Gyo-Bin Lee1, Weon-Dae Cho2, and Wan-Gyu Kim2*
1Plant Disease Control Division, National Institute of Agricultural Sciences, Wanju 55365, Korea
2Global Agro-Consulting Corporation, Suwon 16614, Korea
*Correspondence to wgkim5121@naver.com
Korean Journal of Mycology (Kor J Mycol) 2026 March, Volume 54, Issue 1, pages 33-42.
https://doi.org/10.4489/kjm.2026.54.1.4
Received on January 22, 2026, Revised on March 18, 2026, Accepted on March 20, 2026, Published on March 31, 2026.
Copyright © The Korean Society of Mycology.
This is an Open Access article which is freely available under the Creative Commons Attribution-Non-Commercial 4.0 International License (CC BY-NC) (https://creativecommons.org/licenses/by-nc/4.0/).
During crop disease surveys in October 2022, we encountered leaf spot symptoms in coastal hogfennel (Peucedanum japonicum) grown in a field in Buyeo, Korea. Disease symptom outbreaks in the field ranged from 5 to 10%. The three fungal isolates obtained from leaf lesions were morphologically identified as Didymella sp. Phylogenetic analyses confirmed a close relationship between the isolates and Didymella acutilobae, and the mycological characteristics were generally consistent with those of D. acutilobae. Pathogenicity of the isolates on leaves of coastal hogfennel was confirmed via artificial inoculation test. To date, this is the first report on the onset of D. acutilobae-induced leaf spot in coastal hogfennel.
Coastal hogfennel, Didymella acutilobae, Leaf spot, Peucedanum japonicum
Coastal hogfennel (Peucedanum japonicum Thunb.) is perennial and a member of the family Apiaceae. It is native to countries in East Asia, including China, Korea, Japan, the Philippines, and Taiwan, and typically thrives in temperate biomes [1]. Coastal hogfennel is usually used as an edible wild vegetable in cuisine [2], with its phytochemical constituents having been studied for their potential anti-inflammatory and oxidant activities [3,4].
Phoma is one of the largest fungal genera, encompassing approximately 3,000 taxa that exhibit diverse lifestyles, ranging from saprobic and opportunistic to endophytic and pathogenic across various substrates, including animals, plants, and even inorganic matter [5–8]. The identification of Phoma spp. has undergone significant changes with the development and application of new molecular tools and techniques. Many species in the genus Phoma (anamorphs), identified using traditional methods, have been reclassified within the genus Didymella (teleomorphs) of the family Didymellaceae. As a teleomorph, the genus Didymella produces eight didymospores in cylindrical bitunicate asci in a pseudothecium. Didymellaceae is the largest family, encompassing more than 30 genera, including Ascochyta and Phoma with over 5,400 taxa [9]. Nowadays, integrated identification methods combining traditional approaches with molecular phylogenetic analysis are mandatory and have become routine for the identification of ambiguous species in families.
Leaf spot symptoms were observed in coastal hogfennel plants grown in a field located in Buyeo, Korea, during a survey of crop diseases in October 2022. We investigated fungi isolated from the symptoms and found that they produced conidia within the pycnidia. The morphological features of the pycnidial fungi were generally consistent with those of the genus Phoma [5].
Leaf spot in coastal hogfennel caused by other fungi (Alternaria peucedani and Ascochyta sp.) has been reported in Korea [10]. However, the disease caused by Phoma sp. has not yet been reported. In this study, to identify Phoma sp. isolates from diseased leaves of coastal hogfennel, the multilocus phylogenetic characteristics, morphological, and cultural characteristics of the isolates were analyzed. In addition, pathogenicity of the isolates on coastal hogfennel was assessed to determine the disease-inducing pathogen.
In October 2022, disease surveys of in-field crops were conducted in Buyeo, Korea. During which, we encountered leaf spot symptoms in coastal hogfennel plants grown in a field and collected these diseased leaves. Leaf lesions were cut into small pieces (3 × 3 mm) and immersed in 1% sodium hypochlorite for 1 min for surface sterilization. Surface-sterilized lesion pieces were plated on 2% water agar (WA; FUJIFILM Wako Pure Chemical Corporation, Osaka, Japan). The WA plates were incubated at 25℃ for 3–4 days. Three single-conidium isolates were obtained from pycnidia produced in lesion pieces on the WA and used for identification and pathogenicity tests.
