Ju-Heon Lee1,2, Youngsoo Kim1, Jong-Taek Park1, Dong-Hyuk Lee1, and Hee-Young Jung2,*
1Apple Research Center, National Institute of Horticultural & Herbal Science, Gunwi 43100, Korea
2Department of Plant Medicine, Kyungpook National University, Daegu 41566, Korea
*Correspondence to heeyoung@knu.ac.kr
Korean Journal of Mycology (Kor J Mycol) 2026 March, Volume 54, Issue 1, pages 43-54.
https://doi.org/10.4489/kjm.2026.54.1.5
Received on February 11, 2026, Revised on March 22, 2026, Accepted on March 23, 2026, Published on March 31, 2026.
Copyright © The Korean Society of Mycology.
This is an Open Access article which is freely available under the Creative Commons Attribution-Non-Commercial 4.0 International License (CC BY-NC) (https://creativecommons.org/licenses/by-nc/4.0/).
Fungal isolate ARI-25-A10 was obtained from an ambrosia beetle collected in an apple orchard in Korea. The isolate was identified through an integrative taxonomic framework incorporating cultural and morphological characterization, growth responses under different temperature and pH conditions, and multilocus phylogenetic analyses. When cultured on potato dextrose agar at 25°C for 5 days, colonies reached 65.3–67.3 mm in diameter. Colony morphology transitioned from white to pale smoke grey to light olivaceous as incubation progressed, and sclerotia developed during extended culture. In temperature-dependent growth assays, colony development was minimal at 10°C and 30°C, whereas optimal growth was observed between 15°C and 25°C. In pH-response assays, optimal growth occurred at pH 5, with stable growth also observed between pH 4 and 6, indicating an optimal pH range of 4–5. Hyphae were hyaline, irregularly septate, and averaged 2.2 μm in width. BLAST analyses of the internal transcribed spacer, actin, RNA polymerase II second largest subunit, translation elongation factor 1-α, and β-tubulin gene sequences revealed high sequence similarity between ARI-25-A10 and Cytospora ceratosperma. In addition, multilocus phylogenetic reconstruction using concatenated datasets placed ARI-25-A10 within a strongly supported clade comprising previously reported and characterized C. ceratosperma strains. This study represents the first report of C. ceratosperma associated with an ambrosia beetle collected from an apple orchard in Korea.
Ambrosia beetles, Cytospora ceratosperma, Korean apple orchard, Mycological characterization
The genus Cytospora, belonging to the family Cytosporaceae, was originally proposed by Ehrenberg [1] with the inclusion of C. betulina, C. epimyces, C. resinae, and C. ribis. Subsequently, C. chrysosperma was designated as the lectotype species of the genus [2]. Most Cytospora species are recognized as plant pathogens responsible for canker and dieback diseases affecting a wide range of deciduous and coniferous trees, although occasional occurrences on herbaceous plants have also been documented. Diseases caused by Cytospora are commonly referred to as Valsa canker, Leucostoma canker, or perennial canker [3].
Several Cytospora species are considered facultative wound parasites, primarily infecting hosts that are stressed or weakened. The genus exhibits considerable ecological versatility, and some species have been isolated as endophytes from asymptomatic bark, xylem tissues, or foliar tissues of woody plants [4]. Depending on host condition and environmental factors, Cytospora species may occur as xylotropic (xylem-inhabiting) endophytes, saprophytes, or epiphytes [5].
Traditionally, species identification within Cytospora relied primarily on morphological traits and host association. However, these criteria have often proven inadequate. For example, Cytospora chrysosperma has been reported as a major causal agent of canker disease on poplar and willow trees in China [6], whereas C. kuanchengensis, together with five additional Cytospora species, has been identified as the principal pathogen complex associated with chestnut (Castanea mollissima) canker in the same region [7]. In the United States, fifteen Cytospora species have been identified from woody hosts exhibiting canker and dieback symptoms [8].
The extensive global distribution of Cytospora and its exceptionally broad host range, encompassing angiosperms and gymnosperms, and both woody and non-woody plants, has complicated species delimitation based solely on host affiliation [5]. Consequently, frequent synonymization and repeated taxonomic revisions have resulted in continued uncertainty among plant pathologists and mycologists. To resolve these issues, integrative taxonomic approaches that combine molecular phylogenetic analyses with morphological characteristics and host information have become the standard framework for species identification, as supported by recent multigene phylogenetic studies [9,10].
In the present study, a fungal isolate (ARI-25-A10), presumed to belong to the genus Cytospora, was obtained from an ambrosia beetle sampled in a Korean apple orchard. The objective of this study was to determine the taxonomic identity and phylogenetic placement of the isolate through an integrative approach combining morphological characterization with multigene molecular phylogenetic analyses.
