1RPTU Kaiserslautern-Landau, Department of Biology, Kaiserslautern 67663, Germany
2Genomics & Transcriptomics Labor (BMFZ), Heinrich-Heine-University Düsseldorf, Düsseldorf 40225, Germany
3Research Institute for Forest Ecology and Forestry Rheinland-Pfalz, Trippstadt 67705, Germany
*Correspondence to saeed@rptu.de
Korean Journal of Mycology (Kor J Mycol) 2026 March, Volume 54, Issue 1, pages 11-17.
https://doi.org/10.4489/kjm.2026.54.1.2
Received on November 19, 2025, Revised on February 02, 2026, Accepted on March 11, 2026, Published on March 31, 2026.
Copyright © The Korean Society of Mycology.
This is an Open Access article which is freely available under the Creative Commons Attribution-Non-Commercial 4.0 International License (CC BY-NC) (https://creativecommons.org/licenses/by-nc/4.0/).
Powdery mildew caused by Erysiphe alphitoides is a persistent constraint on European oaks, yet the host responses in pedunculate oak (Quercus robur) that determine susceptibility or resistance remain poorly understood. We profiled transcriptomic changes at 30 hours post-inoculation in detached Q. robur leaves using RNA-seq, identifying 195 up- and 40 down-regulated genes. Up-regulated transcripts included receptor-like kinases (RLKs), ethylene-responsive factor (ERF)/ethylene components, calmodulin-like (CML) sensors, germin-like proteins, peroxidases, and phenylpropanoid pathway enzymes. Gene Ontology enrichment highlighted defense response and response to biotic stimulus, and Kyoto Encyclopedia of Genes and Genomes indicated plant–pathogen interaction. RT-qPCR experiments confirmed ERF017, mUCP5, CML37-like, and one of two CML19 prime targets. These data indicate the induction of RLK-linked, Ca2+/CML- and ethylene-associated defense response during early powdery mildew infection and suggest targeted time-course and functional tests of RLK/CML candidates.
Calmodulin-like protein, Erysiphe alphitoides, ethylene, Quercus robur, RNA-seq
Powdery mildew is among the most widespread foliar diseases of broadleaf trees. In the two European oak species, pedunculate oak (Quercus robur) and sessile oak (Q. petraea), Erysiphe alphitoides is the main cause of oak powdery mildew (OPM) [1–3]. Recurrent epidemics of OPM reduce photosynthetic capacity and lead to stunted growth of seedlings, thereby increasing management costs in nurseries and restoration plantings [4–6]. Oaks are of great ecological and economic importance across Europe [7], yet the molecular events that lead to OPM resistance or susceptibility, particularly within the early phase of contact, are poorly understood [8–10]. Plant immunity relies on rapid pathogen perception and signaling [11]. At the cell surface, pattern-recognition receptors and receptor-like kinases (RLKs) detect microbe-associated signals and trigger downstream cascades [12]. Canonical early outputs include mitogen-activated protein kinase activation and cytosolic Ca²+ fluxes decoded by calmodulin-like proteins (CMLs), transcriptional reprogramming via WRKY/ ethylene-responsive factors (ERFs) and cell-wall reinforcement with reactive oxygen species metabolism [13]. Hormonal crosstalk among salicylic acid, jasmonic acid, and ethylene modulates these responses in a context-dependent manner [14]. Phenylpropanoid biosynthesis contributes to both structural (lignin-related) and defensive (antimicrobial) metabolites [15]. In long-lived woody hosts, these processes may be temporally shifted or prolonged relative to annuals, underscoring the value of time-resolved studies in trees [16]. Despite these advances, molecular studies capturing the early responses of Q. robur to E. alphitoides remain scarce. Prior work has largely emphasized visible symptoms, priming effects, or later transcriptional adjustments in oak [8,9], and notably the transcriptomic study of Streptomyces-induced resistance by Kurth et al. [10] focused on systemic responses rather than the immediate signaling events that follow pathogen contact. However, identifying these early signaling pathways is essential for designing durable and sustainable management strategies against powdery mildew.