The mycological characteristics of the isolates were investigated in accordance with previously described protocols [5,11]. Malt extracted agar (MEA; Sigma-Aldrich, St. Louis, MO, USA), oatmeal agar (OA; Sigma-Aldrich), and potato dextrose agar (PDA; BD Difco, Sparks, USA) were used to investigate cultural features of the isolates, which were described according to the Rayner color chart [12]. To determine growth rates, the colony diameters of the isolates were measured after 7 days of incubation in quadruplicate. Morphological characteristics were assessed utilizing 14-day-old cultures of the isolates on OA through collecting 30 pycnidia and 30 conidia and using a light microscope (Nikon Eclipse Ci-L, Tokyo, Japan). Additionally, 15 pycnidial sections per isolate were prepared and examined using a previously described method [13]. One-week-old cultures of the isolates on MEA were used for the NaOH spot reactions. All reaction tests were triplicated.
The protocol described by Dong et al. [14] was used to extract genomic DNA from the isolates, with a slight modification: boiling time was shortened. The 28S large subunit of the nrDNA gene (LSU), internal transcribed spacer regions 1 and 2 including 5.8S nrDNA gene (ITS), β-tubulin (TUB2), and RNA polymerase II second largest subunit (RPB2) gene regions of the isolates were amplified using specific primer sets [15‒21] (Table 1) and cycling conditions [8,22]. PCR amplification was conducted using the DNA Free-Taq Master Mix (CellSafe, Yongin, Korea), while DNA purification was performed using the Universal DNA Purification Kit (Tiangen, Beijing, China); both procedures were performed according to the manufacturers’ protocols. The same primers were used for sequencing by Bionics Co., Ltd. (Seoul, Korea). Sequencing data were deposited in the National Center for Biotechnology Information (NCBI) GenBank database.
Table 1. Information of the primer sets used for molecular identification of Didymella sp. isolates from coastal hogfennel
| Gene region | Primer | Sequence (5’–3′) | Reference |
|---|---|---|---|
| LSU | LR0R | ACCCGCTGAACTTAAGC | 16 |
| LR7 | TACTACCACCAAGATCT | 17 | |
| ITS | V9G | TTACGTCCCTGCCCTTTGTA | 18 |
| ITS4 | TCCTCCGCTTATTGATATGC | 19 | |
| TUB2 | Btub2Fd | GTBCACCTYCARACCGGYCARTG | 20 |
| Btub4Rd | CCRGAYTGRCCRAARACRAAGTTGTC | 20 | |
| RPB2 | RPB2-5f2 | GGGGWGAYCAGAAGAAGGC | 21 |
| fRPB2-7cR | CCCATRGCTTGYTTRCCCAT | 22 |
LSU: 28S large subunit of the nrDNA gene; ITS: internal transcribed spacer regions 1 and 2 including the 5.8S nrDNA gene; TUB2: β-tubulin; RPB2: RNA polymerase II second largest subunit.
Isolates were sequenced using SeqMan II (DNASTAR Inc., Madison, WI, USA), and manually adjusted, if necessary. Relevant sequences of Didymella spp. were retrieved from previous studies [23–25]. MUSCLE within the MEGA 7 software [26] was used to evaluate the performance of multiple sequence alignments. The outgroup taxa, Ascochyta koolunga (CBS 373.84) and Ascochyta rabiei (CBS 534.65) were used to construct the phylogenetic tree. The phylogenetic tree was constructed from the concatenated datasets using the maximum-likelihood method (ML) in RAxML 8.2.4 [27], employing a general time-reversible gamma-distributed rate variation model. Branch support was assessed using 1,000 bootstrap replicates, and only values ≥ 50% were shown at the nodes.
The most suitable substitution models were selected using MrModelTest v.2.4 [28] and implemented in MrBayes v.3.2.4 [29] for Bayesian analysis, which was continued until the average standard deviation of the split frequencies fell below 0.01. Generated trees underwent a 25% burn-in procedure to calculate posterior probabilities (PP). Probabilities ≥ 0.9 were displayed at the nodes. The phylogenetic tree was visualized using FigTree version 1.4.4 software [30].
