Seong-Eun Kim, Jae-Eui Cha, and Ahn-Heum Eom*
Department of Biology Education, Korea National University of Education, Cheongju, 28173, Korea
*Correspondence to eomah@knue.ac.kr
Korean Journal of Mycology (Kor J Mycol) 2026 March, Volume 54, Issue 1, pages 55-63.
https://doi.org/10.4489/kjm.2026.54.1.6
Received on February 23, 2026, Revised on March 23, 2026, Accepted on March 24, 2026, Published on March 31, 2026.
Copyright © The Korean Society of Mycology.
This is an Open Access article which is freely available under the Creative Commons Attribution-Non-Commercial 4.0 International License (CC BY-NC) (https://creativecommons.org/licenses/by-nc/4.0/).
Endophytic fungi inhabit healthy plant tissues without causing apparent disease symptoms and contribute to host fitness through diverse ecological functions. In this study, two endophytic fungal isolates obtained from asymptomatic leaves of Stephanandra incisa and Celtis sinensis in Korea were identified based on their morphological characteristics and multilocus phylogenetic analyses. The isolates were confirmed to be Microascus brunneosporus and Colletotrichum camelliae-japonicae, which are unrecorded fungal species in Korea. Their cultural, morphological, and phylogenetic characteristics are described. These findings expand our understanding of the endophytic fungal diversity in Korea.
Colletotrichum camelliae-japonicae, Endophytic fungi, Microascus brunneosporus, New record, Phylogenetic analysis
Endophytic fungi are defined as fungi that colonize healthy plant tissues without causing any apparent disease symptoms [1]. They are widely distributed across diverse plant taxa, most of which belong to the phylum Ascomycota [2]. Depending on the host species and environmental conditions, endophytic fungi may establish mutualistic, commensalistic, or latent pathogenic relationships with their hosts. Many endophytic fungi are known to enhance host growth and stress tolerance and to produce a variety of bioactive secondary metabolites [3].
Forest ecosystems, predominantly composed of woody plants, harbor diverse endophytic fungal communities. Endophytic fungi associated with woody plants may contribute to plant fitness by producing plant growth regulators, enzymes, and secondary metabolites, as well as by increasing resistance to biotic and abiotic stresses [4]. Stephanandra incisa (Thunb.) Zabel is a deciduous shrub belonging to the family Rosaceae that is distributed in Korea, China, and Japan and has been used as an ornamental and medicinal plant [5]. Celtis sinensis Pers., a deciduous tree belonging to the family Ulmaceae, is widely distributed in East Asia and has long been used in traditional medicine [6].
During a survey of endophytic fungi associated with native woody plants in Korea, two fungal isolates were obtained from the asymptomatic leaf tissues of S. incisa and C. sinensis. Based on morphological characteristics and multilocus phylogenetic analyses, these isolates were identified as Microascus brunneosporus and Colletotrichum camelliae-japonicae. To the best of our knowledge, these species have not been recorded previously in Korea. This study documents these two fungal species as new records in Korea and provides baseline information for future studies on endophytic fungal diversity.
On April 28, 2023, the asymptomatic leaves of S. incisa were collected from Gaphasan Mountain, Gapdong, Yuseong-gu, Daejeon, Korea. On July 13, 2023, asymptomatic leaves of C. sinensis were collected from Jeungdo-myeon, Sinan-gun, Jeollanam-do, Korea. The collected samples were placed in polyethylene bags and transported to the laboratory within 24 h.
Leaf samples were washed under running tap water to remove surface debris. Surface sterilization was performed by immersion in 35% H2O2 for 40 s, followed by 70% ethanol for 30 s, as previously described [7]. The sterilized leaf tissues were cut into approximately 0.5 × 1.5 cm segments and placed on potato dextrose agar (PDA; Difco Laboratories, Detroit, MI, USA). The plates were then incubated at 25°C in the dark until fungal growth was observed. Pure isolates were then cultured on PDA and malt extract agar (MEA; Kisan Bio, Seoul, Korea) and incubated at 25°C in the dark for 7 d. Colony morphology was examined using a stereomicroscope (SZX7, Olympus, Tokyo, Japan). Microscopic characteristics were observed by mounting the fungal structures on glass slides with lactic acid and examining them under a light microscope (Axio Imager A2; Carl Zeiss, Oberkochen, Germany). Measurements were based on at least 20 conidia.