In this study, we present the results of gene expression changes in Q. robur detached leaves 30 hours after inoculation with E. alphitoides, to capture early host reprogramming events. Our results provide an early snapshot of the gene expression map of the oak-PM interaction that will guide future functional assays and inform selection of markers for screening pedigrees or provenances.
Pedunculate oak (Q. robur) seedlings were raised from acorns under controlled conditions. Healthy, fully expanded leaves were collected from independent seedlings of similar age and size prior to inoculation. Leaves were placed in sterile 90-100 mm Petri dishes lined with moistened tissue. For each condition (OPM-inoculated, mock), three biological replicates were used, each consisting of two leaves. Detached Q. robur leaves were inoculated on the adaxial surface with 2.5 µL droplets containing ~150 conidia of E. alphitoides or a water mock control, ensuring broad coverage of the leaf area [17]. Leaves were harvested at 30 hours post-inoculation (hpi) to capture early host transcriptional signaling following pathogen perception, after initial penetration but prior to extensive colonization or symptom development. For RNA-seq, we used three biological replicates per condition, each consisting of two pooled leaves. Total RNA was extracted, DNase-treated, poly(A)-selected, and sequenced (2 × 150 bp; NextSeq 2000; Illumina, Inc., San Diego, CA, USA). Reads were quality-checked, adapter- and quality-trimmed, and aligned to the Q. robur v2 reference genome [18] using CLC Genomics Workbench v24.0.1 (Qiagen, Hilden, Germany) with default RNA-seq analysis parameters. Gene expression was quantified at the gene level and normalized as transcripts per million values. Differential expression analysis was performed using a negative binomial model with Benjamini–Hochberg false discovery rate (FDR) correction; genes with |log2 fold change (FC)| ≥ 1 and FDR ≤ 0.05 were considered significant. Isoforms were collapsed to the longest transcript per gene prior to downstream analyses. The collapsed gene set comprised 12,946 genes. Evolutionary genealogy of genes by Non-supervised Orthologous Groups (eggNOG)-mapper v2 [19] returned annotations for 12,641 genes (Table S1). Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) over-representation analyses (one-sided Fisher’s exact test; BH-FDR) used the annotated subsets: GO universe N = 5,717 genes with ≥1 GO term, and KEGG universe N = 5,819 genes with ≥1 K number: Gene-level eggNOG annotations for the differentially expressed gene (DEG) sets are provided in Table S2 (up-regulated) and Table S3 (down-regulated).
Six up-regulated RNA-seq targets (five DEGs with |log2 FC| ≥ 1 and FDR ≤ 0.05, plus one up-regulated candidate with FDR > 0.05) were selected based on (i) statistical significance and pathway coverage across RLK/Ca²+/ethylene modules and (ii) practical measurability; ensuring significant expression in mock controls to avoid zero-inflated fold change artifacts. Two CML19 loci were assayed with distinct primer pairs: CML19-81 (LOC126712681) and CML19-84 (LOC126712684). Relative expression was calculated by the Pfaffl method [20] using GAPDH and Actin as references; significance was assessed with two-sided Student’s t-tests. Primer sequences are provided in Table S4.
We profiled the leaf transcriptome of Q. robur at 30 hpi (n = 3 mock: S13–S15, n = 3 OPM: S16–S18). A principal component analysis of log2 (counts per million (CPM)+1) values showed separation between samples (PC1 = 34.9%, PC2 = 31.4%; Fig. 1A), indicating that condition effects dominated. The volcano plot (Fig. 1B) was shifted toward positive log2FC values, indicating an induction-biased response. With thresholds |log2 FC| ≥ 1 and FDR ≤ 0.05, we identified 195 up-regulated and 40 down-regulated genes. Effect sizes among up-regulated DEGs were moderate to strong (median log2 FC = 2.18; IQR 1.52–3.09; maximum = 6.62). A heat map highlighted a dominant induction module across inoculated samples with a comparatively smaller set of repressed transcripts (Fig. 1C). Gene-level eggNOG annotations for these DEGs are listed in Tables S2–S3.
