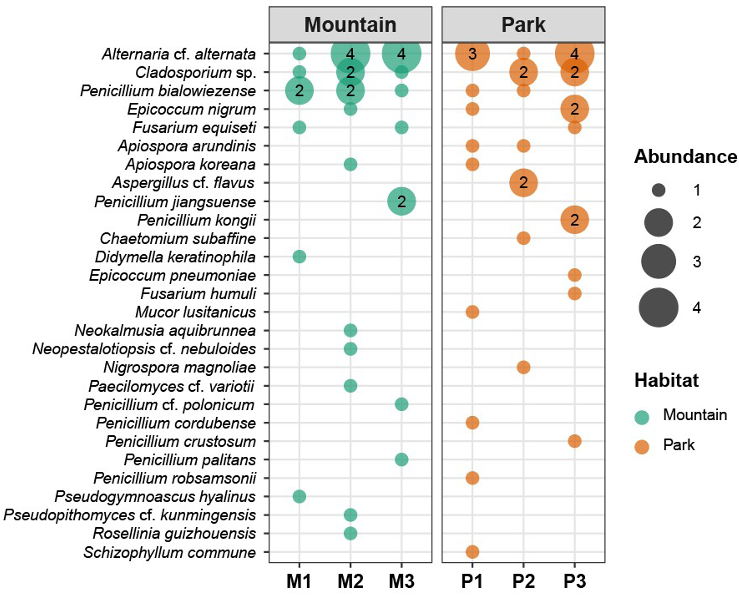
Fig. 2. Bubble plot showing the number of isolates for each species at individual sampling sites. Bubble size represents the abundance of isolates, and numbers ≥ 2 are shown inside the bubbles. Sampling sites are grouped by habitat: mountain (M1–M3) and park (P1–P3).
Alpha diversity analyses were conducted to compare fungal communities between urban parks (n = 3 sites, 20 spiders, and 34 strains) and mountain forests (n = 3 sites, 18 spiders, and 33 strains) (Fig. 3A). Mean species richness was 8.00 ± 1.00 in urban parks and 7.67 ± 2.08 in mountains, with no significant difference (t-test, t = −0.25, p = 0.82). The Chao1 estimator showed values of 18.94 ± 15.60 for parks and 13.33 ± 3.21 for mountains (t = −0.61, p = 0.60). Shannon–Wiener diversity was 2.85 ± 0.16 in parks and 2.73 ± 0.33 in mountains (t = −0.60, p = 0.59), indicating moderate diversity in both habitats without marked disparities. Pielou’s evenness was similarly comparable at 0.66 ± 0.01 for parks and 0.65 ± 0.02 for mountains (t = −0.64, p = 0.57). These results indicate that the alpha diversity of the fungal community associated with T. clavata did not vary significantly between urban and natural habitats. Beta diversity was assessed using Bray–Curtis dissimilarities, followed by NMDS ordination. The NMDS plot showed overlapping clusters for the urban park and mountain forest communities, with no clear separation along the axes (Fig. 3B). PERMANOVA confirmed the absence of significant differences between habitats (F = 0.936, R² = 0.190, p = 0.6). This lack of differentiation may reflect the shared fungal pools in both environments.