Ambrosia beetles were collected from an apple orchard located at the Apple Research Center in Gunwi, Republic of Korea (36°16′44.2″N, 128°27′56.3″E) using standard beetle traps. Following collection, adult beetles were surface-sterilizedby immersion in 70% ethanol for 1 min and subsequently allowed to dry at ambient temperature for approximately 10 min. Individual beetles were then aseptically transferred to potato dextrose agar (PDA; Difco, Detroit, MI, USA) plates and incubated at 25°C for 3 days to facilitate fungal emergence.
Hyphal growth emerging from the plated beetles was transferred to fresh PDA plates to establish axenic cultures, which were subsequently incubated at the same temperature (25°C) for an additional two weeks. After repeated subculturing and verification of culture purity, a single isolate was selected and designated ARI-25-A10. The isolate was preserved in 20% (v/v) glycerol and stored at −80°C for long-term preservation. The isolate was deposited in the Korean Agricultural Culture Collection (KACC) under the accession number KACC 411281.
Isolate ARI-25-A10 was cultured to assess its temperature and pH-dependent growth, cultural characteristics, and morphological features. The isolate was grown on PDA, malt extract agar (MEA; Difco, Detroit, MI, USA), and oatmeal agar (OA; Difco, Detroit, MI, USA) in 90 mm Petri dishes, with plates incubated at 25°C under dark conditions. Colony diameters were measured throughout incubation to assess growth, and changes in colony pigmentation were recorded. Growth assays were conducted on PDA at temperatures ranging from 10 to 30°C at 5°C intervals and pH values from 4 to 9 at 1 pH intervals. Morphological observations of hyphae, conidia, and conidiophores were conducted using a light microscope (CX-43; Olympus, Tokyo, Japan), and measurements of the dimensions of these structures were recorded.
Total genomic DNA of isolate ARI-25-A10 was extracted using the HiGene Genomic DNA Preparation Kit (Biofact, Daejeon, Korea) to determine its molecular identity and phylogenetic placement. PCR amplification targeted the internal transcribed spacer (ITS) region and four genes commonly used for species-level identification in Cytospora: actin (ACT), the second largest subunit of RNA polymerase II (RPB2), translation elongation factor 1-α (TEF1-α), and β-tubulin (β-TUB).
Primers ITS1F and ITS4 [11] were used to amplify the ITS region, whereas ACT, RPB2, TEF1-α, and β-TUB were amplified with the primer pairs ACT512F/ACT728R, RPB2-5F/RPB2-7cR, EF1-728F/ EF1-986R, and Bt2a/Bt2b, respectively [12–15]. PCR products were verified by electrophoresis in 1% agarose gels stained with ethidium bromide. Sequencing was carried out by SolGent Co. (Daejeon, Korea), and sequence assembly and editing were conducted using SeqMan Lasergene software (DNAStar Inc., Madison, WI, USA). All sequences were deposited in GenBank under accession numbers LC911265 (ITS), LC911266 (ACT), LC911267 (RPB2), LC911268 (TEF1-α), and LC911269 (β-TUB).
Phylogenetic analyses were conducted to determine the taxonomic position of isolate ARI-25-A10 based on the sequence data generated in this study. For comparative analyses, reference sequences of representative species within the genus Cytospora were retrieved from the National Center for Biotechnology Information (NCBI) database and incorporated into the analyses (Table 1). Sequences were aligned in MEGA version 7 using the Clustal X version 2.0 algorithm [16].
Phylogenetic trees were inferred from the ITS region alone and from a concatenated dataset comprising partial sequences of the ACT, RPB2, TEF1-α, and β-TUB genes. Tree inference was carried out using the maximum likelihood (ML) method [17] under Kimura’s two-parameter substitution model [18]. Alignment gaps were excluded from the analysis, and tree topology was optimized using the nearest neighbor interchange algorithm. Bootstrap resampling (1,000 iterations) was performed to assess the robustness of inferred clades.