Leaf spot symptoms were found on coastal hogfennel plants grown in a field during the crop disease survey. The symptoms began as small, brown, and circular to irregular spots exhibiting yellow halos. As the disease progressed, the spots coalesced into irregular dark-brown lesions (Fig. 1A and B). The incidence of disease symptoms in the field ranged from 5 to 10%.
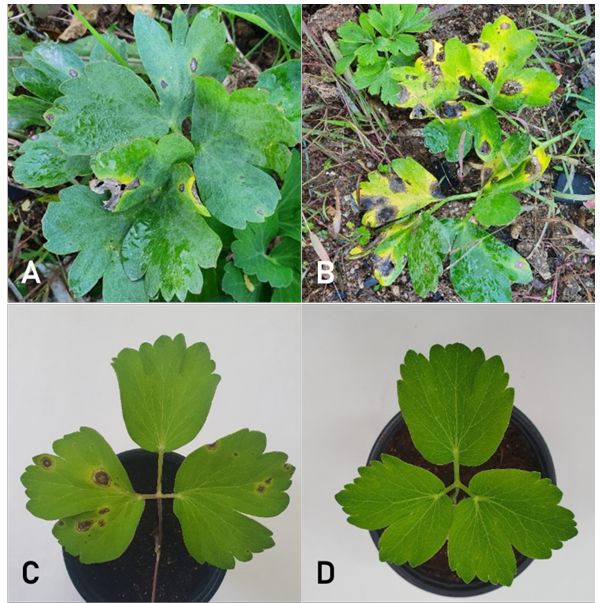

Fig. 1. Leaf spot symptoms of coastal hogfennel plants. (A, B) Symptoms observed in the investigated field. (C) Induced symptoms by artificial inoculation with the isolate (PEJA-2207) of Didymella acutilobae. (D) A non-inoculated control plant.
Three single-conidium isolates (PEJA-2203, PEJA-2205, and PEJA-2207) from coastal hogfennel were analyzed for their phylogenetic characteristics. The optimal models for the LSU, ITS, TUB2, and RPB2 alignments were identified as HKY + I + G, SYM + I + G, GTR + I + G, and GTR + I + G, respectively, based on model tests. Overall, 2,470 characters, including alignment gaps (LSU, 958; ITS, 558; RPB2, 600; and TUB2, 354), were obtained from the concatenated four-gene sequence alignments of the 24 ingroup taxa. The ML and Bayesian trees showed no significant topological differences. Therefore, an ML tree with bootstrap and PP values at the nodes in the phylogenetic tree was presented. Phylogenetic analyses confirmed that the three isolates clustered with the reference strain KACC 410302 of Didymella acutilobae G.B. Lee and W.G. Kim [31] (Fig. 2). The isolates formed a well-supported cluster with acceptable bootstrap and PP values, distinguishing them from other closely related Didymella spp. [11,13,25,32]. The sequence data of LSU, ITS, TUB2, and RPB2 obtained from the isolates were deposited in the NCBI GenBank database (Table 2).
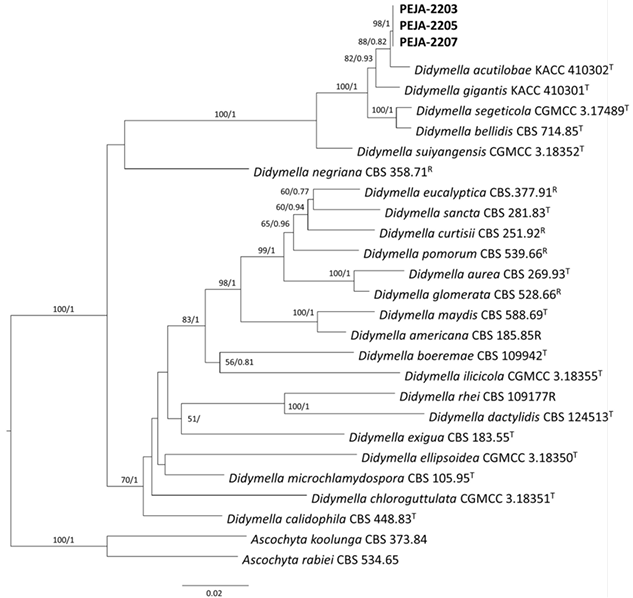
Fig. 2. Phylogenetic tree inferred from the maximum-likelihood analysis with general time-reversible model based on concatenated alignments of 28S large subunit of the nrDNA gene, internal transcribed spacer regions 1 and 2 including 5.8S nrDNA gene, β-tubulin, and RNA polymerase II second largest subunit sequences of three isolates (PEJA-2203, PEJA-2205, and PEJA-2207) from coastal hogfennel and Didymella spp. strains retrieved from the NCBI GenBank database. The phylogenetic tree was rooted to Ascochyta koolunga (CBS 373.84) and Ascochyta rabiei (CBS 534.65). RAxML bootstrap values (BS) and Bayesian posterior probabilities (PP) are provided at nodes (BS/PP). Values of BS/ PP more than 50/0.5 are shown at the nodes. The bar at 0.08 represents the number of nucleotide substitutions per site. T: type or ex-type strains; R: reference strains.
