Yeon-Su Jeong1, Seong-Keun Lim1, Seung-Yeol Lee1,2, and Hee-Young Jung1,2,*
1Department of Plant Medicine, Kyungpook National University, Daegu 41566, Korea
2Institute of Plant Medicine, Kyungpook National University, Daegu 41566, Korea
*Correspondence to heeyoung@knu.ac.kr
Korean Journal of Mycology (Kor J Mycol) 2026 March, Volume 54, Issue 1, pages 1-9.
https://doi.org/10.4489/kjm.2026.54.1.1
Received on January 20, 2026, Revised on March 01, 2026, Accepted on March 04, 2026, Published on March 31, 2026.
Copyright © The Korean Society of Mycology.
This is an Open Access article which is freely available under the Creative Commons Attribution-Non-Commercial 4.0 International License (CC BY-NC) (https://creativecommons.org/licenses/by-nc/4.0/).
Strain KNUF-20-NI020 was recovered from soil collected during a survey of soil fungi in Korea in Gunwi-eup, Gunwi-gun, Daegu. The strain formed pale luteous colonies with rust-colored reverses on potato dextrose agar, reaching a diameter of 25.2 ± 1.0 mm after incubation for 14 days. Mycelial growth was most pronounced at 25°C and was relatively higher at pH values ranging from 6 to 8. For molecular identification, multiple gene regions were analyzed. The internal transcribed spacer (ITS) region showed 99.8% sequence similarity to Keissleriella quadriseptata, and the large subunit ribosomal RNA (LSU), small subunit ribosomal RNA (SSU), and translation elongation factor 1-alpha (TEF1-α) gene sequences were also most similar to those of K. quadriseptata. Phylogenetic analysis based on concatenated ITS, LSU, SSU, and TEF1-α sequences placed the KNUF-20-NI020 strain in the same clade as K. quadriseptata, clearly separated from other related species. Based on the combined cultural characteristics and multilocus molecular phylogenetic analyses, the KNUF-20-NI020 strain was identified as K. quadriseptata. This study represents the first report of K. quadriseptata from Korea and the first record of the genus Keissleriella in Korea.
Keissleriella quadriseptata, Molecular phylogeny, Morphological analysis, Soilinhabiting fungi
The order Pleosporales comprises more than 300 genera and is commonly found in saprobic habitats, such as on dead plant material, and as parasites on plant leaves and stems. Traditionally, identification of this order has relied on the morphological characteristics of sexual structures, particularly asci and ascospores [1]. The type species of the genus Keissleriella is Keissleriella aesculi, which was first described in 1919 [2]. Meanwhile, approximately 50 species have been reported in this genus [3]. A major morphological characteristic of Keissleriella is that the ostiolar neck of the associated ascomata is covered with short dark setae [4].
Species identification within Keissleriella has traditionally relied on asci morphology. Indeed, Trichometasphaeria was previously distinguished from Keissleriella mainly through host specificity or morphological differences; however, recent phylogenetic analyses have led to the integration of these genera [4]. Likewise, the genus Pleurophoma was originally identified based on the observed anamorphic form; however, phylogenetic studies revealed that Pleurophoma belongs to the Keissleriella clade and is now regarded as the anamorph of Keissleriella. Accordingly, Keissleriella has become the preferentially used name [3].
Members of Keissleriella have been isolated from woody plants such as Populus tremula and Rosa canina, as well as from herbaceous plants, including Setaria faberii and Typha latifolia [5,6]. Keissleriella species have also been reported from soil samples [3]. Notably, an unidentified fungal isolate from a freshwater environment in Korea was reported to be closely related to Keissleriella [7]; however, an official record of Keissleriella in Korea has not been confirmed. Furthermore, polysaccharides isolated from some Keissleriella strains have been reported to enhance antioxidant enzyme activity in cellular systems, suggesting a potential applicability for these strains in biotechnology and related industries [8].
Therefore, this study aimed to survey fungal diversity in Korean soils and to isolate the KNUF-20-NI020 strain, which was identified using a combination of morphological features and molecular phylogenetic data.