Table 1. The species included in the phylogenetic analysis and their corresponding GenBank accession numbers
| Species | Strain | GenBank accession numbers | ||||
|---|---|---|---|---|---|---|
| ITS | ACT | RPB2 | TEF1-α | β-TUB | ||
| Cytospora ceratosperma | ARI-25-A10 | LC911265 | LC911266 | LC911267 | LC911268 | LC911269 |
| Cytospora ceratosperma | CBS 109777 | PP988755 | KX964687 | KX965464 | KX965053 | EU219136 |
| Cytospora ceratosperma | CBS 192.42 | KY051900 | KX964785 | KX965529 | KX965149 | KX964959 |
| Cytospora ceratosperma | CFCC 89627 | KR045648 | KU711012 | KU710979 | KU710935 | KR045689 |
| Cytospora ceratosperma | CFCC 89626 | KR045647 | KU711011 | KU710978 | KU710934 | KR045688 |
| Cytospora ceratosperma | CBS 217.81 | PP988760 | KX964799 | KX965541 | KX965163 | KX964972 |
| Cytospora ceratosperma | CBS 512.76 | KY051941 | KX964821 | KX965552 | KX965184 | KX964994 |
| Cytospora juniperiicola | CBS 654.97 | KY051944 | KX964824 | KX965555 | KX965187 | KX964995 |
| Cytospora juniperiicola | CBS 196.97 | KY051906 | KX964791 | KX965535 | KX965154 | KX964964 |
| Cytospora myricicola | CFCC 59324 | OR769869 | OR767325 | OR767339 | OR767365 | OR767352 |
| Cytospora myricicola | CFCC 59325 | OR769870 | OR767326 | OR767340 | OR767366 | OR767353 |
| Cytospora myricicola | CFCC 59323 | OR769868 | OR767324 | OR767338 | OR767364 | OR767351 |
| Cytospora pavettae | CBS 145562 | NR_165558 | MK876457 | MK876483 | MK876497 | MK876503 |
| Cytospora platyclada | CFCC 50505 | MH933646 | MH933553 | MH933611 | MH933517 | MH933582 |
| Cytospora platyclada | CFCC 50504 | MH933645 | MH933552 | MH933610 | MH933516 | MH933581 |
| Cytospora tamaricicola | CFCC 50507 | MH933651 | MH933559 | MH933616 | MH933525 | MH933587 |
| Cytospora tamaricicola | CFCC 50508 | NR_171068 | MH933560 | MH933617 | MH933523 | MH933588 |
| Cytospora viridistroma | CBS 116813 | KY051788 | KX964698 | KX965470 | KX965061 | KX964898 |
| Cytospora viridistroma | CBS 116811 | KY051786 | KX964696 | KX965468 | KX965060 | KX964896 |
| Diaporthe eres | CBS 145040 | MK442579 | MK442634 | MK442663 | MK442693 | MK442731 |
ITS: internal transcribed spacer; ACT: actin; RPB2: second largest subunit of RNA polymerase II; TEF1- α: translation elongation factor 1-α; β-TUB: β-tubulin.
The isolated strain is shown in bold.
When cultured on PDA medium at 25°C for 5 days, colonies reached diameters of 65.3–67.3 mm. Colonies were initially white, becoming pale smoke grey to light olivaceous as growth progressed. Colonies were flat with a uniform texture, and no asexual spores were observed. However, when incubation exceeded 30 days, colonies turned smoke grey, and sclerotia were sporadically formed (Fig. 1A–B). On MEA medium, colony diameters ranged from 24.9 to 27.6 mm, indicating slower growth than on PDA. Colonies were initially white and later became light olivaceous, remaining flat and uniform in texture (Fig. 1F–G). On OA medium, colonies reached 69.1–69.9 mm, comparable to growth on PDA. Colonies remained uniformly white throughout the incubation period, with no noticeable color change, and developed flocculent aerial mycelium (Fig. 1H–I).
Temperature-dependent growth assays showed minimal growth at 10°C and 30°C. In particular, at 30°C, growth was negligible even after 7 days of incubation, indicating an inability of the isolate to grow at elevated temperatures above 30°C. In contrast, normal growth was observed at 15°C, 20°C, and 25°C, and by 7 days, colonies at all three temperatures completely covered the 90 mm Petri dishes. These results demonstrate that optimal growth occurred between 15 and 25°C, with the most distinctive cultural features observed at 25°C (Fig. 2A–B).
In pH-dependent growth assays, colonies grown at pH 5 completely covered the 90-mm Petri dish after 7 days of incubation. Relatively good growth was also observed at pH 4 and pH 6, with colony diameters exceeding 60 mm. In contrast, at pH 7 and above, growth progressively declined as the pH increased (Fig. 2C–D).
Hyphae were hyaline, with irregularly distributed septa, averaging 2.2 µm in width (n = 10). Irregular, outward-projecting hyphal structures were observed, and some hyphae darkened to brown with prolonged incubation (Fig. 1D–E). In sclerotial regions, locally swollen hyphae were frequently observed, with an average width of 5.7 μm (n = 10) (Fig. 1C). On PDA, MEA, and OA culture media, conidiophores and conidia were not observed.