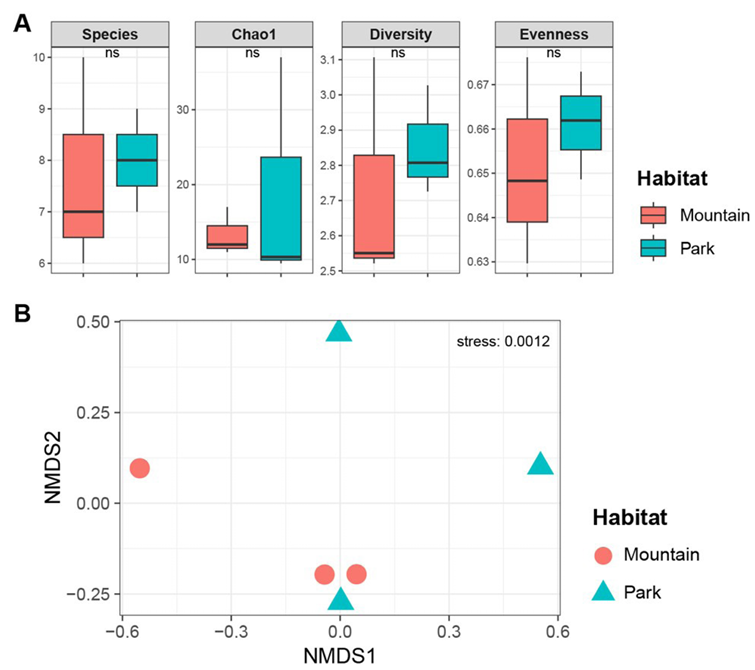
Fig. 3. Diversity analysis comparing habitats. (A) Alpha diversity indices, including Species (number of species), Chao1 (Chao1 richness estimator), Diversity (Shannon–Wiener diversity), and Evenness (Pielou’s evenness). ns indicates statistically not significant differences. (B) Beta diversity based on NMDS ordination using Bray–Curtis dissimilarities. NMDS: nonmetric multidimensional scaling.
However, subtle trends were observed, such as a comparable prevalence of Alternaria cf. alternata between mountains and parks (nine vs. eight strains) and a higher representation of Penicillium bialowiezense in mountains than in parks (five vs. two strains), although these differences were not statistically significant (Fisher’s exact test, p = 0.399).
The fungal communities associated with T. clavata in this study were largely composed of genera such as Alternaria, Cladosporium, and Penicillium, which together accounted for more than 60% of the isolated strains, including 17 Alternaria, 8 Cladosporium, and 16 Penicillium strains. This pattern likely reflects a combination of the ecological traits of these fungi and methodological factors related to culture-based isolation. These genera consist mainly of saprophytic or opportunistic fungi that are common in decaying organic matter, soil, and airborne substrates, environments that overlap with habitats frequently contacted by spiders [28–30]. Alternaria cf. alternata, the most abundant species in this study, is a widely distributed Dematiaceous fungus known for its tolerance to environmental stressors such as ultraviolet radiation and desiccation, which enables persistence on arthropod exoskeletons exposed to outdoor conditions [29,31]. Similarly, Cladosporium species are ubiquitous airborne and soil-associated fungi with a high spore-dispersal capacity, facilitating repeated deposition on spider surfaces during web construction and locomotion [30]. Penicillium species, including P. bialowiezense, are metabolically versatile saprophytes that grow rapidly in nutrient-rich media and tolerate mildly acidic and humid microenvironments that may arise from spider-derived residues or organic deposits [28,32,33].
Previous studies on spider-associated fungi have revealed comparable patterns across different arachnid hosts. For example, a study of Trichonephila pilipes from Indian forests reported frequent isolation of Aspergillus, Penicillium, Trichoderma, and Cladosporium from spider legs and saliva, with higher fungal loads on the legs attributed to contact with soil and vegetation [34]. Broader surveys of spider-associated fungi worldwide have identified Alternaria, Cladosporium, and Pencillium as common epibionts on spiders, which are typically regarded as ubiquitous environmental fungi rather than specialized arthropod pathogens [5,35]. These genera possess functional traits that allow survival or limited exploitation of the spider environment, including the production of enzymes such as chitinases and proteases that can degrade arthropod cuticles or utilize host-derived nutrients without necessarily inducing disease, consistent with commensal or weakly parasitic interactions [36–39]. In contrast, entomopathogenic specialists such as Gibellula and Beauveria were rarely recovered, which is consistent with the known biases of culture-based methods that favor fast-growing saprophytes over obligate pathogens [8,40,41].