Table 2. Accession number of sequence data obtained from Didymella acutilobae isolates and reference species of Didymella and outgroup taxa deposited in GenBank
| Species | Strain/Isolate | Host/Substrate | Locality | Genbank accession number | |||
|---|---|---|---|---|---|---|---|
| TUB2 | RPB2 | LSU | ITS | ||||
| D. acutilobae | PEJA-2203 | Peucedanum japonicum | Korea | PX555481 | PX555484 | PX578144 | PX593789 |
| PEJA-2205 | Peucedanum japonicum | Korea | PX555482 | PX555485 | PX578145 | PX593790 | |
| PEJA-2207 | Peucedanum japonicum | Korea | PX555483 | PX555486 | PX578146 | PX593791 | |
| D. acutilobae | KACC 410302 | Angelica acutiloba | Korea | OQ744073 | OQ744071 | OQ749983 | OQ749981 |
| D. americana | CBS 185.85 | Zea mays | Russia | KT389594 | FJ427088 | GU237990 | FJ426972 |
| D. aurea | CBS 269.93 | Medicago polymorpha | New Zealand | KT389599 | GU237557 | GU237999 | GU237818 |
| D. bellidis | CBS 714.85 | Bellis perennis | The Netherlands | KP330417 | GU237586 | GU238046 | GU237904 |
| D. boeremae | CBS 109942 | Medicago littoralis | Australia | KT389600 | FJ427097 | GU238048 | FJ426982 |
| D. calidophila | CBS 448.83 | Desert soil | Egypt | MT018170 | FJ427168 | GU238052 | FJ427059 |
| D. chloroguttulata | CGMCC 3.18351 | Air | China | KY742142 | KY742299 | KY742211 | KY742057 |
| D. curtisii | CBS 251.92 | Nerine sp. | The Netherlands | MT018131 | FJ427148 | GU238013 | FJ427038 |
| D. dactylidis | CBS 124513 | Dactylis glomerata | USA | MT018173 | GU237599 | GU238061 | GU237766 |
| D. ellipsoidea | CGMCC 3.18350 | Air | China | KY742145 | KY742302 | KY742214 | KY742060 |
| D. eucalyptica | 377.91 | Eucalyptus sp. | Australia | KT389605 | GU237562 | GU238007 | GU237846 |
| D. exigua | CBS 183.55 | Rumex arifolius | France | EU874850 | GU237525 | EU754155 | GU237794 |
| D. gigantis | KACC 410301 | Angelica gigas | Korea | OQ731405 | OQ731407 | OQ746316 | OQ746336 |
| D. glomerata | CBS 528.66 | Chrysanthemum sp. | The Netherlands | GU371781 | FJ427124 | EU754184 | FJ427013 |
| D. ilicicola | CGMCC 3.18355 | Ilex chinensis | Italy | KY742150 | KY742307 | KY742219 | KY742065 |
| D. maydis | CGMCC 3.17489 | Zea mays | USA | GU371782 | FJ427190 | EU754192 | FJ427086 |
| D. microchlamydospora | CBS 105.95 | Eucalyptus sp. | UK | GU238143 | GU237787 | GU237657 | MT018177 |
| D. negriana | CBS 358.71 | Vitis vinifera | Germany | KT389610 | GU237635 | GU238116 | GU237838 |
| D. pomorum | CGMCC 3.18352 | Polygonum tataricum | The Netherlands | KT389618 | FJ427166 | GU238028 | FJ427056 |
| D. rhei | CBS 109177 | Rheum rhaponticum | New Zealand | KP330428 | GU237653 | GU238139 | GU237743 |