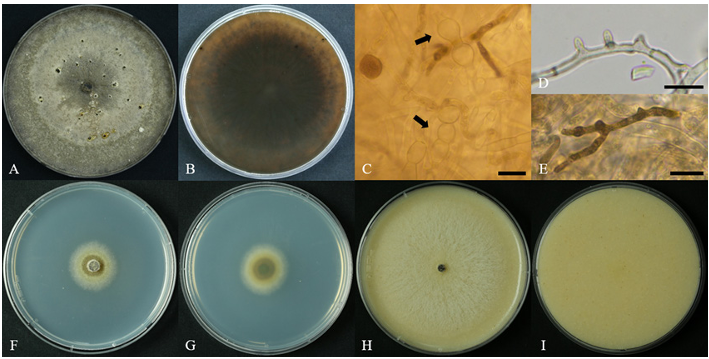
Fig. 1. Cultural and morphological characteristics of ARI-25-A10 (Cytospora ceratosperma). A, B: Colonies on potato dextrose agar after more than 30 days of incubation at 25°C, showing the formation of black sclerotia. C: Swollen regions of hyphae within the sclerotia (indicated by black arrows). D: Protrusion and branching observed at irregular regions of hyphae. E: Dark brown pigmentation observed in some hyphae after prolonged incubation. F, G: Colonies on malt extract agar after 5 days of incubation at 25°C. H, I: Colonies on oatmeal agar after 5 days of incubation at 25°C. Scale bars: C–E = 10 μm.
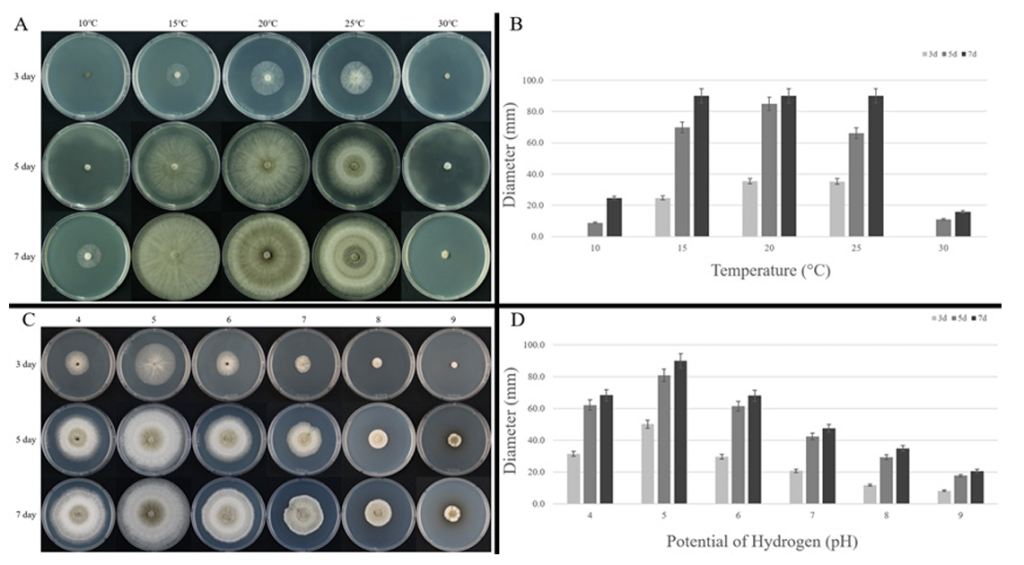
Fig. 2. Growth of ARI-25-A10 on potato dextrose agar under different temperature and pH conditions. A: Colony morphology observed after incubation for 3, 5, and 7 days under different temperature conditions (10, 15, 20, 25, and 30°C). B: Colony diameters measured under different temperature conditions. C: Colony morphology observed after incubation for 3, 5, and 7 days under different pH conditions (pH 4, 5, 6, 7, 8, and 9). D: Colony diameters measured under different pH conditions.
The cultural characteristics of isolate ARI-25-A10 were largely congruent with those reported for Cytospora ceratosperma, particularly with respect to colony color, morphology, and texture. In addition, the growth response of ARI-25-A10 to temperature conditions showed a pattern similar to that reported previously. C. ceratosperma has been reported to exhibit very slow growth at temperatures below 10°C, while growth at temperatures above 30°C is markedly reduced compared with that at 25°C [19]. In the present study, ARI-25-A10 displayed a comparable temperature-dependent growth pattern, further supporting its close cultural affinity with C. ceratosperma.
In contrast, clear cultural differences were observed between ARI-25-A10 and the phylogenetically related species C. junipericola and C. myricicola. C. junipericola reportedly forms greenish ash-colored colonies on MEA, exhibiting undulate margins and a woolly texture. However, ARI-25-A10 formed pale olivaceous colonies on MEA with a distinct central olivaceous band, exhibiting flat, uniform surfaces and lacking undulate or woolly margins.
Similarly, C. myricicola is known to produce colonies on PDA that change in color from white to dark, whereas ARI-25-A10 exhibited a color transition from pale olivaceous to grey, indicating a clear distinction in colony pigmentation between the two taxa (Table 2).