Soil samples were collected in Gunwi-eup, Gunwi-gun, Daegu, Republic of Korea (36°11′31.4″N, 128°33′29.5″E), from which the KNUF-20-NI020 strain was isolated. Fungal isolation involved serial dilution of soil samples, plating 100 μL aliquots onto potato dextrose agar (PDA; Difco, Detroit, MI, USA), and incubation in the dark at 25°C for 14 days [9]. Representative colonies were subcultured on fresh PDA to obtain pure cultures. Preliminary identification was based on colony morphology and sequence data from the internal transcribed spacer (ITS) regions. Based on these observations, the KNUF-20-NI020 strain was tentatively considered an unrecorded species in Korea. This strain (NIBRFGC000507888) has been deposited in a metabolically inactive state at the National Institute of Biological Resources (NIBR), Republic of Korea.
KNUF-20-NI020 was maintained on PDA in the dark at 20°C and 25°C for four weeks. Colony morphology was recorded photographically using a Canon EOS 5D Mark III digital camera (Canon, Tokyo, Japan). Growth responses were examined under various temperature conditions and different culture media: 10, 15, 20, 25, 30, and 37°C, and PDA, nutrient agar (NA; Difco, Detroit, MI, USA), malt extract agar (MEA; Sigma-Aldrich, St. Louis, MO, USA), Sabouraud dextrose agar (SDA; MBcell, Seoul, Korea), and Czapek-Dox agar (CZA; Sigma-Aldrich, St. Louis, MO, USA) for 15 days. To evaluate the effect of pH, PDA was adjusted to a pH range of 4–9 and incubated at 25°C for 15 days. For sporulation, the strain was cultivated on rice straw medium in the dark at 20°C [4]. Microscopic features were examined using a BX50 light microscope (Olympus, Tokyo, Japan).
Genomic DNA was extracted from KNUF-20-NI020 using the HiGene Genomic DNA Prep kit (BIOFACT, Daejeon, South Korea) according to the manufacturer’s protocol. The ITS regions were amplified using primers ITS1F and ITS4 [10], the large subunit ribosomal RNA (LSU) gene with primers LR0R and LR7 [11], the small subunit ribosomal RNA (SSU) gene with primers NS1 and NS4 [10], and the translation elongation factor 1-alpha (TEF1-α) gene with primers EF1-983F and EF1-2218R [12]. Amplicons were purified using ExoSAP-IT reagent (Thermo Fisher Scientific, Waltham, MA, USA). Sequencing services were provided by Macrogen (Seoul, South Korea). The resulting ITS, LSU, SSU, and TEF1-α sequences were deposited in GenBank under accession numbers LC890403 (ITS), LC890404 (LSU), LC890405 (SSU), and LC890406 (TEF1-α), respectively.
Reference sequences related to the KNUF-20-NI020 strain were identified through BLAST searches against the GenBank database (Table 1). Alignments were performed in ClustalX version 2.0. Phylogenetic relationships were inferred based on a concatenated dataset of ITS, LSU, SSU, and TEF1-α sequences using the maximum likelihood (ML) method implemented in MEGA version 7.0 [13]. Branch support was evaluated with 1000 bootstrap replicates, and genetic distances were calculated under the Kimura two-parameter model [14].
Table 1. GenBank accession numbers for the sequences used in the phylogenetic analyses in this study
| Species name | Strain | GenBank accession numbers | |||
|---|---|---|---|---|---|
| ITS | LSU | SSU | TEF1-α | ||
| Keissleriella breviasca | KT 649T | AB811455 | AB807588 | AB797298 | AB808567 |
| Keissleriella camporesiana | MFLUCC 15-0029T | MN401745 | MN401741 | MN401743 | MN397907 |
| Keissleriella cirsii | MFLUCC 16-0454T | KY497783 | KY497780 | KY497782 | KY497786 |
| Keissleriella culmifida | KT 2642 | LC014562 | AB807592 | AB797302 | AB808571 |
| Keissleriella gloeospora | KT 829 | LC014563 | AB807589 | AB797299 | AB808568 |
| Keissleriella poagena | KUNCC 25-19140 | PV608542 | PV607392 | PV607289 | PV626435 |
| Keissleriella quadriseptata | CBS 139692T | AB811456 | AB807593 | AB797303 | AB808572 |
| Keissleriella quadriseptata | KNUF-20-NI020 | LC890403 | LC890404 | LC890405 | LC890406 |
| Keissleriella taminensis | KT 571 | LC014564 | AB807595 | AB797305 | AB808574 |
ITS: internal transcribed spacer regions; LSU: partial sequence of large subunit 28S rDNA; SSU: partial sequence of small subunit 18s rDNA; TEF1-α: translation elongation factor 1-alpha.