We performed over-representation analysis separately on up- and down-regulated DEGs using a onesided Fisher’s exact test with Benjamini–Hochberg correction, against the GO-annotated background for GO terms and the KEGG-annotated background for KEGG pathways (ranked visualizations), shown in Fig. S1. Among up-regulated genes, enrichment clustered in defense-related Biological Process terms—most prominently defense response, response to chitin, defense response to other organism, and aligned response to biotic stimulus/wounding—with little signal in Molecular Function and sparse Cellular Component terms pointing to cell wall/external encapsulating structure, consistent with apoplastic/wall-associated defenses at this timepoint. For the downregulated DEGs, nominal enrichments in specialized metabolism and auxin transport were detected; however, none remained significant after FDR correction (q > 0.05), and the large apparent fold-enrichments reflect small background annotation counts (K). A summary of the major GO enrichment patterns is shown in Table 1; ranked visualizations are in Fig. S1A (GO) and Fig. S1B (KEGG); full outputs in Tables S5–S8; per-gene (one-per-LOC protein) eggNOG annotations for the DEG sets in Tables S2–S3.
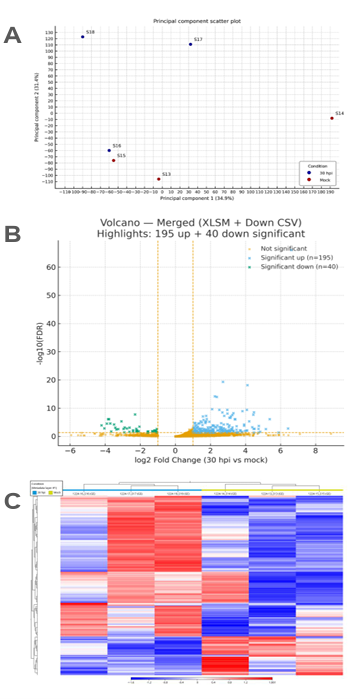
Fig. 1. Global transcriptional structure and differential expression at 30 h post-inoculation (hpi). (A) Principal component analysis of log2(CPM+1) expression for all genes (n = 6 libraries: 3 mock, 3 OPM; samples S13–S18). PC1 = 34.9%, PC2 = 31.4% of variance. Points are colored by treatment and labeled with sample IDs. (B) Volcano plot for 30 hpi vs. mock showing log2 fold change (x-axis) versus − log10(FDR) (y-axis). Vertical lines mark ±1 log2FC, and the horizontal line marks FDR = 0.05 (BenjaminiHochberg). Genes passing both thresholds were called DEGs (195 up-regulated, 40 down-regulated). (C) Heat map of the 235 DEGs (|log2FC| ≥ 1; FDR ≤ 0.05). Values are row-wise z-scores of log2(CPM+1). Genes and samples are hierarchically clustered (Euclidean distance, complete linkage), separating OPM from mock and revealing treatment-associated modules (red = higher, blue = lower relative to each gene’s mean). CPM: counts per million; OPM: oak powdery mildew; FDR: false discovery rate; FC: fold change; DEG: differentially expressed gene.
Across two independent RT-qPCR experiments at 30 hpi, four targets—ERF017, mUCP5, CML37like, and one of two CML19 loci, CML19 (84)—showed infection-induced expression consistent with the RNA-seq data (Fig. 2; Table S9). Upregulation of the second CML19 locus was reproduced in one of the two experiments but not confirmed for 1-aminocyclopropane-1-carboxylate oxidase 4.