| D. sancta | CBS 281.83 | Ailanthus altissima | South Africa | MT018132 | MT005619 | GU238030 | FJ427063 |
| D. segeticola | CGMCC 3.17489 | Cirsium segetum | China | KP330414 | KP330399 | KP330455 | KP330443 |
| D. suiyangensis | CGMCC 3.18352 | Air | China | MT018174 | MT005643 | MN943747 | MN973540 |
| Ascochyta koolunga | CBS 373.84 | Pisum sativum | Australia | KT389775 | KT389560 | KT389698 | KT389481 |
| A. rabiei | CBS 534.65 | Cicer arietinum | India | GU237533 | KP330405 | GU237970 | GU237886 |
TUB2: β-tubulin; RPB2: RNA polymerase II second largest subunit; LSU: 28S large subunit of the nrDNA gene; ITS: internal transcribed spacer regions 1 & 2 including 5.8S nrDNA gene; CBS: Westerdijk Fungal Biodiversity Institute, Utrecht, The Netherlands; CGMCC: China General Microbiological Culture Collection, Beijing, China; KACC: Korean Agricultural Culture Collection, National Institute of Agricultural Sciences, Wanju, Korea.
The colony diameter of a representative isolate (PEJA-2207) grown on MEA, OA, and PDA for 7 days was 66–69, 67–69, and 71–72 mm, respectively. Colonies on MEA displayed white floccose and fawn mycelium with pycnidia in the aged zones (Fig. 3A). Colonies on OA displayed white floccose and buff mycelium in the middle (Fig. 3B). Colonies on PDA displayed rosy buff-to-buff mycelium with dark brown concentric rings covered with whitish mycelia (Fig. 3C). An NaOH spot test on the MEA showed negative reaction.
No teleomorphs were produced in culture. Pycnidia were non-papillate or papillate, globose, brown to black, solitary (Fig. 3D), and measured 67–223 μm in diameter. They possessed 1–3 ostioles and sometimes exhibited elongated necks. Pycnidial walls were pseudoparenchymatous, comprising round cells in 2–3 layers, and measured 7–17 μm in thickness (Fig. 3E). Conidiogenous cells were hyaline, phialidic, and varied in shape from globose to ampulliform or flask-shaped, and measured 3.3–4.0 × 3.74.3 μm (Fig. 3E). Conidia were smooth-walled, ellipsoidal, aseptate, typically containing multiple guttules (Fig. 3F), and measured 3.5–6.4 × 1.7–2.7μm (av. 4.5 × 2.1 μm). The conidial matrices were saffron to cinnamon in PDA culture. Chlamydospores were not detected.
The mycological characteristics of the D. acutilobae isolate from coastal hogfennel were generally consistent with the descriptions of the holotype species [31] (Table 3), although they differed slightly in pycnidial neck shape and growth rates. The slight differences in pycnidial neck shape and growth rate highlight the intraspecific variation among D. acutilobae strains, suggesting that further comparative studies are needed.