The newly generated sequences are indicated in bold. TType strain.
After incubation on PDA for 14 days, the KNUF-20-NI020 colonies reached a diameter of 25.2 ± 1.0 mm (n = 3). The colony was pale luteous, and the colony reverse ranged from pale luteous to rust-colored. The rust-colored coloration on the reverse was more pronounced at 25°C than at 20°C. No pigment was produced (Fig. 1A–D). Colony growth was observed after 15 days of incubation under various culture media and temperature conditions. Growth occurred at temperatures ranging from 10 to 25°C, whereas no growth was observed at 37°C. Meanwhile, limited growth was detected only on CZA medium at 30°C. The largest colony diameters were recorded at 25°C across all tested media. The maximum growth was observed on NA medium, reaching 28.5 ± 1.1 mm (n = 3), while comparable levels of growth were noted on the other media (Fig. 2A). In the pH-dependent growth experiment, the largest colonies were observed at pH 8, with a diameter of 28.6 ± 0.8 mm (n = 3). Colony growth was similar within the pH range of 6–8. Overall, relatively higher growth was observed under neutral to slightly alkaline conditions (pH 6–8) (Fig. 2B). Hyphae were hyaline, branched, and septate, with a width of 2.3 ± 0.5 μm (n = 30) (Fig. 1E, F). Ascospores and asexual spores were not observed in the cultures grown on PDA or rice straw. While the colony characteristics of the KNUF-20-NI020 strain were generally similar to those of K. quadriseptata (CBS 139692T), differences in colony color and growth rate were observed when compared with other closely related Keissleriella species (Table 2).
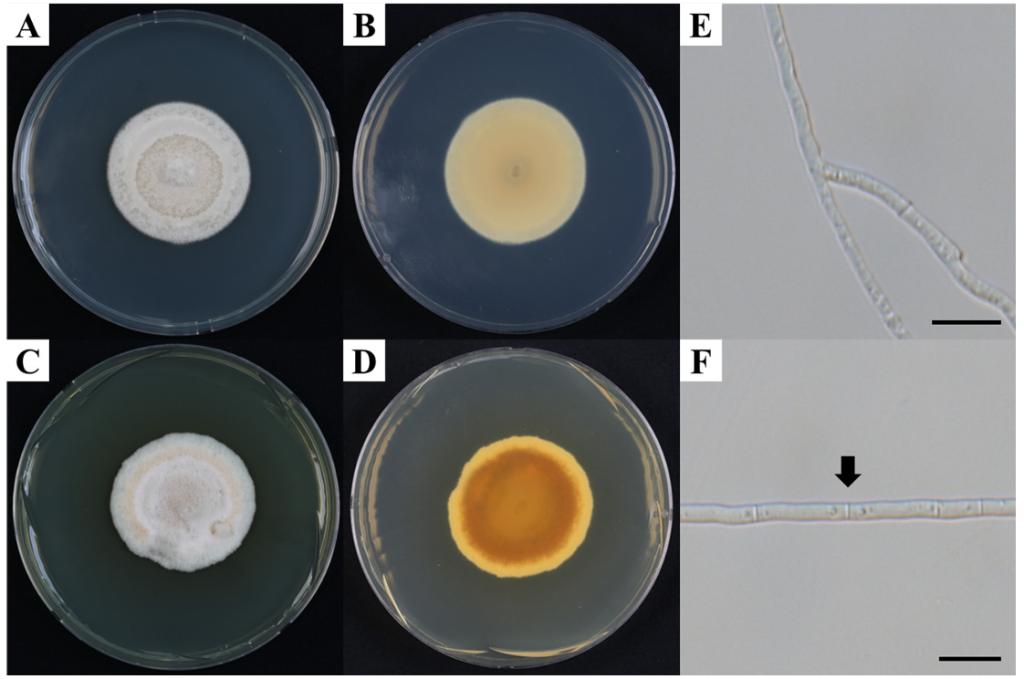
Fig. 1. Cultural and morphological characteristics of KNUF-20-NI020 (Keissleriella quadriseptata). A, B: Colony on potato dextrose agar (PDA) at 20°C (A, surface; B, reverse); C, D: Colony on PDA at 25°C (C, surface; D, reverse); E, F: Hyphae. Arrows indicate septa. Scale bars = 10 μm.