The absence of significant differences in fungal diversity and community composition between urban parks and mountain forests suggests that habitat-level environmental variations exert a limited influence on host-related filtering processes. Both alpha-diversity indices and beta-diversity analyses indicated overlapping communities between habitats, despite apparent differences in the surrounding vegetation and land use. This pattern is consistent with a host filtering mechanism, whereby the characteristics of the spider exoskeleton, chemical secretions, and behaviors, such as web maintenance and grooming, constrain fungal colonization and persistence [3,7]. The large body size and orb-weaving lifestyle of T. clavata may generate relatively consistent microhabitat conditions, including moist but nutrient-poor surfaces that favor stress-tolerant generalist fungi, such as Alternaria and Penicillium, regardless of the broader habitat context. Similar host-driven structuring has been documented in other arthropods, where mycobiome composition tends to be more closely associated with host identity than with sampling location, potentially due to antimicrobial compounds or physical barriers present on the cuticle [42–45]. In urban environments, elevated pollution levels may reduce ambient fungal spore densities, but spider mobility and repeated movement among microhabitats may promote the homogenization of associated fungal communities.
A key limitation of this study is its reliance on culture-dependent methods, which are known to bias results toward easily cultivable, fast-growing fungi while underrepresenting slow-growing or unculturable taxa. Isolation of PDA favors mesophilic saprophytes such as Penicillium and Alternaria, potentially excluding obligate entomopathogens such as Cordyceps or Gibellula, which may require specialized media or host-specific conditions for growth [8]. Culture-independent approaches, including environmental DNA or metabarcoding analyses of materials recovered from spider surfaces, are likely to reveal additional diversity, including rare or viable but non-culturable taxa. Future studies integrating culture-independent and molecular approaches will provide a more comprehensive understanding of spider-associated fungal assemblages.
In conclusion, this study revealed that T. clavata hosts a diverse assemblage of associated fungi with slight variation in community composition between urban and forest habitats. Although taxonomic resolution was limited for some isolates, several species (e.g., Cladosporium) may represent previously unrecorded or poorly characterized taxa, warranting further phylogenetic and morphological investigations. These findings highlight spider-associated fungi as underexplored biological resources, particularly given that fungi linked to insects and arachnids can produce antimicrobial compounds, secondary metabolites, and industrially relevant enzymes, such as cellulases and lipases [46,47]. Collectively, these results support the ecological relevance and potential application of arachnid-associated mycobiomes and emphasize the need for expanded surveys to better characterize and utilize these fungal communities.
The authors declare that there are no conflicts of interest.
This research was supported by Changwon National University in 2025–2026.
1. Nyffeler M, Birkhofer K. An estimated 400-800 million tons of prey are annually killed by the global spider community. Sci Nat 2017;104:30. https://doi.org/10.1007/s00114-017-1440-1
[DOI]
2. Foelix R. Biology of spiders. 3rd ed. New York: Oxford University Press; 2010.
3. Vega FE, Blackwell M. Insect-fungal associations: Ecology and evolution. New York: Oxford University Press; 2005.
https://doi.org/10.1093/oso/9780195166521.001.0001
[DOI]
4. Gibbons AT, Idnurm A, Seiter M, Dyer PS, Kokolski M, Goodacre SL, Gorb SN, Wolff JO. Amblypygid-fungal interactions: The whip spider exoskeleton as a substrate for fungal growth. Fungal Biol 2019;123:497-506. https://doi.org/10.1016/j.funbio.2019.05.003
[DOI]
5. Yoder JA, Benoit JB, Christensen BS, Croxall TJ, Hobbs III HH. Entomopathogenic fungi carried by the cave orb weaver spider, Meta ovalis (Araneae, Tetragnathidae), with implications for mycoflora transfer to cave crickets. J Cave Karst Stud 2009;71:116-20.
