1Species Diversity Research Division, National Institute of Biological Resources, Incheon 22689, Korea
2Strategic Planning Division, National Institute of Biological Resources, Incheon 22689, Korea
*Correspondence to snubull@korea.kr
Korean Journal of Mycology (Kor J Mycol) 2025 June, Volume 53, Issue 2, pages 65-77.
https://doi.org/10.4489/kjm.2025.53.2.2
Received on April 16, 2025, Revised on May 28, 2025, Accepted on May 29, 2025, Published on June 30, 2025.
Copyright © The Korean Society of Mycology.
This is an Open Access article which is freely available under the Creative Commons Attribution-NonCommercial 4.0 International License (CC BY-NC) (https://creativecommons.org/licenses/by-nc/4.0/).
The genus Lachancea holds significant potential for industrial applications in fermentation due to its high sugar tolerance. Globally, 11 species of Lachancea have been identified, with only one species previously reported in Korea. However, research on the genus Lachancea in Korea remains limited. In this study, a phylogenetic analysis was performed on 58 Lachancea strains, including those isolated during yeast diversity studies in Korea and strains preserved at the National Institute of Biological Resources (NIBR). This analysis revealed four yeast species, three of which had not been previously reported in Korea. Physiological and morphological characterization of these three newly identified species confirmed their identity as being consistent with their respective type strains, establishing them as indigenous to Korea. This paper provides a taxonomic description of these newly reported Lachancea species—L. fermentati, L. kluyveri, and L. waltii—isolated from natural environments in Korea.
Lachancea, Unrecorded species, Yeast diversity
The genus Lachancea was proposed by Kurtzman (2003), following a reclassification of the “Saccharomyces complex” based on multigene sequence analyses [1]. The five species assigned to this novel genus are Kluyveromyces thermotolerans, K. waltii, Saccharomyces kluyveri, Zygosaccharomyces cidri, and Z. fermentati. Following the proposal of the Lachancea genus, several species have been added to it, including L. meyersii [2], L. dasiensis [3], L. nothofagi [4], L. mirantina [5], L. lanzarotensis [6], and L. quebecensis [7]. The 11 species currently comprising the genus Lachancea are characterized by the absence of true hyphae, 1–4 spherical ascospores, ability to ferment glucose and at least one other sugar, and an inability to assimilate nitrate [1,4]. The species of this genus are cosmopolitan and inhabit various niche, including soil, seawater, plants, animals, food, and even humans [8].
In recent years, the interest in application of non-Saccharomyces species in wine-making has increased, to generate higher quantities and greater diversities of metabolites [9]. In this regard, the most studied nonSaccharomyces species are Hanseniaspora uvarum, Lachancea thermotolerans, Metschnikowia pulcherrima, Schizosaccharomyces pombe, and Pichia kluyveri [10]. Lachancea thermotolerans is useful in oenological applications [11], and its various commercial strains, such as strain Kt 421 (Viniflora® CONCERTO™; Chr. Hansen, Hoersholm, Denmark), strain Laktia (LAKTIA™; Lallemand, Montreal, Canada), are currently available. The application of L. thermotolerans in wine fermentation is being investigated to address the problem of increasing sugar content and decreasing acidity of grapes due to global warming, which affects the acidity of wine [12]. Research has been conducted on the oenological characteristics of L. fermentati and L. lanzarotensis as well [13].
In Korea, Hyun et al. [14] isolated L. thermotolerans (synonym Kluyveromyces thermotolerans) from wild flowers, which was an unrecorded yeast species. Since then, L. thermotolerans has been reported in several regions and from various sources, including plants [15], soil, and water [16] in Korea. In addition to L. thermotolerans, other species such as L. cidri, L. fermentati, and L. kluyveri have also been isolated in Korea. However, information on their taxonomic characteristics or preserved strains remains scarce.
During our investigation into the diversity of indigenous yeasts in Korea, we found that the genus Lachancea accounted for the highest number of isolates in environments with high sugar concentrations (10%) and the presence of alcohol (5%), highlighting the physiological traits of this genus.
In this study, we investigated yeast strains isolated from natural environments and those preserved at the National Institute of Biological Resources (NIBR). Our objective was to identify the species of the genus Lachancea indigenous to Korea and to describe the taxonomic characteristics of the three unrecorded species: L. fermentati, L. kluyveri, and L. waltii.
A phylogenetic analysis was conducted on a total of 70 strains. These included 57 strains from the Lachancea genus isolated in this study, 1 strain preserved at the NIBR, and 12 strains selected from the data in the literature (Table 1). Considering the diversity of the isolated sources, five strains of three Lachancea species were selected as representative strains for morphological, physiological, and phylogenetic analyses.