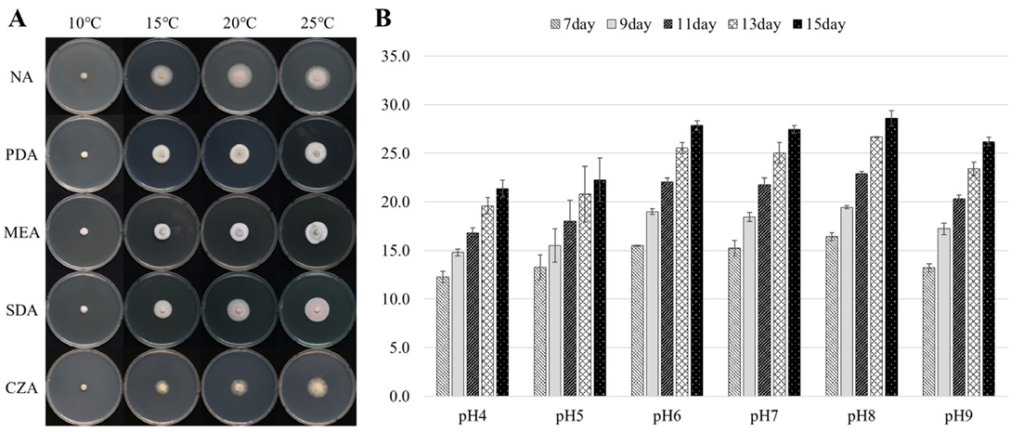
Fig. 2. Dependence of colony diameter and density on growth conditions. A: Influence of temperature and media; B: Influence of pH of the potato dextrose agar (PDA) medium on the growth of culture at 25°C. NA: nutrient agar; PDA: potato dextrose agar; MEA: malt extract agar; SDA: Sabouraud dextrose agar; CZA: Czapek-Dox agar.
Table 2. Comparison of cultural characteristics of the KNUF-20-NI020 strain and phylogenetically closely related Keissleriella species
| Characteristics | K. quadriseptata KNUF-20-NI020a |
K. quadriseptata CBS 139692Tb |
K. gloeospora UESTCC24.0217c |
K. breviasca CBS 139691Tb |
K. culmifida KT 2308b |
K. poagena CBS 136767Td |
|
|---|---|---|---|---|---|---|---|
| Colony | Color | PDA: pale luteous; reverse rust; no pigment produced MEA: white to greyish white; reverse pale yellow; no pigment produced |
PDA: pale luteous; reverse rust; no pigment produced | PDA: pale brown-milky white; reverse pale brown-milky white | PDA: white margin; reverse red to flesh; sienna pigment produced | N/A | PDA: smoke-grey; reverse buff MEA: pale olivaceous-grey; reverse umber |
| Growth | 11.5 mm at 20°C for 1 week 13.3 mm at 25°C for 1 week 25.2 mm at 20°C for 2 weeks 26.3 mm at 25°C for 2 weeks |
17–22 mm at 20°C for 2 weeks | 12–14 mm at 25°C for 1 week | 10–13 mm at 20°C for 4 weeks | N/A | 15 mm at 25°C for 2 weeks | |
PDA: potato dextrose agar; MEA: malt extract agar; N/A: not available.
aFungal strain used in this study; bSource of description [4]; cSource of description [17]; dSource of description [18]. TType strain.
