Jin-Woo Park1, Ji-Hyun Park1,2,*, Lamiya Abasova3,4, In-Young Choi3,4,*, and Hyeon-Dong Shin2,5
1Department of Forest Resources, Kookmin University, Seoul 02707, Korea
2Department of Forestry, Environment, and Systems, Kookmin University, Seoul 02707, Korea
3Department of Plant Medicine, Jeonbuk National University, Jeonju 54896, Korea
4Research Center for Plant Medicine, Jeonbuk National University, Jeonju 54896, Korea
5Division of Environmental Science and Ecological Engineering, Korea University, Seoul 02841, Korea
*Correspondence to jhpark10@kookmin.ac.kr, iychoi@jbnu.ac.kr
Korean Journal of Mycology (Kor J Mycol) 2026 March, Volume 54, Issue 1, pages 77-82.
https://doi.org/10.4489/kjm.2026.54.1.8
Received on February 06, 2026, Revised on March 24, 2026, Accepted on March 30, 2026, Published on March 31, 2026.
Copyright © The Korean Society of Mycology.
This is an Open Access article which is freely available under the Creative Commons Attribution-Non-Commercial 4.0 International License (CC BY-NC) (https://creativecommons.org/licenses/by-nc/4.0/).
Erysiphe zelkovae is one of the main pathogens causing significant damage to Zelkova serrata in East Asia. Despite the global diversity of Zelkova spp., E. zelkovae has been reported only on Z. serrata from East Asian countries (Japan, China, and Korea). Mycological information regarding this fungus is insufficient in terms of sequenced nucleotides and detailed morphological characteristics. This study presents a detailed characterization of the holomorph of E. zelkovae and its taxonomic placement through molecular phylogenetic analyses using the internal transcribed spacer region and large subunit gene sequences from three representative Korean specimens. This study confirmed the identity of E. zelkovae as a distinct species among the Erysiphe section Uncinula species on host plants belonging to the family Ulmaceae.
Cannabaceae, Celtis, Erysiphe kusanoi, Ulmaceae, Uncinula zelkovae
The genus Zelkova Spach, belonging to the family Ulmaceae, consists of six species distributed across Europe and East Asia. Although Zelkova species are widespread, only Zelkova serrata (Thunb.) Makino has been reported as a host for powdery mildew fungus Erysiphe zelkovae (Henn.) U. Braun [as ʻzelkowae’], which is also encountered only in East Asian countries, such as Japan, China, and Korea [1]. Furthermore, E. zelkovae is the only known powdery mildew pathogen found on Zelkova globally. Traditionally, the powdery mildew pathogen of Z. serrata was identified as a variety of Uncinula kusanoi Syd. & P. Syd. [currently Erysiphe kusanoi (Syd. & P. Syd.) U. Braun & S. Takam.] based on the similarities in their morphological traits [2,3]. Japanese scientists followed this concept and recorded Zelkova powdery mildew as U. kusanoi var. zelkovae (P. Henn) U. Braun [4]. In Korea, E. zelkovae was first recorded as Uncinula zelkovae P. Henn. [5]. Shin [5] recognized U. zelkovae as a distinct species differing from U. kusanoi var. zelkovae on the basis of the number of ascospores in an ascus, while also emphasizing the close affinity of the anamorph morphology of the two powdery mildew species.
Erysiphe kusanoi is an obligate parasite of Celtis L. species [1]. Previously, Celtis spp. were classified within the elm (Ulmaceae) family, but were later transferred to the Cannabaceae (hemp) based on the molecular phylogenetic analysis [6]. Both the Ulmaceae and Cannabaceae families belong to the order Rosales; therefore, the molecular and morphological similarities of E. zelkovae and E. kusanoi can be understood within the framework of the phylogenetic relationship between host families. Considering the above-mentioned issues between these two powdery mildew species, detailed mycological characterization is required to distinguish them. Furthermore, a single sequence from a Japanese specimen (MUMH 403) for the internal transcribed spacer (ITS) and large subunit ribosomal DNA (LSU) regions provides insufficient molecular evidence to recognize E. zelkovae as a distinct species. Therefore, this study aimed to characterize the holomorph morphology of E. zelkovae in detail, to elucidate its molecular phylogenetic position using ITS and LSU sequence data derived from Korean specimens, and to compare its phylogenetic relationships with other closely related Erysiphe powdery mildews.