Table 2. Comparative analysis of the cultural characteristics of ARI-25-A10 and phylogenetically related species within the genus Cytospora.
| Species | Cultural characteristics | ||
|---|---|---|---|
| Color | Shape | Size (mm) | |
| Cytospora ceratosperma (ARI-25-A10)a | On PDA: Initially white, later becoming pale olivaceous to grey On MEA: Initially white, becoming pale olivaceous On OA: white |
On PDA: Flat with a uniform texture, showing white and olivaceous bands. Black sclerotia formed after prolonged incubation (>30 days) On MEA: Flat with a uniform texture, with a central olivaceous band On OA: Flat with uniform growth and sparse flocculent aerial mycelium. No asexual reproductive structures were observed |
On PDA, 66.2 mm in diameter after 5 days at 25°C On MEA, 26.3 mm in diameter after 5 days at 25°C On OA, 69.4 mm in diameter after 5 days at 25°C |
| Cytospora ceratosperma (CFCC 56503)b | Initially white, becoming smoke grey with olivaceous pigment. | Colonies were thick, with felty aerial mycelium and regular edges | 9 cm in diameter after 5 days |
| Cytospora ceratosperma (CFCC 55991)b | Initially white, becoming grey-olivaceous and smoky | Colonies flat with a uniform texture, with sparse aerial mycelium, and sterile | 9 cm in diameter after 4 days |
| Cytospora juniperiicolac | On MEA: Greenish ash | On MEA, colonies were moderately fast growing, irregular, flat, undulate, woolly, curled, and loosely attached to the medium | On MEA, 10 mm in diameter after 7 days at 18°C |
| Cytospora myricicolac | On PDA: Initially white, becoming dark | On PDA, colonies were flat, with flocculent aerial mycelium and entire margins, sterile | On PDA, 90 mm in diameter after 10 days at 25°C |
Partial nucleotide sequences of the ITS region and the ACT, RPB2, TEF1-α, and β-TUB genes were generated for isolate ARI-25-A10. The obtained sequences were compared with reference sequences in the NCBI database using BLAST searches to assess sequence similarity.
For the ITS region (637 base pairs), ARI-25-A10 showed the highest similarity to Cytospora ceratosperma NEFU2019 (99.8%, percent identity), followed by C. ceratosperma NEFU2020 (99.0%). Lower similarity values were observed with C. ceratophora CBS 109777 (98.2%), C. mali TS08-168 (98.0%), and C. pinastri CPC 28381 (97.8%).
ACT gene analysis (257 bp) indicated the highest similarity to C. ceratosperma CFCC 53154 (97.9%), followed by C. ceratosperma MFLU 17-0327 (94.5%) and C. ceratosperma CFCC 89626 (92.6%). In contrast, lower similarity values were found with C. rosicola CF 20197024 (86.8%), C. tamaricicola CFCC 50508 (85.8%), and C. ulmicola MFLUCC 18-1227 (85.8%).
For the RPB2 gene (876 bp), ARI-25-A10 exhibited the highest similarity to C. ceratosperma CFCC 53155 (99.7%), followed by C. ceratosperma Ct-95 (98.1%) and C. ceratosperma AR3426 (97.8%). Comparatively lower similarities occurred with C. cedri MFLU 17-0835 (94.1%), C. pavettae CBS 145562 (91.4%), and C. mali ARI-15-US (90.3%).
The TEF1-α gene sequence (317 bp) also showed the highest similarity to C. ceratosperma NEFU2020 (98.1%) and C. ceratosperma CFCC 53155 (98.0%), while C. ceratosperma CFCC 89624 showed 93.3% similarity. Lower similarity values were detected for C. spiraeicola CFCC 55830 (88.8%) and C. thailandica MFLUCC 17-0353 (86.3%).
For β-TUB (533 bp), ARI-25-A10 was most similar to C. ceratosperma CFCC 53155 (98.2%), followed by C. ceratosperma HMBF281 (95.0%) and C. ceratosperma CFCC 89626 (94.6%). In contrast, lower similarities were observed with C. pingbianensis XG107a (86.7%), C. diopuiensis CFCC 56756 (86.1%), and C. anacardii RM10 (85.4%).
Across all analyzed loci, ARI-25-A10 consistently showed the highest similarity to Cytospora ceratosperma. However, substantial intraspecific variation was evident. Notably, the act1 gene exhibited over a 5% difference in sequence similarity between C. ceratosperma CFCC 53154 (97.9%) and CFCC 89626 (92.6%), highlighting considerable genetic variation within the species.
Overall, BLAST analyses confirmed that ARI-25-A10 is most closely related to C. ceratosperma based on ITS, ACT, RPB2, TEF1-α, and β-TUB gene sequences, and is clearly distinct from other Cytospora species.
Phylogenetic analysis using the ML method based on concatenated sequences of the ITS region and the ACT, RPB2, TEF1, and β-TUB genes, placed ARI-25-A10 within a major clade together with C. ceratosperma. Although some variation was observed among strains within C. ceratosperma, they collectively constituted a single, well-supported clade that was clearly separated from other Cytospora species (Fig. 3).