The ITS, LSU, SSU, and TEF1-α sequence lengths for the KNUF-20-NI020 strain were 553 bp, 1303 bp, 1007 bp, and 921 bp, respectively. BLAST analysis of the ITS sequence in the NCBI database showed 99.8% similarity to Keissleriella quadriseptata (CBS 139692; NR_145135), 99.2% similarity to K. gloeospora (U19; PV394908), and 97.1% similarity to K. trichophoricola (CBS 136770; NR_156276). BLAST analysis of the LSU sequence revealed 99.7% similarity to K. quadriseptata (KT 2292; NG_059401), 99.6% similarity to K. gloeospora (KT 829; AB807589), and 99.5% similarity to K. culmifida (KT 2308; AB807591). The SSU sequence showed 99.9% similarity to K. quadriseptata (KT 2292; NG_064855), 98.8% similarity to K. rosarum (MFLUCC 15-0089; NG_063685), and 98.8% similarity to Pleurophoma pleurospora (TASM 6115; MG829159). The TEF1-α sequence showed 99.5% similarity to K. quadriseptata (KT 2292; AB808572), 97.7% similarity to K. gloeospora (KT 829; AB808568), and 96.3% similarity to Phragmocamarosporium hederae (KUMCC 18-0165; MK214378).
To determine the phylogenetic position of KNUF-20-NI020, a concatenated dataset of ITS, LSU, SSU, and TEF1-α sequences was analyzed using the ML method. The resulting phylogenetic tree showed that KNUF-20-NI020 clustered with K. quadriseptata and was clearly separated from K. gloeospora. Therefore, KNUF-20-NI020 was identified as K. quadriseptata (Fig. 3).
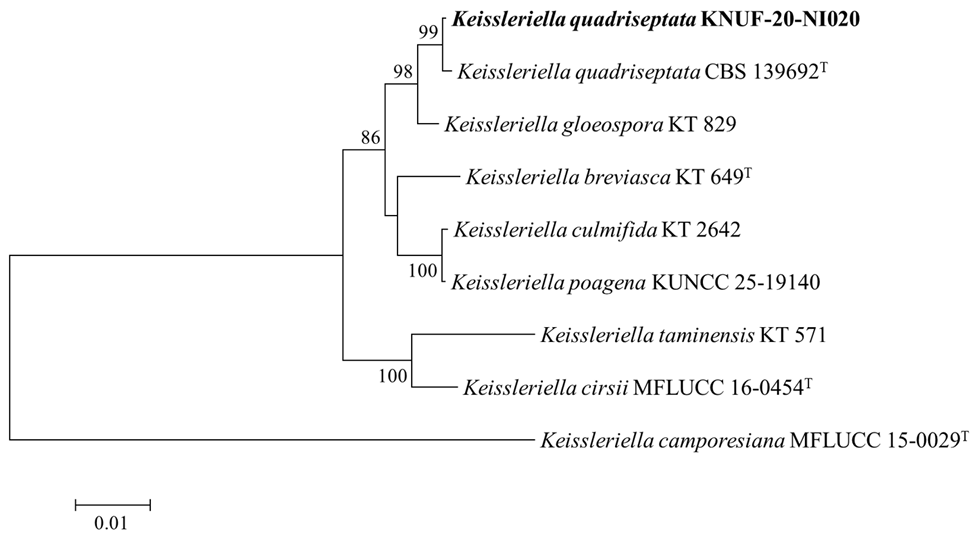
Fig. 3. Maximum likelihood phylogenetic tree based on the internal transcribed spacer (ITS), large subunit ribosomal RNA (LSU), small subunit ribosomal RNA (SSU), and translation elongation factor 1-alpha (TEF1-α) sequences that depicts the phylogenetic position of KNUF-20-NI020 within the genus Keissleriella. K. camporesiana MFLUCC 15-0029T was used as the outgroup. The isolate from this study is highlighted in bold. The numbers above the branches indicate bootstrap values (> 80%) from 1,000 replicates. Scale bar = 0.01 substitutions per nucleotide position.