A total of 28 powdery mildew specimens associated with Z. serrata were collected from multiple localities in Korea and are currently preserved at the Korea University Herbarium (KUS). Accession numbers for the specimens are listed as follows: KUS-F15571 (Oct 27, 1998; Seoul), F15636 (Nov 03, 1998; Seoul), F17168 (Oct 22, 1998; Namyangju), F17795 (Oct 09, 2000; Dongducheon), F22395 (Oct 28, 2006; Jinju), F23620 (Aug 29, 2008; Gongju), F24645 (Sep 20, 2009; Mungyeong), F25070 (Jul 18, 2010; Jecheon), F25084 (Jul 19, 2010; Youngwol), F25252 (Aug 08, 2010; Dongducheon), F25511 (Oct 18, 2010; Pocheon), F27062 (Oct 06, 2012; Seoul), F27952 (Jul 25, 2014; Hongcheon), F28198 (Sep 22, 2014; Jinju), F28249 (Sep 25, 2014; Busan), F28349 (Oct 09, 2014; Miryang), F28389 (Oct 14, 2014; Hongcheon), F28549 (Nov 27, 2014; Ulsan), F28912 (Oct 12, 2015; Yangpyeong), F29630 (Oct 27, 2016; Yangpyeong), F30353 (Nov 09, 2017; Daegu), F30809 (Sep 11, 2018; Hongcheon), F31296 (Oct 15, 2019; Daegu), F33414 (Oct 23, 2022; Imsil), F33898 (Oct 23, 2023; Uijeongbu), F33905 (Oct 23, 2023; Seoul), F34384 (Oct 21, 2024; Namyangju), F34444 (Nov 15, 2024; Paju), and F34595 (Aug 21, 2025; Chuncheon).
The morphological features of the powdery mildew pathogen were examined using an Olympus BX50 light microscope (Olympus, Tokyo, Japan), and images were captured using a Zeiss AX10 microscope equipped with an AxioCam MRc5 camera (Carl Zeiss, Oberkochen, Germany). Microscopic observations and measurements were primarily conducted on fresh specimens. The lactic acid technique was used to examine the dried herbarium materials [7]. For each diagnostic structure, quantitative assessment was based on 30 measurements.
Mycelial mats with conidiophores and conidia were amphigenous and abundantly epigenous, forming circular to irregular white colonies (Fig. 1A). In the later stages of the disease, chasmothecia were formed abundantly on the lower side of the leaves (Fig. 1B, C). The appressoria on the mycelia were moderately lobed to multi-lobed (Fig. 1D, E). Conidiophores emerging from the side of the hyphae were 55–87 × 7–9 μm, and producing conidia singly, followed by 1–2(–3) cells (Fig. 1F–I). Foot cells of conidiophores were curved or L-shaped from the basal part, cylindrical, and 18–32 μm long. Conidia were hyaline, cylindrically ovoid to oblong, variable in shape and size, 26–36 × 13–16 μm (length/width ratio = 2.0–2.8), occupied by vacuoles and oil drops, and lacking distinct fibrosin bodies (Fig. 1J). Germ tubes were produced at the perihilar position of the conidia (Fig. 1K, L). Chasmothecia were amphigenous, abundantly hypogenous, scattered to gregarious, dark brown, depressed globose, and 91–125 μm in diameter, and containing 4–7 asci per chasmothecium (Fig. 1M, N). Chasmothecial appendages were positioned in the lower half of the chasmothecium, numerous, 12–24 in number, not branched, substraight to mildly curved, apex circinate to uncinate, 4–6 μm wide but up to 9 μm wide at the uncinate part, 1–1.5 times as long as the chasmothecial diameter, aseptate, and hyaline throughout (Fig. 1O). Asci were obovoid, saccate, subsessile, 42–56 × 32–42 µm, and moderately thick-walled, thinner at the upper part (Fig. 1P, Q). Ascospores were 3–4 in an ascus, oval to cylindrically ovoid, 24–30 × 11–14 µm (length/width ratio = 1.8–2.2), and containing oil drops (Fig. 1R, S).