Genomic DNA was extracted from fresh mycelia using the HiGene Genomic DNA Prep Kit (Biofact, Daejeon, Korea) according to the manufacturer’s instructions. The primer sets used for PCR amplification were as follows: internal transcribed spacer (ITS) region using ITS1F/ITS4 [8]; large subunit ribosomal DNA (LSU) using NL1/NL4 [9]; β-tubulin (TUB2) using Bt2a/Bt2b [10] and T1/Bt2b [10,11]; translation elongation factor 1-α (TEF) using EF1-983F/EF1-2218R [12]; actin (ACT) using ACT512F/ACT783R [12]; chitin synthase 1 (CHS-1) using CHS-79F/CHS-345R [13]; and glyceraldehyde-3-phosphate dehydrogenase (GAPDH) using GDF/GDR [14].
PCR amplification was performed with an initial denaturation at 95°C for 2 min, followed by 35 cycles of denaturation at 95°C for 20 s, annealing at primer-specific temperatures for 40 s, and extension at 72°C for 1 min, with a final extension at 72°C for 5 min. PCR products were verified by agarose gel electrophoresis and purified prior to Sanger sequencing (Macrogen Co., Ltd., Sejong, Korea).
The obtained sequences were compared with reference sequences in the National Center for Biotechnology Information (NCBI) database using BLAST to identify the closest related taxa and perform preliminary species identification based on sequence similarity. Sequence alignments and phylogenetic analyses were performed using MEGA version 11 [15]. Phylogenetic trees were constructed using the maximum likelihood method with 1,000 bootstrap replicates to assess branch support. Appropriate nucleotide substitution models were selected based on the best-fitting model. Newly identified fungal strains were deposited in the National Institute of Biological Resources (NIBR) and the corresponding DNA sequences were submitted to GenBank.
Multilocus phylogenetic analyses were performed to determine the taxonomic placement of the two study isolates. Based on the analyses of the ITS, LSU, TEF, and TUB2 regions, isolate KNUE 23N902 was identified as M. brunneosporus. Sequence comparison revealed a high similarity to the ex-type strain M. brunneosporus CBS 138276, with 99.2% (ITS), 98.5% (LSU), 98.5% (TEF), and 97.9% (TUB2) identity. In the ML phylogenetic tree inferred from the combined ITS–LSU–TEF–TUB2 dataset, the KNUE 23N902 isolate formed a strongly supported clade with the ex-type strain (bootstrap value of 100%) (Fig. 1), confirming its taxonomic identity. Similarly, isolate KNUE 23P320 was identified as C. camelliae-japonicae based on ITS, TUB2, ACT, CHS-1, and GAPDH sequence data. The isolate showed 99.8% (ITS), 100% (TUB2), 99.5% (ACT), 99.5% (CHS-1), and 99.5% (GAPDH) similarity with the ex-type strain C. camelliae-japonicae CGMCC 3.18118. In the ML phylogenetic analysis based on the combined ITS–TUB2–ACT–CHS-1–GAPDH dataset, isolate KNUE 23P320 clustered with the ex-type strain with moderately to well supported (78%) (Fig. 2), supporting its identification. These phylogenetic results clearly indicate that both isolates correspond to previously described species that have not been recorded in Korea.
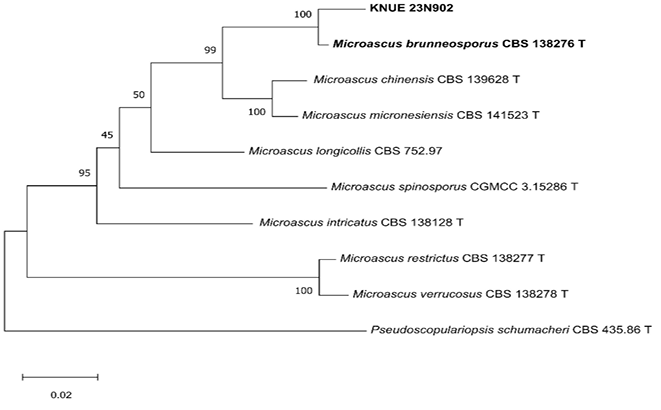
Fig. 1. A maximum likelihood phylogenetic tree (TN93+G) for Microascus brunneosporus KNUE 23N902 based on a concatenated alignment of internal transcribed spacer (ITS), large subunit ribosomal RNA (LSU), translation elongation factor 1-α (TEF), and β-tubulin (TUB2) sequences, with Pseudoscopulariopsis schumacheri used as an outgroup. The numbers on branches indicate bootstrap value (1,000 replicates) greater than 50%. The fungal strains isolated in this study are shown in bold type. T indicates an ex-type culture.