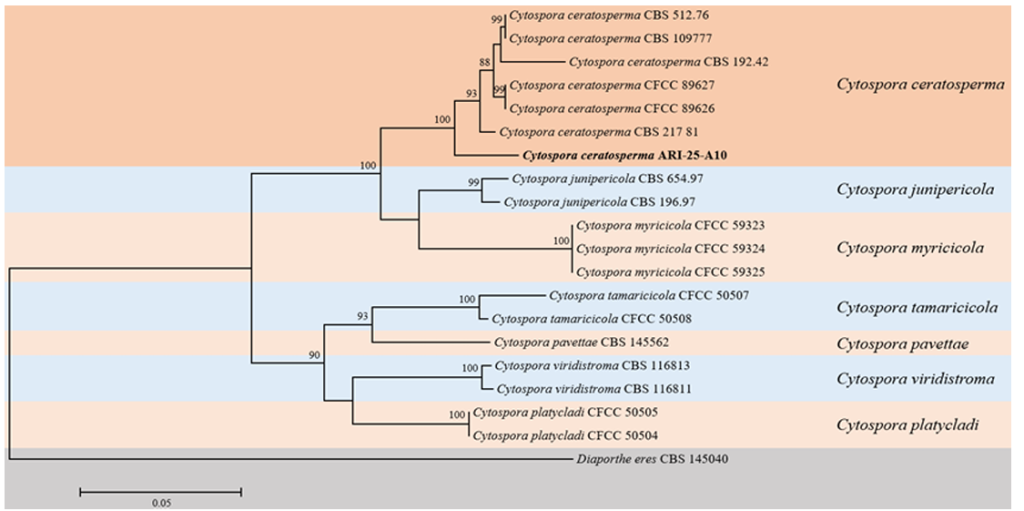
Fig. 3. Phylogenetic position of isolate ARI-25-A10 within the genus Cytospora. A multilocus phylogenetic tree was inferred using the maximum-likelihood approach based on concatenated partial sequences of the internal transcribed spacer, actin, RNA polymerase II second largest subunit, translation elongation factor 1-α, and β-tubulin. Only bootstrap support values exceeding 70%, obtained from 1,000 resampling replicates, are displayed at the corresponding nodes. The strain obtained in this study is indicated in bold. Diaporthe eres CBS 145040 was selected as the outgroup taxon. Branch lengths are proportional to the number of substitutions per site, as indicated by the scale bar (0.05).
The type species of Cytospora is C. chrysosperma, as clarified by Donk [2]. This species represents the asexual morph of Valsa sordida and is widely recognized as a causal agent of canker diseases, particularly on hosts in the family Salicaceae [20]. By contrast, the type species of Valsa, V. ambiens, has been demonstrated to be taxonomically conspecific with C. leucospora, the type species of Leucostoma [21]. Furthermore, the type species of Leucostoma, L. massarianum, is phylogenetically positioned within Cytospora, closely related to C. mali and C. persoonii [19,22]. On the basis of these phylogenetic relationships among type species, Leucostoma and related genera have long been regarded as synonyms of Cytospora.
Considering these taxonomic relationships, Rossman et al. [20] concluded that the generic name Cytospora should be adopted in preference to Valsa, as it is the older name and has been the focus of extensive recent taxonomic and phylogenetic studies. Consequently, they proposed the use of Cytospora (described in 1818) instead of Valsa (1825), Valsella (1870), Leucostoma (1917), Valseutypella (1919), and Leucocytospora (1927).
As of 2025, the MycoBank database lists a total of 852 species under Cytospora. However, many species described prior to 2000 lack DNA sequence data, and several taxa cannot be verified due to the absence of original material. These limitations substantially hinder reliable species identification. In addition, species of Cytospora exhibit remarkable morphological and ecological diversity. Even within a single species, considerable morphological variation may occur, and conidial dimensions are known to vary with culture medium or host plant [7,9,23]. To address these issues, Lin et al. [5] established a comprehensive phylogenetic framework based on multilocus sequence data (ITS, ACT, RPB2, TEF1-α, and β-TUB) and reorganized species of Cytospora, including the type species, using a polyphasic approach integrating molecular phylogeny, morphology, host range, and geographical distribution.
In the present study, a fungal isolate was obtained from ambrosia beetles collected in an apple orchard and designated ARI-25-A10. Based on its cultural characteristics and multilocus phylogenetic analyses conducted following the framework of Lin et al. [5], the isolate was identified as Cytospora ceratosperma. However, ARI-25-A10 failed to produce conidia in culture, precluding direct assessment of conidial morphology. To compensate for this limitation, detailed cultural characteristics, including growth responses under different temperature and pH conditions, were examined.
Within Cytospora, several species are known to include isolates that fail to produce asexual spores in culture. Consequently, species-level identification often relies on a combination of multiple morphological and cultural traits rather than on a single diagnostic character. Cultural features have been shown to provide valuable supplementary criteria for distinguishing species of Cytospora, particularly those associated with Prunus and Malus hosts [22]. Adams et al. [19] demonstrated that temperature-dependent growth rates differ among species and may therefore serve as an additional taxonomic criterion. These observations are consistent with the results of the present study. Cytospora ceratosperma has been reported to grow very slowly at 32°C and to exhibit minimal growth at 37°C, and similarly, ARI-25-A10 showed a pronounced reduction in growth at 30°C.