The order Pleosporales has traditionally been identified based on morphological characteristics of the ascomata and ascospores, such as perithecioid ascomata, papillate apex, cellular pseudoparaphyses, fissitunicate asci, and diverse ascospore morphology and septation [1]. Likewise, the genus Keissleriella has historically been delineated based on sexual morphological features, including ascus size, ascospore morphology, and number of septa. For instance, K. breviasca is characterized according to the noted short asci [4], while K. dactylidis is distinguished by ellipsoid to cylindrical ascospores and relatively small ascomata [6]. More recently, both sexual and asexual morphological features have been utilized in species identification. For example, K. quadriseptata is defined by the observed four-septate ascospores and pycnidial asexual morph [4], while K. phragmiticola forms cylindrical to bone-shaped hyaline conidia [15]. However, the introduction of molecular phylogenetic analyses has shown that morphology-based identification alone is insufficient for accurate classification of both Pleosporales and Keissleriella. Initially, Pleosporales was circumscribed by Barr (1987) to include 22 families based on morphology; however, subsequent molecular studies have proposed various family counts: 13 by Eriksson (2006), 19 by Kirk et al. (2001), and 23 by Kirk et al. (2008) [1]. Likewise, the taxonomy of Keissleriella has undergone substantial revision. Subsequently, species such as Pleurophoma pleurospora, P. ossicola, and P. acaciae have been reclassified as Keissleriella based on phylogenetic evidence indicating that these species represent asexual morphs of this genus [3]. Therefore, morphological characteristics alone are insufficient for accurate species delimitation within Keissleriella, and multilocus molecular phylogenetic analyses are essential for reliable taxonomic resolution, particularly using multiple genetic loci such as ITS, LSU, SSU, and TEF1-α [3,4,15]. Sexual and asexual morphs of strain KNUF-20-NI020 were not observed under the examined culture conditions, limiting morphological comparison to cultural characteristics. In the present study, species identification relied primarily on multilocus molecular phylogenetic analyses. Phylogenetic reconstruction revealed that strain KNUF-20-NI020 formed a well-supported clade with K. quadriseptata (bootstrap = 99%). Based on these molecular data, the strain was determined to be K. quadriseptata.
The genus Keissleriella has been predominantly reported from dried stems of herbaceous plants; however, some species have also been found on the leaves of woody plants, in aquatic plants, and in soil. For example, K. breviasca and K. quadriseptata have reportedly been identified from dried culms of Dactylis glomerata [4], while K. yunnanensis and K. pleurospora were isolated from twigs and leaves of woody plants, respectively [3]. K. yonaguniensis and K. phragmiticola were found in wetland-associated plants [4,15], and K. ossicula was reported from sandy soil near pine trees [3]. The KNUF-20-NI020 strain was also isolated from pine rhizosphere soil. This additional soil-derived isolate further supports the view that Keissleriella occupies a broader ecological range than previously recognized. In Korea, a species presumed to belong to the genus Keissleriella was reported from the freshwater environment of the Nam River in Jinju; however, the species was not identified at the species level due to the lack of morphological and molecular phylogenetic data [7]. Therefore, to our knowledge, this study represents the first confirmed record of Keissleriella in Korea.
Industrial applications of the genus Keissleriella remain limited. However, EPS2, a polysaccharide produced by Keissleriella sp. YS4108, isolated from the Yellow Sea, has demonstrated antioxidant and neuroprotective effects, suggesting potential medical applications in the treatment of neurodegenerative diseases [8]. Furthermore, species of Keissleriella are frequently found in diverse environments, particularly on decomposing plant debris. Although studies on plant-degrading enzymes in this genus are scarce, members of the order Pleosporales have been shown to possess various cell wall-degrading enzymes [16]. This indicates the potential for Keissleriella to be utilized as a biodegrader of plant-derived organic matter. Therefore, Keissleriella may have various industrial applications, warranting further investigation.
The identification of the KNUF-20-NI020 strain confirms the presence of Keissleriella in Korean soil environments, thereby expanding the known fungal diversity of Korean soil ecosystems. Moreover, considering the potential industrial applications of this genus, these findings provide valuable baseline data for future ecological and applied mycological research.
The authors declare that they have no potential conflicts of interest.
This study was supported by the National Institute of Biological Resources, funded by the Ministry of Environment of the Republic of Korea [NIBR202002104].