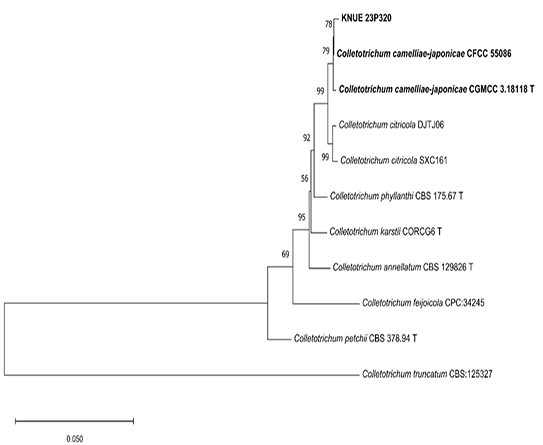
Fig. 2. A maximum likelihood phylogenetic tree (K2+G) for Colletotrichum camelliae-japonicae KNUE 23P320 based on a concatenated alignment of internal transcribed spacer (ITS), β-tubulin (TUB2), actin (ACT), chitin synthase genes (CHS-1), and the glyceraldehyde-3-phosphate dehydrogenase (GAPDH) sequences, with Colletotrichum truncatum used as an outgroup. The numbers on branches indicate bootstrap value (1,000 replicates) greater than 50%. The fungal strains isolated in this study are shown in bold type. T indicates an ex-type culture.
Morphological characteristics: After incubation at 25°C in the dark for 14 d on PDA, colonies were dark blue-gray on the surface and showed no pigmentation of the culture medium. The colonies exhibited an irregular outline with an uneven and wrinkled surface characterized by radially arranged folds. The colony texture was velvety with short, densely packed hyphae forming a thick mycelial mat covering the agar surface. On the reverse side, the colonies were similar in color to the surface and appeared to be embedded in the agar. On MEA, colony morphology was generally similar to that observed on PDA in terms of surface color, texture, and overall appearance on both the surface and reverse sides; however, colony growth on MEA was slower than that on PDA. Conidia were subglobose to navicular, hyaline to light brown, thin- and smooth-walled, measuring (4.5–)5.1(–6.0) × (1.7–)2.3(–3.3) μm (n = 20) (Fig. 3).
Specimen examined: Gaphasan Mountain, Gap-dong, Yuseong-gu, Daejeon, Korea (36.36228°N, 127.28507°E), April 28, 2023, isolated from a Stephanandra incisa (Thunb.) Zabel leaf, strain KNUE 23N902 (NIBRFGC000510701); GenBank accession numbers: PX869737 (ITS), PX869745 (LSU), PX883739 (TUB2), and PZ025072 (TEF).
Notes: The specific epithet brunneus is derived from the Latin word meaning “brown” referring to the coloration of the ascospores. This species was originally isolated from human bronchoalveolar lavage fluid. In the original description, both sexual and asexual morphs were reported [16], whereas only asexual morphs were observed in the present study. The culture characteristics of the isolate, particularly its morphology, size, and conidial chains, were consistent with those of the ex-type strain (Table 1).
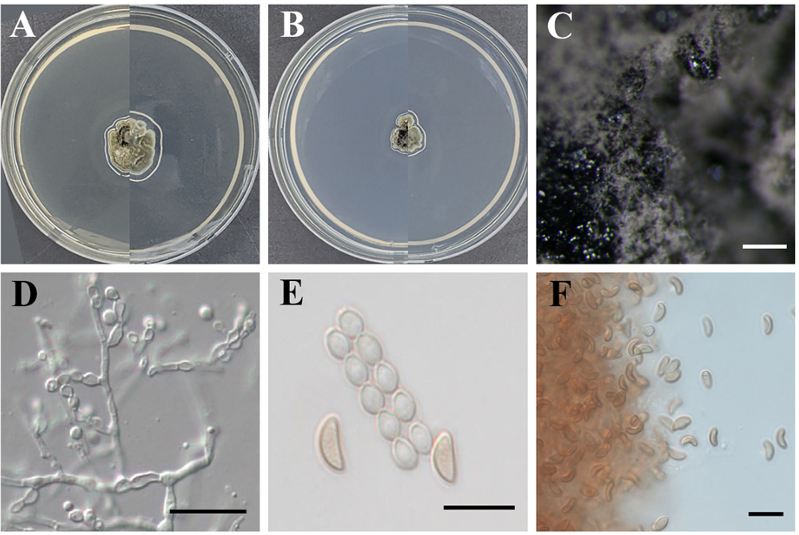
Fig. 3. Morphological characteristics of Microascus brunneosporus KNUE 23N902. Colonies grown for 14 d on potato dextrose agar (A) and malt extract agar (B). Conidiomata (C). Conidiophore (D). Conidia and conidial chain (E). Conidia (F). Scale bars: 200 μm (C), 20 μm (D), and 10 μm (E, F).