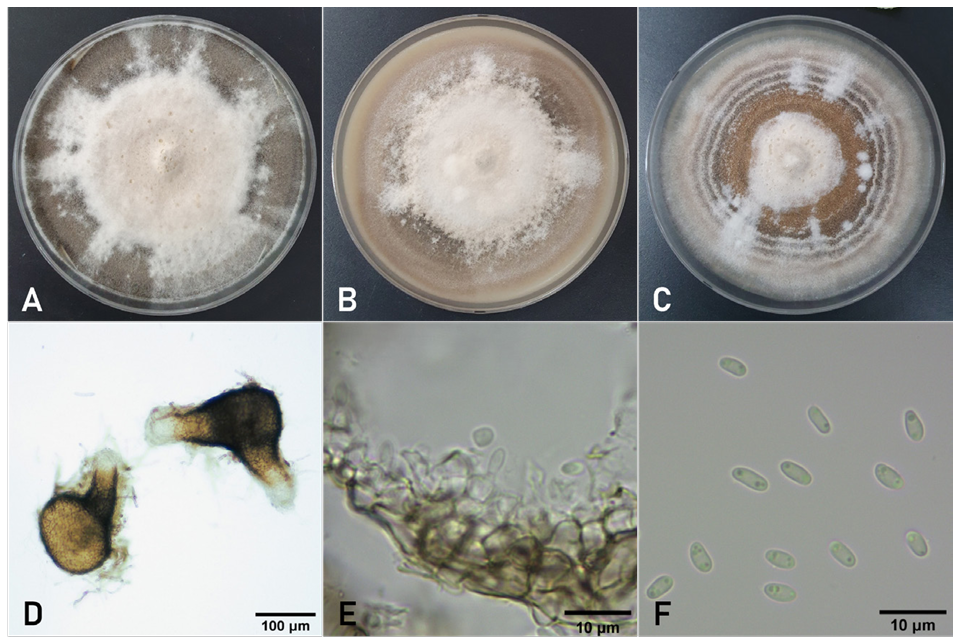
Fig. 3. Cultural and morphological features of Didymella acutilobae isolated from coastal hogfennel. Fourteen-day-old colonies of the fungus on malt extract agar (A), oatmeal agar (B), and potato dextrose agar (C). (D) Pycnidia produced in oatmeal agar. (E) Conidiogenous cells and conidia on a pycnidial wall. (F) Conidia.
Table 3. Morphological and cultural characteristics of Didymella acutilobae isolate from coastal hogfennel and the holotype species
| Isolate (Reference) | Morphological characteristics | Colony on mediaa and result of NaOH spot tests | |
|---|---|---|---|
| Pycnidia | Conidiogenous cells and conidia | ||
| PEJA-2207 (Present study) | 67–223 μm in diameter. Globose, brown to black, solitary, 1–3 ostioles, elongated neck, non-papillate or papillate. | Conidiogenous cells: 3.3–4.0 × 3.7–4.3 μm, globose to ampulliform or flask-shaped, hyaline, phialidic. Conidia: 3.5–6.4 × 1.7–2.7 μm (av. 4.5 × 2.1 μm), ellipsoidal, aseptate, multiple guttules mostly. Conidial matrices saffron to cinnamon in PDA. Chlamydospores absent. | MEA: white floccose, fawn mycelium; 66–69 mm. OA: white floccose, buff mycelium; 67–69 mm. PDA: white spots, rosy buff-to-buff mycelium with dark brown concentric rings; 71–72 mm. NaOH spot test: negative. |
| KACC 410302 (32) | 70–240 μm in diameter. Solitary or confluent, globose, brown to black, with 1–5 ostioles, non-papillate or papillate. | Conidia: 2.9–6.5 × 1.6–3.0 μm, ellipsoidal or slightly curved, aseptate with usually 2 bipolar guttules. Conidial matrices white. Chlamydospores absent. | MEA: brown to black with light concentric rings; 54–61 mm. OA: brown to dark olivaceous; 53–54 mm. PDA: white to light brown with concentric rings; 53–55 mm. NaOH spot test: negative. |
aDiameter of colonies on MEA, OA, and PDA was measured after incubation at 22 ℃ for 7 days. Colony morphology was investigated after incubation at 22 ℃ for 14 days. MEA: malt extracted agar; PDA: potato dextrose agar; OA: oatmeal agar.
The pathogenicity tests confirmed that the three D. acutilobae isolates caused leaf spot symptoms identical to those occurred in the field (Fig. 1C). Control plants were asymptomatic (Fig. 1D). Re-isolation of the fungus from induced lesions confirmed the identity of the inoculated isolates.
D. acutilobae was first reported to cause leaf spot and stem rot in Angelica acutiloba (Siebold & Zucc.) Kitag. [31], and a subspecies of the fungus was recently reported to cause leaf spot in East Asian hogweed [33]. To date, several diseases have been reported in coastal hogfennel in Korea and Japan, including powdery mildew, Fusarium wilt, damping-off, Sclerotinia rot, and leaf spot [10,34]. In Korea, Alternaria peucedani and Ascochyta sp. have been documented as the causal fungi of leaf spot in coastal hogfennel [10], with D. acutilobae having never been documented, until now, to act as the etiologic agent for this disease. Thus, this is the first study to report on D. acutilobae-induced leaf spot in coastal hogfennel.