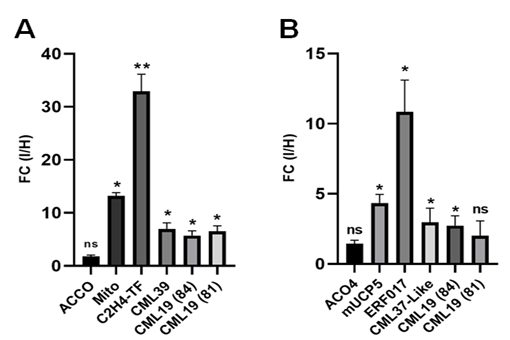
Fig. 2. RT-qPCR validation of 30 h post-inoculation (OPM vs mock). (A) Exp1 (n = 3 per condition; same RNA as used for RNA-seq). (B) Exp2 (n = 5 per condition; independent replications). Bars show mean fold-change (OPM-infected/mock) ± standard deviation computed by the Pfaffl method relative to GAPDH and Actin. Y-axis shows fold change for OPM-infected vs. healthy (I/H). Asterisks indicate two-sided Student’s t-test versus mock (* p < 0.05; ** p < 0.01; ns: not significant). ERF017, mUCP5, CML37-like, and CML19-84 are significant in both experiments; CML19-81 is significant only in (A); ACO4 is not significant. OPM: oak powdery mildew; ERF: ethylene-responsive factor; CML: calmodulin-like; ACO: 1-aminocyclopropane-1-carboxylate oxidase.
Taken together, we identified and validated four genes that are upregulated in Q. robur leaves early after infection with E. alphitoides. These genes could be useful as sensitive molecular markers for OPM infection. Further studies are required to reveal if these genes are involved in the defense against OPM, which seems to be likely based on their predicted functions. In this case, they could be helpful for the identification of Q. robur genotypes that are partially resistant to OPM infection. In addition, more genes from the up- and down-regulated set obtained by RNA-seq can be validated by RT-qPCR, making it a valuable source for infection-related gene mining. Therefore, our study will open further investigations into molecular communication in the oak-OPM pathosystem, and towards the exploration of novel strategies for the protection of oak from this destructive disease.
The following supplementary materials are available on the journal’s website:
· Table S1. Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG)
· Table S2. Functional Annotations of Up-regulated Genes
· Table S3. Functional Annotations of Down-regulated Genes
· Table S4. List of Primer Sequences
· Table S5. Gene Ontology (GO) Enrichment Results for Upregulated Genes
· Table S6. Gene Ontology (GO) Enrichment Results for Down-regulated Genes
· Table S7. Kyoto Encyclopedia of Genes and Genomes (KEGG) Pathway Enrichment Results for Upregulated Genes
· Table S8. Kyoto Encyclopedia of Genes and Genomes (KEGG) Pathway Enrichment Results for Down-regulated Genes
· Table S9. Reverse Transcription Quantitative PCR (RT-qPCR) Validation Results for Infectionresponsive Genes
· Fig. S1. Enrichment Summary for Up-regulated Genes at 30 Hours Post-inoculation.
This project has been funded by the Fachagentur Nachwachsende Rohstoffe e.V. (BMEL), ref. no. 2220WK1684 (2021-2024).
The RNA-seq data generated in this study have been deposited in the NCBI Sequence Read Archive (SRA) under accession number PRJNA1444513.