As noted above, species delimitation in Cytospora based solely on morphological characters is often unreliable, especially for taxa other than the well-studied species associated with Prunus and Malus [22]. For example, Cytospora sibiraeae was described solely from its sexual morph and is currently morphologically indistinguishable from C. sibiraeicola [24]. Nevertheless, Li et al. [25] demonstrated that these two species occurring on Sibiraea angustata are clearly separable based on phylogenetic analyses, further underscoring the critical role of molecular data in Cytospora taxonomy.
Cytospora ceratosperma is a widely distributed and commonly encountered species with a broad host range, yet it remains taxonomically problematic. Adams et al. [19] attempted to resolve morphological overlap among specimens assigned to this species by narrowing the species concept using ITS sequence data. However, the absence of type material and the limited resolving power of ITS-based phylogenies have impeded precise species delimitation. More recently, Li et al. [25] analyzed 21 isolates and showed that they formed a single, well-supported clade, leading to the synonymization of Cytospora ceratospermopsis, C. predappioensis, and C. vinacea with C. ceratosperma. Similarly, although ARI-25-A10 exhibited some sequence divergence from other C. ceratosperma isolates, it clustered with them in a strongly supported clade. Taking into account both its cultural characteristics and its phylogenetic placement, ARI-25-A10 is therefore most appropriately identified as Cytospora ceratosperma.
In this study, C. ceratosperma was isolated from ambrosia beetles, which are known to be xylomycetophagous insects. Previous studies have reported that ambrosia beetles such as Xylosandrus crassiusculus and X. germanus are implicated in the damage and decline of ornamental, fruit, and nut trees in the United States. In addition, a diverse range of fungi has been isolated from beetle galleries in infested host tissues, including opportunistic phytopathogens belonging to genera such as Botryosphaeria, Fusarium, and Diaporthe [26].
Furthermore, Gugliuzzo et al. [27] investigated fungal communities associated with Xylosandrus compactus in carob trees in Sicily, Italy, and identified Ambrosiella xylebori as the primary symbiont. In addition, Acremonium sp., Penicillium sp., and Cytospora sp. were also isolated. Notably, the association of Cytospora sp. with ambrosia beetles was reported for the first time, suggesting a possible relationship between ambrosia beetles and members of the genus Cytospora. However, the ecological roles and interactions of these fungi have not yet been elucidated.
Although C. ceratosperma was also isolated from ambrosia beetles in the present study, its direct interaction with the beetles or their role as vectors could not be determined. Therefore, further studies are required to elucidate the ecological relationship between ambrosia beetles and Cytospora species, particularly their potential role in pathogen transmission and disease development.
The authors declare that they have no potential conflicts of interest.
1. Ehrenberg CG. Sylvae mycologicae Berolinenses. Berlin: Formis Theophili Bruschcke; 1818. pp. 1–32.
2. Donk MA. Nomina conservanda proposita 1. Proposals in fungi. Deuteromycetes. Regnum Veg 1964;34:7–15.
3. Farr DF, Bills GF, Chamuris GP, Rossman AY. Fungi on plants and plant products in the United States. St. Paul: APS Press; 1989.
4. Bills GF, Redlin SC, Carris LM. Isolation and analysis of endophytic fungal communities from woody plants. In: Redlin SC, Carris LM, editors. Endophytic fungi in grasses and woody plants: Systematics, ecology, and evolution. St. Paul: APS Press; 1996. p. 31–65.
5. Lin L, Fan XL, Groenewald JZ, Jami F, Wingfield MJ, Voglmayr H, Jaklitsch W, Castlebury LA, Tian CM, Crous PW. Cytospora: An important genus of canker pathogens. Stud Mycol 2024;109:323–402. https://doi.org/10.3114/sim.2024.109.05
6. Fan XL, Tian CM, Yang Q, Liang YM, You CJ, Zhang YB. Cytospora from Salix in northern China. Mycotaxon 2014;129:303–15. https://doi.org/10.5248/129.303
7. Jiang N, Yang Q, Fan XL, Tian CM. Identification of six Cytospora species on Chinese chestnut in China. MycoKeys 2020;62:1–25. https://doi.org/10.3897/mycokeys.62.47425
8. Lawrence DP, Holland LA, Nouri MT, Travadon R, Abramians A, Michailides TJ, Trouillas FP. Molecular phylogeny of Cytospora species associated with canker diseases of fruit and nut crops in California, with the descriptions of ten new species and one new combination. IMA Fungus 2018;9:333–69. https://doi.org/10.5598/imafungus.2018.09.02.07
9. Fan XL, Bezerra JDP, Tian CM, Crous PW. Cytospora (Diaporthales) in China. Persoonia 2020;45:1–45. https://doi.org/10.3767/persoonia.2020.45.01
10. Shang QJ, Hyde KD, Camporesi E, Maharachchikumbura SSN, Norphanphoun C, Brooks S, Liu JK. Additions to the genus Cytospora with sexual morph in Cytosporaceae. Mycosphere 2020;11:189–224. https://doi.org/10.5943/mycosphere/11/1/2
11. White TJ, Bruns TD, Lee SB, Taylor JW. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, editors. PCR protocols: A guide to methods and applications. San Diego: Academic Press; 1990. p. 315–22.