No potential conflict of interest was reported by the authors.
This study was supported by a research grant (PJ01450701) from the Rural Development Administration, Korea.
1. Plants of the World Online. Peucedanum japonicum. [Internet]. Kew: Royal Botanic Garden; 2025 [cited 2025 Dec 18]. Available from: https://powo.science.kew.org/
2. Hisamoto M, Kikuzaki H, Ohigashi H, Nakatani N. Antioxidant compounds from the leaves of Peucedanum japonicum Thunb. J Agric Food Chem 2003;51:5255-61. https://doi.org/10.1021/jf0262458
3. Uy NP, Kim H, Ku J, Lee S. Regional variations in Peucedanum japonicum antioxidants and phytochemicals. Plants 2024;13:377. https://doi.org/10.3390/plants13030377
4. Park JH, Kim JH, Shin JY, Kang ES, Cho BO. Anti-inflammatory effects of Peucedanum japonicum Thunberg leaves extract in lipopolysaccharide-stimulated RAW264.7 cells. J Ethnopharmacol 2023;309:116362. https://doi.org/10.1016/j.jep.2023.116362
5. Monte E, Bridge PD, Sutton BC. An integrated approach to Phoma systematics. Mycopathologia 1991;115:89–103. https://doi.org/10.1007/bf00436797
6. Boerema GH, de Gruyter J, Noordeloos ME, Hamers MEC. Phoma identification manual. Differentiation of specific and infra-specific taxa in culture. Oxfordshire: CABI Publishing; 2004.
7. Aveskamp MM, de Gruyter J, Crous PW. Biology and recent developments in the systematics of Phoma, a complex genus of major quarantine significance. Fungal Divers 2008;31:1–18.
8. Aveskamp MM, de Gruyter J, Woudenberg JHC, Hamers MEC. Highlights of the Didymellaceae: A polyphasic approach to characterise Phoma and related pleosporalean genera. Stud Mycol 2010;65:1–60. https://doi.org/10.3114/sim.2010.65.01
9. de Gruyter J, Aveskamp MM, Woudenberg JHC, Verkley GJ, Groenewald JZ, Crous PW. Molecular phylogeny of Phoma and allied anamorph genera: Towards a reclassification of the Phoma complex. Mycol Res 2009;113:508–19. https://doi.org/10.1016/j.mycres.2009.01.002
10. List of Plant Diseases in Korea. Peucedanum japonicum [Internet]. Seoul: The Korean Society of Plant Pathology; Wanju: National Institute of Agricultural Sciences; 2025 [cited 2025 Dec 18]. Available from: https://genebank.rda.go.kr/kplantdisease.do
11. Chen Q, Jiang JR, Zhang GZ, Cai L, Crous PW. Resolving the Phoma enigma. Stud Mycol 2015;82:137–217. https://doi.org/10.1016/j.simyco.2015.10.003