6. Araújo JP, Hughes DP. Diversity of entomopathogenic fungi: Which groups conquered the insect body? Adv Genet 2016;94:1-39. https://doi.org/10.1016/bs.adgen.2016.01.001
[DOI]
7. Durkin ES, Cassidy ST, Gilbert R, Richardson EA, Roth AM, Shablin S, Keiser CN. Parasites of spiders: Their impacts on host behavior and ecology. J Arachnol 2021;49:281-98. https://doi.org/10.1636/JoA-S-20-087
[DOI]
8. Kuephadungphan W, Petcharad B, Tasanathai K, Thanakitpipattana D, Kobmoo N, Khonsanit A, Samson RA, Luangsaard JJ. Multi-locus phylogeny unmasks hidden species within the specialised spider-parasitic fungus, Gibellula (Hypocreales, Cordycipitaceae) in Thailand. Stud Mycol 2022;101:245-86. https://doi.org/10.3114/sim.2022.101.04
[DOI]
9. Nyffeler M, Hywel-Jones N. Diversity of spider families parasitized by fungal pathogens: A global review. J Arachnol 2024;52:151-85. https://doi.org/10.1636/JoA-S-23-007
[DOI]
10. Tedersoo L, Bahram M, Põlme S, Kõljalg U, Yorou NS, Wijesundera R, Ruiz LV, Vasco-Palacios AM, Thu PQ, Suija A, et al. Global diversity and geography of soil fungi. Science 2014;346:1256688. https://doi.org/10.1126/science.1256688
[DOI]
11. Baldrian P. Forest microbiome: Diversity, complexity and dynamics. FEMS Microbiol Rev 2017;41:109-30. https://doi.org/10.1093/femsre/fuw040
[DOI]
12. Hui N, Liu X, Kotze DJ, Jumpponen A, Francini G, Setälä H. Ectomycorrhizal fungal communities in urban parks are similar to those in natural forests but shaped by vegetation and park age. Appl Environ Microbiol 2017;83:e01797-17. https://doi.org/10.1128/AEM.01797-17
[DOI]
13. Scholier T, Lavrinienko A, Brila I, Tukalenko E, Hindström R, Vasylenko A, Cayol C, Ecke F, Singh NJ, Forsman JT, et al. Urban forest soils harbour distinct and more diverse communities of bacteria and fungi compared to less disturbed forest soils. Mol Ecol 2023;32:504-17. https://doi.org/10.1111/mec.16754
[DOI]
14. Bills GF, Christensen M, Powell M, Thorn G. Saprobic soil fungi. In: Mueller GM, Bills GF, Foster MS, editors. Biodiversity of fungi: Inventory and Monitoring Methods. Amsterdam: Elsevier; 2004. p. 271-301.
https://doi.org/10.1016/B978-012509551-8/50016-7
[DOI]
15. Hoebeke ER, Huffmaster W, Freeman BJ. Nephila clavata L Koch, the Joro spider of East Asia, newly recorded from North America (Araneae: Nephilidae). PeerJ 2015;3:e763. https://doi.org/10.7717/peerj.763
[DOI]
16. Grabarczyk EE, Querejeta M, Tillman PG, Wallace RD, Barnes BF, Meinecke CD, Villari C, Gandhi KJK, LaForest J, Elliott M, et al. DNA metabarcoding analysis of three material types to reveal Joro spider (Trichonephila clavata) trophic interactions and web capture. Front Ecol Evol 2023;11:1177446. https://doi.org/10.3389/fevo.2023.1177446
[DOI]
17. Russell JE, Mizera N, Brown CG, Chuang A, Coyle DR, Nelsen DR. Mitochondrial and Wolbachia phylogenetics of the introduced Jorō spider, Trichonephila clavata (Araneae: Araneidae) in North America. PeerJ 2025;13:e19952. https://doi.org/10.7717/peerj.19952
[DOI]
18. Oh SY, Jang Y. New Tricoderma species and six unrecorded species of endolichenic fungi isolated from lichens. Mycobiology 2025;53:435-49. https://doi.org/10.1080/12298093.2025.2519854
[DOI]
19. White TJ, Bruns TD, Lee SB, Taylor JW. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, editors. PCR protocols: A guide to methods and applications. New York: Academic Press; 1990. p. 315-22.