12. Carbone I, Kohn LM. A method for designing primer sets for speciation studies in filamentous ascomycetes. Mycologia 1999;91:553–6. https://doi.org/10.1080/00275514.1999.12061051
13. Liu YJ, Whelen S, Hall BD. Phylogenetic relationships among ascomycetes: Evidence from an RNA polymerse II subunit. Mol Biol Evol 1999;16:1799–808. https://doi.org/10.1093/oxfordjournals.molbev.a026092
14. Sung GH, Sung JM, Hywel-Jones NL, Spatafora JW. A multi-gene phylogeny of Clavicipitaceae (Ascomycota, Fungi): Identification of localized incongruence using a combinational bootstrap approach. Mol Phylogenet Evol 2007;44:1204–23. https://doi.org/10.1016/j.ympev.2007.03.011
15. O’Donnell K, Kistler HC, Cigelnik E, Ploetz RC. Multiple evolutionary origins of the fungus causing Panama disease of banana: Concordant evidence from nuclear and mitochondrial gene genealogies. Proc Natl Acad Sci U S A 1998;95:2044–9. https://doi.org/10.1073/pnas.95.5.2044
16. Kumar S, Stecher G, Tamura K. MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 2016;33:1870–4. https://doi.org/10.1093/molbev/msw054
17. Felsenstein J. Evolutionary trees from DNA sequences: A maximum likelihood approach. J Mol Evol 1981;17:368–76. https://doi.org/10.1007/BF01734359
18. Kimura M. A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 1980;16:111–20. https://doi.org/10.1007/bf01731581
19. Adams GC, Wingfield MJ, Common R, Roux J. Phylogenetic relationships and morphology of Cytospora species and related teleomorphs (Ascomycota, Diaporthales, Valsaceae) from Eucalyptus. Stud Mycol 2005;52:1–144.
20. Rossman AY, Adams GC, Cannon PF, Castlebury LA, Crous PW, Gryzenhout M, Jaklitsch WM, Mejia LC, Stoykov D, Udayanga D, et al. Recommendations of generic names in Diaporthales competing for protection or use. IMA Fungus 2015;6:145–54. https://doi.org/10.5598/imafungus.2015.06.01.09
21. Spielman LJ. A monograph of Valsa on hardwoods in North America. Can J Bot 1985;63:1355–78.
22. Adams GC. Inoculating trees in nature: A problem of environmental interactions. In: Uotila A, Ahola V, editors. Shoot and foliage diseases. Helsinki: Finnish Forest Research Institute; 2002. p. 8–16.
23. Lin L, Pan M, Bezerra JDP, Tian C, Fan X. Re-evaluation of the fungal diversity and pathogenicity of Cytospora species from Populus in China. Plant Dis 2023;107:83–96. https://doi.org/10.1094/pdis-02-22-0260-re
24. Liu JK, Hyde KD, Jones EBG, Ariyawansa HA, Bhat DJ, Boonmee S, Maharachchikumbura SSN, McKenzie EHC, Phookamsak R, Phukhamsakda C, et al. Fungal diversity notes 1–110: Taxonomic and phylogenetic contributions to fungal species. Fungal Divers 2015;72:1–197. https://doi.org/10.1007/s13225-015-0324-y
25. Li J, Li J, Jiang N. Morphology and phylogeny of Cytospora (Cytosporaceae, Diaporthales) species associated with plant cankers in Tibet, China. MycoKeys 2024;104:51–70. https://doi.org/10.3897/mycokeys.104.113567
26. Gresham SDM, Ranger CM, Walgenbach JF, Villani SM. Exotic ambrosia beetles and phytopathogenic fungi associated with rapid apple decline in North Carolina. Journal of Economic Entomology 2025;118:3029–41. https://doi.org/10.1093/jee/toaf191
27. Gugliuzzo A, Criscione G, Biondi A, Aiello D, Vitale A, Polizzi G, Tropea Garzia G. Seasonal changes in population structure of the ambrosia beetle Xylosandrus compactus and its associated fungi in a southern Mediterranean environment. PLoS One 2020;15:e0239011. https://doi.org/10.1371/journal.pone.0239011