Table 1. Morphological characteristics of Microascus brunneosporus KNUE 23N902 compared with the original description [16]
| Strain | M. brunneosporus KNUE 23N902 | M. brunneosporus CBS 138276 [16] |
|---|---|---|
| Colony | PDA: 25 °C, 7 d; MEA: 25 °C, 14 d. | OA: 25 °C, 14 d; PCA: 25 °C, 14 d. |
| Color | PDA: grayish green; reverse grayish green MEA: dark grayish green; reverse dark grayish green. |
OA: dull green to olive brown; reverse dark green. PCA: dull green, with a white margin; reverse dull green. |
| Size | PDA: 11–19 mm; MEA: 9–14 mm. | OA: 21–25 mm; PCA: 15–17 mm. |
| Shape | PDA: irregularly wrinkled surface, velvety; reverse firmly embedded. MEA: irregularly wrinkled surface, velvety; reverse firmly embedded. |
OA: flat, velvety, granular at the center due to the presence of ascomata. PCA: slightly elevated, downy, fasciculate at the center. |
| Conidia | subglobose, navicular, hyaline, light brown, thin and smooth-walled, (4.5–)5.1(–6.0) × (1.7–)2.3(–3.3) μm (n=20). | subglobose, ellipsoidal or navicular, 4–5 × 2.5–5 μm, with truncate base, light green-brown, thin and smooth-walled, arranged in long chains. |
PDA: potato dextrose agar; MEA: malt extract agar; OA: oatmeal agar; PCA: potato carrot agar.
Morphological characteristics: After incubation at 25°C in the dark for 7 d, colonies on PDA reached 70–76 mm in diameter. The colonies grew radially and were circular. The surfaces of the colonies had a cottony texture. The colony color was brilliant orange-yellow at the center, with a white marginal zone, and black radial pigment streaks were observed on the surface. On MEA, the colonies reached 64–67 mm in diameter and grew radially and were circular. The marginal zone was white, whereas the central area was yellowish-white. The reverse side showed a coloration similar to that of the surface. The conidia were hyaline, oblong, unicellular, smooth-walled, and aseptate, measuring (11.6–)13.4(–15.5) × (5.2–)5.6(–6.3) μm (n = 20) (Fig. 4).
Specimen examined: Jeungdo, Jeungdo-myeon, Sinan-gun, Jeollanam-do, Korea (34.99026° N, 126.13743°E), July 13, 2023, isolated from a Celtis sinensis Pers. leaf, strain KNUE 23P320 (NIBRFGC000510705); GenBank accession numbers: PX938858 (ITS); PZ025073 (TUB2), PZ025075 (ACT), PZ025071 (CHS-1), and PZ025074 (GAPDH).
Notes: This species was originally isolated from Camellia japonica imported from Japan, and its specific epithet is derived from the scientific name of the host plant from which it was first described. It was detected during quarantine inspections conducted by the Ningbo Entry-Exit Inspection and Quarantine Bureau in China [17]. In the present study, this fungus was isolated as an endophyte from Celtis sinensis. The cultural characteristics and asexual conidial morphology were consistent with the original description (Table 2), including colony color changes on PDA owing to the production of orange conidial masses and similar conidial shape and dimensions.
M. brunneosporus was originally isolated from human bronchoalveolar lavage fluid [16]. However, in the present study, it was isolated from asymptomatic leaf tissues, suggesting its ecological versatility and ability to occupy diverse habitats. Although C. camelliae-japonicae is primarily recognized as a plant pathogen, it has also been reported to be an endophyte or saprobe [18,19], indicating ecological flexibility within the genus. The recovery of C. camelliae-japonicae from healthy leaf tissues in the present study further supports this hypothesis. Taken together, these findings expand the known geographic and ecological distributions of both species and contribute to the documentation of endophytic fungal diversity associated with woody plants in Korea.