Marçais B, Desprez-Loustau ML. European oak powdery mildew: Impact on trees, effects of environmental factors, and potential effects of climate change. Ann For Sci 2014;71:633–61. https://doi.org/10.1007/s13595-012-0252-x
Desprez-Loustau ML, Massot M, Toïgo M, Fort T, Kaya AGA, Boberg J, et al. From leaf to continent: The multi-scale distribution of an invasive cryptic pathogen complex on oak. Fungal Ecol 2018;36:39–50. https://doi.org/10.1016/j.funeco.2018.08.001
Feau N, Lauron-Moreau A, Piou D, Marçais B, Dutech C, Desprez-Loustau ML. Niche partitioning of the genetic lineages of the oak powdery mildew complex. Fungal Ecol 2012;5:154–62. https://doi.org/10.1016/j.funeco.2011.12.003
Hajji M, Dreyer E, Marçais B. Impact of Erysiphe alphitoides on transpiration and photosynthesis in Quercus robur leaves. Eur J Plant Pathol 2009;125:63–72. https://doi.org/10.1007/s10658-009-9458-7
Skwarek-Fadecka M, Nawrocka J, Sieczyńska K, Patykowski J, Posmyk MM. Effect of oak powdery mildew on ascorbate–glutathione cycle and other antioxidants in plant–Erysiphe alphitoides interaction. Cells 2024;13:1035. https://doi.org/10.3390/cells13121035
Marković Č, Kanjevac B, Perišić U, Dobrosavljević J. The effect of the oak powdery mildew, oak lace bug, and other foliofagous insects on the growth of young pedunculate oak trees. Front For Glob Change 2024;6:1297560. https://doi.org/10.3389/ffgc.2023.1297560
Annighöfer P, Beckschäfer P, Vor T, Ammer C. Regeneration patterns of European oak species (Quercus petraea (Matt.) Liebl., Quercus robur L.) in dependence of environment and neighborhood. PLoS One 2015;10:e0134935. https://doi.org/10.1371/journal.pone.0134935
Bartholomé J, Brachi B, Marçais B, Mougou-Hamdane A, Bodénès C, Plomion C, et al. The genetics of exapted resistance to two exotic pathogens in pedunculate oak. New Phytol 2020;226:1088–103. https://doi.org/10.1111/nph.16319
Sanchez-Lucas R, Bosanquet JL, Henderson J, Catoni M, Pastor V, Luna E. Elicitor specific mechanisms of defence priming in oak seedlings against powdery mildew. Plant Cell Environ 2025;48:4455–74. https://doi.org/10.1111/pce.15419
Kurth F, Mailänder S, Bönn M, Feldhahn L, Herrmann S, Große I, et al. Streptomyces-induced resistance against oak powdery mildew involves host plant responses in defense, photosynthesis, and secondary metabolism pathways. Mol Plant Microbe Interact 2014;27:891–900. https://doi.org/10.1094/MPMI-10-13-0296-R
Jones JDG, Staskawicz BJ, Dangl JL. The plant immune system: From discovery to deployment. Cell 2024;187:2095–116. https://doi.org/10.1016/j.cell.2024.03.045
Zipfel C. Plant pattern-recognition receptors. Trends Immunol 2014;35:345–51. https://doi.org/10.1016/j.it.2014.05.004
DeFalco TA, Zipfel C. Molecular mechanisms of early plant pattern-triggered immune signaling. Mol Cell 2021;81:3449–67. https://doi.org/10.1016/j.molcel.2021.07.029
Pieterse CMJ, Van der Does D, Zamioudis C, Leon-Reyes A, Van Wees SCM. Hormonal modulation of plant immunity. Annu Rev Cell Dev Biol 2012;28:489–521. https://doi.org/10.1146/annurev-cellbio-092910-154055
Wilson SK, Pretorius T, Naidoo S. Mechanisms of systemic resistance to pathogen infection in plants and their potential application in forestry. BMC Plant Biol 2023;23:404. https://doi.org/10.1186/s12870-023-04391-9
Dong NQ, Lin HX. Contribution of phenylpropanoid metabolism to plant development and plant–environment interactions. J Integr Plant Biol 2021;63:180–209. https://doi.org/10.1111/jipb.13054
Saeed M, Sharma S, Seegmüller S, Hahn M. First report of powdery mildew on Quercus rubra in Germany. Kor J Mycol 2025;53:29–35. https://doi.org/10.4489/kjm.2025.53.1.4
Plomion C, Aury JM, Amselem J, Leroy T, Murat F, Duplessis S, et al. Oak genome reveals facets of long lifespan. Nat Plants 2018;4:440–52. https://doi.org/10.1038/s41477-018-0172-3
Cantalapiedra CP, Hernández-Plaza A, Letunic I, Bork P, Huerta-Cepas J. eggNOG-mapper v2: Functional annotation, orthology assignments, and domain prediction at the metagenomic scale. Mol Biol Evol 2021;38:5825–9. https://doi.org/10.1093/molbev/msab293
Pfaffl MW. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res 2001;29:e45. https://doi.org/10.1093/nar/29.9.e45