12. Rayner RW. A mycological colour chart. Kew: Commonwealth Mycological Institute; 1970.
13. Lee GB, Kim KD, Cho WD, Kim WG. Didymella gigantis sp. nov. causing leaf spot in Korean angelica. Mycobiology 2023;51:393–400. https://doi.org/10.1080/12298093.2023.2289259
14. Dong L, Liu S, Li J, Tharreau D, Liu P, Tao D, Yang Q. A rapid and simple method for DNA preparation of Magnaporthe oryzae from single rice blast lesions for PCR-based molecular analysis. Plant Pathol J 2022;38:679–84. https://doi.org/10.5423/PPJ.NT.02.2022.0017
15. Rehner SA, Samuels GJ. Taxonomy and phylogeny of Gliocladium analysed from nuclear large subunit ribosomal DNA sequences. Mycol Res 1994;98:625–34. https://dx.doi.org/10.1016/S0953-7562(09)80409-7
16. Vilgalys R, Hester M. Rapid genetic identification and mapping of enzymatically amplified ribosomal DNA from several Cryptococcus species. J Bacteriol 1990;172:4238–46. https://doi.org/10.1128/jb.172.8.4238-4246.1990
17. de Hoog GS, Gerrits van den Ende AH. Molecular diagnostics of clinical strains of filamentous Basidiomycetes. Mycoses 1998;41:183–9. https://doi.org/10.1111/j.1439-0507.1998.tb00321.x
18. White TJ, Bruns T, Lee SB, Taylor JW. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ, editors. PCR protocols: A guide to methods and applications. San Diego: Academic Press; 1990. p. 315–22. https://doi.org/10.1016/B978-0-12-372180-8.50042-1
19. Woudenberg JHC, Aveskamp MM, de Gruyter J, Spiers AG, Crous PW. Multiple Didymella teleomorphs are linked to the Phoma clematidina morphotype. Persoonia 2009;22:56–62. https://doi.org/10.3767/003158509X427808
20. Sung GH, Sung JM, Hywel-Jones NL, Spatafora JW. A multi-gene phylogeny of Clavicipitaceae (Ascomycota, Fungi): Identification of localized incongruence using a combinational bootstrap approach. Mol Phylogenet Evol 2007;44:1204–23. https://doi.org/10.1016/j.ympev.2007.03.011
21. Liu YJ, Whelen S, Hall BD. Phylogenetic relationships among ascomycetes: Evidence from an RNA polymerase II subunit. Mol Biol Evol 1999;16:1799–808. https://doi.org/10.1093/oxfordjournals.molbev.a026092
22. Chen Q, Zhang K, Zhang G, Cai L. A polyphasic approach to characterise two novel species of Phoma (Didymellaceae) from China. Phytotaxa 2015;197:267–81. https://doi.org/10.11646/phytotaxa.197.4.4
23. Hou LW, Groenewald JZ, Pfenning LH, Yarden O, Crous PW, Cai L. The phoma-like dilemma. Stud Mycol 2020;96:309–96. https://doi.org/10.1016/j.simyco.2020.05.001
24. Chen Q, Bakhshi M, Balci Y, Broders KD, Cheewangkoon R, Chen SF, Fan XL, Gramaje D, Halleen F, Jung MH, et al. Genera of phytopathogenic fungi: GOPHY 4. Stud Mycol 2022;101:417–564. https://doi.org/10.3114/sim.2022.101.06
25. Chen Q, Hou LW, Duan WJ, Crous PW, Cai L. Didymellaceae revisited. Stud Mycol 2017;87:105–59. https://doi.org/10.1016/j.simyco.2017.06.002
26. Kumar S, Stecher G, Tamura K. MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 2016;33:1870-4. https://doi.org/10.1093/molbev/msw054
27. Stamatakis A. RAxML version 8: A tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 2014;30:1312-13. https://doi.org/10.1093/bioinformatics/btu033
28. Nylander JAA. MrModeltest version 2.4. Uppsala: Evolutionary Biology Centre; 2004.
29. Ronquist F, Teslenko M, van der Mark P, Ayres DL, Darling A, Höhna S, Larget B, Liu L, Suchard MA, Huelsenbeck JP. MrBayes 3.2: Efficient Bayesian phylogenetic inference and model selection across a large model space. Syst Biol 2012;61:539–42. https://doi.org/10.1093/sysbio/sys029
30. Rambaut A. FigTree version 1.4.4. Edinburgh: Institute of Evolutionary Biology; 2018.
31. Lee GB, Kim KD, Cho WD, Kim WG. Didymella acutilobae sp. nov. causing leaf spot and stem rot in Angelica acutiloba. Mycobiology 2023;51:313–9. https://doi.org/10.1080/12298093.2023.2254052
32. de Gruyter J, Noordeloos ME, Boerema GH. Contributions towards a monograph of Phoma (Coelomycetes) – I. 2. Section Phoma: Additional taxa with very small conidia and taxa with conidia up to 7 μm long. Persoonia 1993;15:369–400.
33. Lee GB, Cho WD, Kim WG. A novel subspecies of Didymella acutilobae causing leaf spot in East Asian hogweed. Mycobiology 2024;52:446–53. https://doi.org/10.1080/12298093.2024.2424016
34. Farr DF, Rossman AY, Castlebury LA. Peucedanum japonicum [Internet]. Beltsville: USDA ARS Mycology and Nematology Genetic Diversity and Biological Laboratory; 2025 [cited 2025 Dec 18]. Available from: https://fungi.ars.usda.gov/