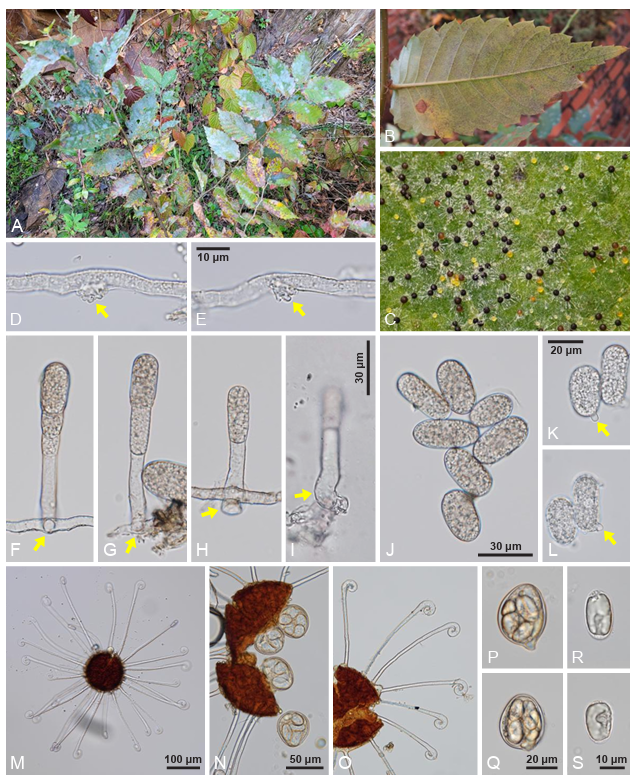

Fig. 1. Erysiphe zelkovae, a powdery mildew fungus found on Zelkova serrata (KUS-F34595). A: Powdery mildew symptoms on the leaves. B: Chasmothecia of E. zelkovae formed on the lower leaf surface. C: Close-up view of chasmothecia. Note yellowish, immature chasmothecia. D and E: Appressoria formed on the hyphae (arrows). F–I: Conidiophores. Note the lower part of conidiophores emerging from the side of creeping hyphae (arrows). J: Conidia. K and L: Conidia germinating (arrows). M: Chasmothecium with appendages. N: Asci protruding from the chasmothecium. O: Chasmothecial appendages with enlarged apex. P and Q: Mature asci. R and S: Ascospores.
To confirm the morphology-based identification, genomic DNA was obtained from three herbarium samples (KUS-F30809, F31296, and F33414). The LSU gene and ITS region, including the 5.8S rDNA gene, were amplified and sequenced using the primer pairs PM3/NLP2 and ITS1/PM6, respectively [8]. Forward and reverse reads were combined in MEGA11 [9], and the resulting sequences were submitted to the National Center for Biotechnology Information database (accession numbers: PX916213–PX916215 for ITS, and PX916216–PX916218 for LSU). Sequence similarity searches were performed against the GenBank database using BLASTn search tool. The results showed 100% identity in the ITS region with the sequences of E. zelkovae (AB475121) and in the LSU region with E. zelkovae (AB475114). The sequences of the LSU region were found to be 98.3% similar to those of E. nothofagi (AB378736) and 97.4% to Erysiphe sp. (OQ221121). To determine the phylogenetic placement of E. zelkovae among other closely related uncinuloid powdery mildews, a concatenated dataset of ITS and LSU sequences was assembled using MEGA11 for 20 Erysiphe sequences, including 17 sequences retrieved from GenBank. Sequence of Erysiphe patagoniaca Havryl. & S. Takam. (AB378747) was used as an outgroup taxon. Maximum parsimony (MP) analysis was conducted in PAUP* 4.0.a using the heuristic searches with the “tree-bisection-reconstruction” algorithm and 100 random sequence additions. All characters were treated as unordered and unweighted, and gaps were interpreted as missing data [10]. Tree statistics, including tree length, consistency index, retention index, and rescaled consistency index, were calculated. The analyzed alignment comprised 20 sequences spanning 870 characters, including 69 variable sites (7.9%) and 101 parsimony-informative sites (11.6%). Moreover, a maximum likelihood (ML) tree was constructed using raxmlGUI 2.0.13 based on the GAMMA model and GTR substitution [11]. Bootstrap (BS) support values were estimated from 1,000 replicates for both MP and ML analyses. In the inferred topology, all three sequences generated in this study formed a separate clade together with E. zelkovae, supported by 100% BS values from both analyses (Fig. 2). Based on the resulting phylogenetic tree presented in this study, it can be assumed that E. zelkovae and Erysiphe powdery mildews on Celtis spp. diverged from the same ancestor, which explains the similarities in their morphology.
Erysiphe zelkovae was first described as Uncinula zelkovae in 1901, based on a Japanese sample from Zelkova serrata [12]. Later, this species was reduced to a variety rank of U. kusanoi, as U. kusanoi var. zelkovae [2]. In accordance with the change in generic concepts within the Erysiphaceae, this fungus was renamed Erysiphe kusanoi var. zelkovae (Henn) U. Braun [12]. In 2009, Heluta et al. [13] provided the first sequence data for E. kusanoi var. zelkovae based on a Japanese specimen, suggesting that the Zelkova mildew pathogen (E. kusanoi var. zelkovae) be differentiated from the typical Celtis mildew pathogen (E. kusanoi var. kusanoi). No additional sequence data for Zelkova mildew were obtained. In 2012, the fungus was elevated to the species level as Erysiphe zelkovae based on the morphological differences from typical Erysiphe kusanoi and the taxonomic position of the host genera [1]. Therefore, the current study provides the first sequence data from Korean samples and supports the concept of E. zelkovae sensu Braun by confirming 99–100% similarity between the Japanese data and three Korean sequences. In conclusion, this study confirmed the identity of E. zelkovae as a distinct species in the genus Erysiphe section Uncinula species on diverse ulmaceous host plants [1,14].