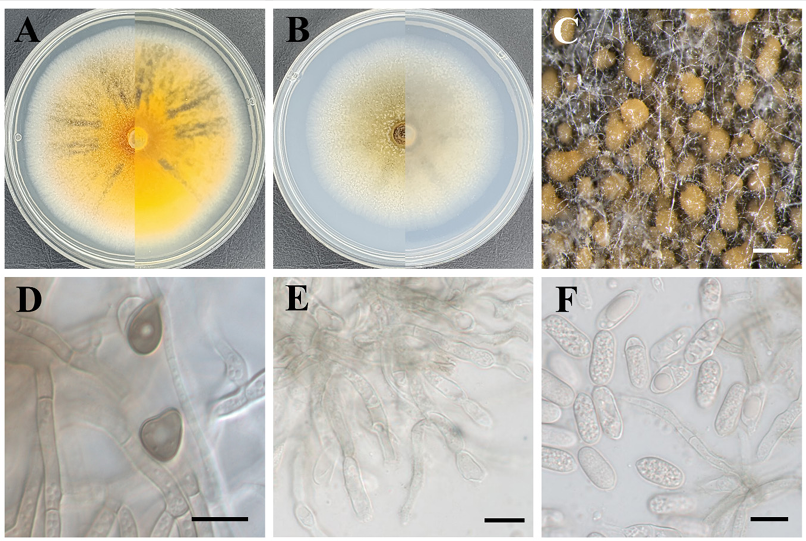
Fig. 4. Morphological characteristics of Colletotrichum camelliae–japonicae KNUE 23P320. Colonies grown for 7 d on potato dextrose agar (A) and malt extract agar (B). Conidiomata (C). Appressoria (D). Conidiophore (E). Conidia (F). Scale bars: 500 μm (C) and 10 μm (D, E, F).
Table 2. Morphological characteristics of Colletotrichum camelliae-japonicae KNUE 23P320 with the original description [17]
| Strain | C. camelliae-japonicae KNUE 23P320 | C. camelliae-japonicae CGMCC 3.18118 [17] |
|---|---|---|
| Colony | PDA: 25°C, 7 d. MEA: 25°C, 7 d. |
PDA: 25°C, 7 d. |
| Color | PDA: brilliant orange-yellow at the center, with a white marginal zone, and black radial pigment streaks were observed on the surface. MEA: marginal zone was white, central area was yellowish-white; reverse same coloration as the surface. |
SNA: surface covered with orange or pale yellow conidiomata; reverse hyaline. PDA: white, becoming grayish and finally covered with orange conidia mass. Reverse pale brown or grayish. |
| Size | PDA: 7.0–7.6 cm. MEA: 6.4–6.7 cm. |
PDA: 4–4.2 cm. |
| Shape | PDA: radially growing and circular in outline. surface was even, with a cottony texture. MEA: radially growing and circular. |
SNA: flat, lacking aerial mycelium. |
| Conidia | hyaline, oblong, unicellular, smooth-walled, and aseptate. (11.6–)13.4(–15.5) × (5.2–)5.6(–6.3) μm (n = 20). |
hyaline, oblong, single-celled, apex and base rounded, with a prominent scar, smooth-walled, aseptate, most contents granular or guttulate; 11–14.5 × 5–6.5 μm. |
PDA: potato dextrose agar; MEA: malt extract agar; SNA: synthetic nutrient-poor agar.
The authors declare no competing interests.
Funding: This study was supported by a grant from the National Institute of Biological Resources (NIBR202602103).