https://doi.org/10.1016/B978-0-12-372180-8.50042-1
[DOI]
20. Gardes M, Bruns TD. ITS primers with enhanced specificity for basidiomycetes – application to the identification of mycorrhizae and rusts. Mol Ecol 1993;2:113-8. https://doi.org/10.1111/j.1365-294X.1993.tb00005.x
[DOI]
21. Glass NL, Donaldson GC. Development of primer sets designed for use with the PCR to amplify conserved genes from filamentous ascomycetes. Appl Env Microbiol 1995;61:1323-30. https://doi.org/10.1128/aem.61.4.1323-1330.1995
[DOI]
22. Carbone I, Kohn LM. A method for designing primer sets for speciation studies in filamentous ascomycetes. Mycologia 1999;91:553-6. https://doi.org/10.1080/00275514.1999.12061051
[DOI]
23. Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S. MEGA5: Molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 2011;28:2731-9. https://doi.org/10.1093/molbev/msr121
[DOI]
24. Katoh K, Standley DM. MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Mol Biol Evol 2013;30:772-80. https://doi.org/10.1093/molbev/mst010
[DOI]
25. R Core Team. R: A language and environment for statistical computing. Vienna: R Foundation for Statistical Computing; 2021.
26. Wickham H. Data analysis. In: Wickham H, editor. ggplot2: Elegant graphics for data analysis. New York: Springer-Verlag; 2016. p. 189-201.
https://doi.org/10.1007/978-3-319-24277-4_9
[DOI]
27. Oksanen J, Blanchet FG, Friendly M, Kindt R, Legendre P, McGlinn D, Minchin PR, O’Hara B, Simpson GL, Solymos P, et al. vegan: Community ecology package. Community Ecology Package; 2020. p. 1-263.
28. Visagie CM, Houbraken J, Frisvad JC, Hong SB, Klaassen CHW, Perrone G, Seifert KA, Varga J, Yaguchi T, Samson RA. Identification and nomenclature of the genus Penicillium. Stud Mycol 2014;78:343-71. https://doi.org/10.1016/j.simyco.2014.09.001
[DOI]
29. Woudenberg JHC, Seidl MF, Groenewald JZ, de Vries M, Stielow JB, Thomma BPHJ, Crous PW. Alternaria section Alternaria: Species, formae speciales or pathotypes? Stud Mycol 2015;82:1-21. https://doi.org/10.1016/j.simyco.2015.07.001
[DOI]
30. Sandoval-Denis M, Gené J, Sutton DA, Wiederhold NP, Cano-Lira JF, Guarro J. New species of Cladosporium associated with human and animal infections. Persoonia 2016;36:281-98. https://doi.org/10.3767/003158516X691951
[DOI]
31. Ko YM, Lee D, Seok MS, Kwak YS. Diversity of fungal genera associated with shot-hole disease in cherry blossoms across Korea. Mycobiology 2025;53:495-506. https://doi.org/10.1080/12298093.2025.2522491
[DOI]
32. Kim MG, Lim SK, Choi JW, Goh J, Lee CS, Mun HY, Lee SY, Jung HY. Identification and characterization of five previously unrecorded Penicillium species of subgenus Aspergilloides isolated in Korea. Mycobiology 2025;53:731-46. https://doi.org/10.1080/12298093.2025.2550814
[DOI]
33. Lee W, Kim JS, Jo S, Lim YW. Expanding the inventory of seven unrecorded marine Penicillium with morphological descriptions and phenotypic variability. Mycobiology 2025;53:648-60. https://doi.org/10.1080/12298093.2025.2536911
[DOI]
34. Jalajakshi S, Anagha J, Buwaneshwari S, Kalpana R, Poojitha PL. Study on diversity of fungus associated with Nephila pilipes. Plant Health Arch 2023;1:14-7. https://doi.org/10.54083/PHA/1.2.2023/14-17