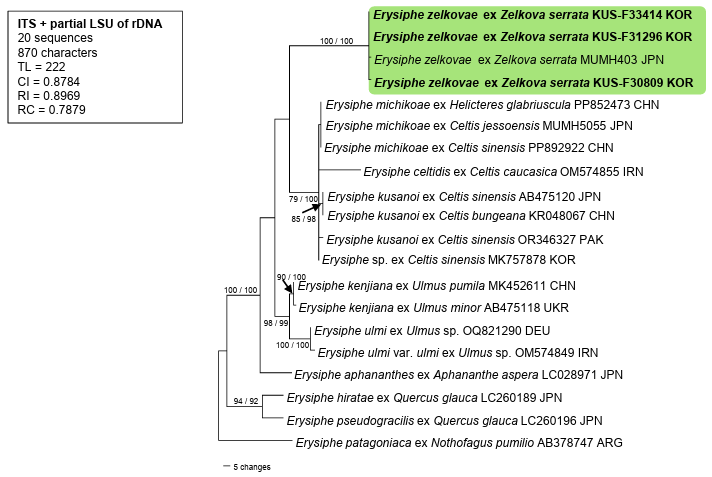
Fig. 2. Phylogenetic position of Erysiphe zelkovae among other closely related species of the Erysiphe sect. Uncinula is presented in the maximum parsimony tree derived from a combined data matrix of internal transcribed spacer (ITS) and large subunit ribosomal DNA (LSU) sequences. The sequence of E. patagoniaca (AB378747) is an outgroup here. Isolates obtained in this study are highlighted in bold. Bootstrap values (> 70%) obtained from the maximum parsimony and maximum likelihood analyses are indicated on the related branches, respectively. TL: tree length; CI: consistency index; RI: retention index; RC: rescaled consistency index.
The authors declare that there are no conflicts of interest.
This study was carried out with the support of the R&D Program for Forest Science Technology (Project No. RS-2024-00404816) provided by Korea Forest Service (Korea Forestry Promotion Institute).
2. Braun U. Taxonomic notes on some powdery mildews. Mycotaxon 1982;15:138–54.
3. Braun U. A monograph of the Erysiphales (powdery mildews). Beih Nova Hedwigia 1987;89:1–700.
4. Nomura Y. Taxonomic study of Erysiphaceae of Japan. Tokyo: Yokendo Ltd.; 1997.
5. Shin HD. Erysiphaceae of Korea. Suwon: National Institute of Agricultural Science and Technology; 2000.
6. The Angiosperm Phylogeny Group, Chase MW, Christenhusz MJM, Fay MF, Byng JW, Judd WS, Soltis DE, Mabberley DJ, Sennikov AN, Soltis PS, et al. An update of the Angiosperm Phylogeny Group classification for the orders and families of flowering plants: APG IV. Bot J Linn Soc 2016;181:1–20. https://doi.org/10.1111/boj.12385
7. Shin HD, La YJ. Morphology of edge lines of chained immature conidia on conidiophores in powdery mildew fungi and their taxonomic significance. Mycotaxon 1993;46:445–51.
8. Bradshaw M, Tobin PC. Sequencing herbarium specimens of a common detrimental plant disease (powdery mildew). Phytopathology 2020;110:1248–54. https://doi.org/10.1094/phyto04-20-0139-per
9. Tamura K, Stecher G, Kumar S. MEGA11: Molecular evolutionary genetics analysis version 11. Mol Biol Evol 2021;38:3022–7. https://doi.org/10.1093/molbev/msab120
10. Swofford DL. PAUP: Phylogenetic analyses using parsimony (and other methods) 4.0b10. Sunderland: Sinauer Associates; 2002.
11. Silvestro D, Michalak I. raxmlGUI: A graphical front-end for RAxML. Org Divers Evol 2012;12:335–7. https://doi.org/10.1007/s13127-011-0056-0
12. Braun U, Takamatsu S. Phylogeny of Erysiphe, Microsphaera, Uncinula (Erysipheae) and Cystotheca, Podosphaera and Sphaerotheca (Cystotheceae) inferred from rDNA ITS sequences – some taxonomic consequences. Schlechtendalia 2000;4:1–33.
13. Heluta V, Takamatsu S, Voytyuk S, Shiroya Y. Erysiphe kenjiana (Erysiphales), a new invasive fungus in Europe. Mycol Prog 2009;8:367–75. https://doi.org/10.1007/s11557-009-0610-8
14. Fragnière Y, Song YG, Fazan L, Manchester SR, Garfì G, Kozlowski G. Biogeographic overview of Ulmaceae: Diversity, distribution, ecological preferences, and conservation status. Plants 2021;10:1111. https://doi.org/10.3390/plants10061111