1. Suryanarayanan TS, Wittlinger SK, Faeth SH. Endophytic fungi associated with cacti in Arizona. Mycol Res 2005;109:635–9. https://doi.org/10.1017/s0953756205002753
2. Arnold AE. Understanding the diversity of foliar endophytic fungi: Progress, challenges, and frontiers. Fungal Biol Rev 2007;21:51–66. https://doi.org/10.1016/j.fbr.2007.05.003
3. Verma VC, Kharwar RN, Strobel GA. Chemical and functional diversity of natural products from plant associated endophytic fungi. Nat Prod Commun 2009;4:1511–32. https://doi.org/10.1177/1934578X0900401114
4. Hong L, Wang Q, Zhang J, Chen X, Liu Y, Asiegbu FO, Wu P, Ma X, Wang K. Advances in the beneficial endophytic fungi for the growth and health of woody plants. For Res 2024;4:e028. https://doi.org/10.48130/forres-0024-0025
5. Zhang CP, Xu ZZ, Xie XM, Lu YZ, Li W, Wang KL, Jiang XQ. The complete chloroplast genome sequence of Stephanandra incisa. Mitochondrial DNA B Resour 2021;6:363–4. https://doi.org/10.1080/23802359.2020.1867012
6. Wang L, Fang X, Hu Y, Zhang Y, Qi Z, Li J, Zhao L. Efficient extraction of bioactive flavonoids from Celtis sinensis leaves using deep eutectic solvent as green media. RSC Adv 2021;11:17924–35. https://doi.org/10.1039/d1ra01848e
7. Choi JW, Lee JM, Park SY, Eom AH. Acrodontium burrowsianum and Pestalotiopsis humicola: Two previously unrecorded fungal species isolated from conifer leaves in Korea. Kor J Mycol 2022;50:311–8. https://doi.org/10.4489/KJM.20220033
8. White TJ, Bruns TD, Lee SB, Taylor JW. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ, editors. PCR protocols: A guide to methods and applications. San Diego: Academic Press; 1990. p. 315–22. https://doi.org/10.1016/B978-0-12-372180-8.50042-1
9. Linton CJ, Borman AM, Cheung G, Holmes AD, Szekely A, Palmer MD, Bridge PD, Campbell CK, Johnson EM. Molecular identification of unusual pathogenic yeast isolates by large ribosomal subunit gene sequencing: 2 years of experience at the United Kingdom mycology reference laboratory. J Clin Microbiol 2007;45:1152–8. https://doi.org/10.1128/JCM.02061-06
10. Glass NL, Donaldson GC. Development of primer sets designed for use with the PCR to amplify conserved genes from filamentous ascomycetes. Appl Environ Microbiol 1995;61:1323–30. https://doi.org/10.1128/aem.61.4.1323-1330.1995
11. O’Donnell K, Cigelnik E. Two divergent intragenomic rDNA ITS2 types within a monophyletic lineage of the fungus Fusarium are nonorthologous. Mol Phylogenet Evol 1997;7:103–16. https://doi.org/10.1006/mpev.1996.0376
12. Rehner SA, Buckley E. A Beauveria phylogeny inferred from nuclear ITS and EF1-α sequences: Evidence for cryptic diversification and links to Cordyceps teleomorphs. Mycologia 2005;97:84–98. https://doi.org/10.1080/15572536.2006.11832842
13. Carbone I, Kohn LM. A method for designing primer sets for speciation studies in filamentous ascomycetes. Mycologia 1999;91:553–6. https://doi.org/10.1080/00275514.1999.12061051
14. Templeton MD, Rikkerink EH, Solon SL, Crowhurst RN. Cloning and molecular characterization of the glyceraldehyde-3-phosphate dehydrogenase-encoding gene and cDNA from the plant pathogenic fungus Glomerella cingulata. Gene 1992;122:225–30. https://doi.org/10.1016/0378-1119(92)90055-t
15. Tamura K, Stecher G, Kumar S. MEGA11: Molecular evolutionary genetics analysis version 11. Mol Biol Evol 2021;38:3022–7. https://doi.org/10.1093/molbev/msab120
16. Sandoval-Denis M, Gené J, Sutton DA, Cano-Lira JF, de Hoog GS, Decock CA, Wiederhold NP, Guarro J. Redefining Microascus, Scopulariopsis and allied genera. Persoonia 2016;36:136. https://doi.org/10.3767/003158516X688027
17. Hou LW, Liu F, Duan WJ, Cai L. Colletotrichum aracearum and C. camelliae-japonicae, two holomorphic new species from China and Japan. Mycosphere 2016;7:1111–23. https://doi.org/10.5943/mycosphere/si/2c/4
18. Jayawardena RS, Hyde KD, Damm U, Cai L, Liu M, Li XH, Zhang W, Zhao WS, Yan JY. Notes on currently accepted species of Colletotrichum. Mycosphere 2016;7:1192–260. https://doi.org/10.5943/mycosphere/si/2c/9
19. Cannon PF, Damm U, Johnston PR, Weir BS. Colletotrichum: Current status and future directions. Stud Mycol 2012;73:181–213. https://doi.org/10.3114/sim0014