[DOI]
35. Del Fiol F, Tosi S, Groppali R. Fungal spores and pollen as potential nutritional additives for the cross spider Araneus diadematus Clerck (Araneae, Araneidae). Bol Micol 2007;22:47-50. https://doi.org/10.22370/bolmicol.2007.22.0.132
[DOI]
36. Al Abboud MA, Al-Rajhi AM, Shater ARM, Alawlaqi MM, Mashraqi A, Selim S, Al Jaouni SK, Abdelghany TM. Halostability and thermostability of chitinase produced by fungi isolated from salt marsh soil in subtropical region of Saudi Arabia. BioResources 2022;17:4763-80. https://doi.org/10.15376/biores.17.3.4763-4780
[DOI]
37. Xie XH, Fu X, Yan XY, Peng WF, Kang LX. A broad-specificity chitinase from Penicillium oxalicum k10 exhibits antifungal activity and biodegradation properties of chitin. Mar Drugs 2021;19:356. https://doi.org/10.3390/md19070356
[DOI]
38. Barbosa RN, Felipe MTC, Silva LF, Silva EA, Silva SA, Herculano PN, Prazeres JFSA, Lima JMS, Bezerra JDP, Moreira KA, et al. A review of the biotechnological potential of cave fungi: A toolbox for the future. J Fungi 2025;11:145. https://doi.org/10.3390/jof11020145
[DOI]
39. Romero-Hernández L, Betanzo-Gutiérrez I, Vazquez-Duhalt R, Figueroa M, Riquelme M, Velez P. Degradation of extra-heavy crude oil by fungi isolated from hydrothermal vents fields in the Gulf of California. Mycobiology 2025;53:684-95. https://doi.org/10.1080/12298093.2025.2545629
[DOI]
40. Meyling NV, Eilenberg J. Ecology of the entomopathogenic fungi Beauveria bassiana and Metarhizium anisopliae in temperate agroecosystems: Potential for conservation biological control. Biol Control 2007;43:145-55. https://doi.org/10.1016/j.biocontrol.2007.07.007
[DOI]
41. Joseph RA, Masoudi A, Valdiviezo MJ, Keyhani NO. Discovery of Gibellula floridensis from infected spiders and analysis of the surrounding fungal entomopathogen community. J Fungi 2024;10:694. https://doi.org/10.3390/jof10100694
[DOI]
42. Cha Y, Oh SY. Fungal diversity associated with Armadillidium isopods: A case study in Central Park of Gwacheon, South Korea. Diversity 2023;15:533. https://doi.org/10.3390/d15040533
[DOI]
43. Noh Y, Cha Y, Oh SY. Comparative analysis of the mycobiomes of two terrestrial isopods, Armadillidium vulgare and Spherillo obscurus. Mycobiology 2025;53:520-30. https://doi.org/10.1080/12298093.2025.2524925
[DOI]
44. Oh SY, Jang Y. First report of Metarhizium viridulum isolated from the cicada (Cryptotympana atrata) in South Korea. Kor J Mycol 2024;52:413-7. https://doi.org/10.4489/kjm.520419
[DOI]
45. Oh SY, Cha Y. Influence of host sex on mycobiome structure in Porcellio laevis using metabarcoding analysis. Kor J Mycol 2024;52:331-40. https://doi.org/10.4489/kjm.520412
[DOI]
46. Helaly SE, Kuephadungphan W, Phongpaichit S, Luangsa-ard JJ, Rukachaisirikul V, Stadler M. Five unprecedented secondary metabolites from the spider parasitic fungus Akanthomyces novoguineensis. Molecules 2017;22:991. https://doi.org/10.3390/molecules22060991
[DOI]
47. Kuephadungphan W, Macabeo APG, Luangsa-ard JJ, Stadler M. Discovery of novel biologically active secondary metabolites from Thai mycodiversity with anti-infective potential. Curr Res Biotechnol 2021;3:160-72. https://doi.org/10.1016/j.crbiot.2021.05.003
[DOI]