Table 1. Strains and isolation sources of yeast species used to analyze the diversity within the Lachancea genus
Species | Strain | Source | Origin | NCBI Acc. No. | |
|---|---|---|---|---|---|
| ITS | LSU | ||||
| Lachancea cidri | CBS 5666 | Plant (Araucaria sp.) | Chile | KY103940 | KY108207 |
| NRRL Y-12634T | Beverage (cider) | France | NR165978 | NG055075 | |
| L. fermentati | NIBRFGC000500447 | Air | Gongju, CN | OQ326709 | OQ318459 |
| NIBRFGC000500449 | Air | Jeonju, JB | OQ326710 | OQ318460 | |
| NIBRFGC000500450 | Air | Jeonju, JB | PP588996 | PP589417 | |
| NIBRFGC000500487 | Plant (Rosa multiflora) | Jeonju, JB | OQ326712 | OQ318462 | |
| NIBRFGC000509731 | Air & insect | Incheon | OQ326721 | OQ318471 | |
| NIBRFGC000509732 | Air & insect | Incheon | OQ326722 | OQ318472 | |
| NIBRFGC000509733 | Air & insect | Incheon | OQ326723 | OQ318473 | |
| NIBRFGC000509734 | Plant (rotten wood) | Gwangju | OQ326724 | OQ318474 | |
| NIBRFGC000509735 | Plant (Crataegus pinnatifida) | Yangpyeong, GG | OQ326725 | OQ318475 | |
| NIBRFGC000509736 | Plant (Lindera obtusiloba) | Yangpyeong, GG | OQ326726 | OQ318476 | |
| NIBRFGC000511204 | Plant (Euscaphis japonica) | Jeju | PP589007 | PP589504 | |
| NIBRFGC000511205 | Plant (Aralia elata) | Jeju | PP589009 | PP589608 | |
| NIBRFGC000511216 | Plant (Huperzia serrata) | Jeju | PP589006 | PP589503 | |
| NIBRFGC000512338 | Plant (Liriope platyphylla) | Gochang, JB | PP589003 | PP589420 | |
| NIBRFGC000512339 | Plant (Commelina communis) | Gochang, JB | PP589010 | PP589419 | |
| NIBRFGC000512340 | Plant (Callicarpa japonica) | Gochang, JB | PP589002 | PP589418 | |
| NIBRFGC000512341 | Plant (Persicaria filiformis) | Gochang, JB | PP588997 | PP589421 | |
| NIBRFGC000512342 | Plant (Vicia unijuga) | Gochang, JB | PP589005 | PP589423 | |
| NIBRFGC000512343 | Plant (Vicia venosavar. cuspidata) | Gochang, JB | PP588994 | PP589359 | |
| NIBRFGC000512344 | Plant (Vicia venosavar. cuspidata) | Gochang, JB | PP588995 | PP589578 | |
| NIBRFGC000512345 | Plant (Agrimonia pilosa) | Gochang, JB | PP589000 | PP589373 | |
| NIBRFGC000512346 | Plant (Scutellaria indica) | Gochang, JB | PP588998 | PP589502 | |
| NIBRFGC000512347 | Plant (Platycarya strobilacea) | Gochang, JB | PP589004 | PP589422 | |
| NIBRFGC000512348 | Plant (Ligustrum obtusifolium) | Gochang, JB | PP589001 | PP589536 | |
| NIBRFGC000512372 | Air | Gwangyang, JN | PP588999 | PP589607 | |
| NIBRFGC000512373 | Air | Gwangyang, JN | PP589008 | PP589546 | |
| NRRL Y-1559T | Beverage (sediment of peppermint) | Unknown | NR130666 | NG055076 | |
| L. kluyveri | NIBRFGC000511155 | Plant (Pinus koraiensis) | Taebaek, GW | PP589027 | PP589424 |
| NRRL Y-12651T | Insect (Drosophila pinicola) | Unknown | NR138159 | NG055066 | |
| L.meyersii | CBS 8951T | Seawater | USA | AY645657 | AY645656 |
| CBS 9924 | Seawater | Bahamas | AY645659 | AY645658 | |
| L. nothofagi | CBS 11399 | Plant (Nothofagus antarctica) | Unknown | KY103991 | KY108256 |
| CBS 11611T | Plant (Nothofagus spp.) | Argentina | NR155331 | NG058329 | |
| L. thermotolerans | NIBRFGC000500451 | Air | Jeonju, JB | OQ326711 | OQ318461 |
| NIBRFGC000500466 | Plant (Dendranthema boreale) | Gongju, CN | MZ575158 | MZ575124 | |
| NIBRFGC000500467 | Plant (Smilax china) | Gongju, CN | MZ575159 | MZ575125 | |
| NIBRFGC000500468 | Plant (Dendranthema zawadskii var. latilobum) | Gongju, CN | MZ575157 | MZ575123 | |
| NIBRFGC000500488 | Plant (Rosa multiflora) | Jeonju, JB | OQ326713 | OQ318463 | |
| NIBRFGC000506493 | Mushroom (Geastrum sp.) | Sungju, GB | OQ326719 | OQ318469 | |
| NIBRFGC000509938 | Plant (Prunus padus) | Yangpyeong, GG | OQ326727 | OQ318477 | |
| NIBRFGC000509939 | Plant (Lindera obtusiloba) | Yangpyeong, GG | OQ326728 | OQ318478 | |
| NIBRFGC000509940 | Plant (Sorbus alnifolia) | Yangpyeong, GG | OQ326729 | OQ318479 | |
| NIBRFGC000509941 | Plant (Rubus crataegifolius) | Yangpyeong, GG | OQ326730 | OQ318480 | |
| NIBRFGC000509942 | Plant (Tilia mandshurica) | Yangpyeong, GG | OQ326731 | OQ318481 | |
| L. thermotolerans | NIBRFGC000500468 | Plant (Dendranthema zawadskii var. latilobum) | Gongju, CN | MZ575157 | MZ575123 |
| NIBRFGC000500488 | Plant (Rosa multiflora) | Jeonju, JB | OQ326713 | OQ318463 | |
| NIBRFGC000506493 | Mushroom (Geastrum sp.) | Sungju, GB | OQ326719 | OQ318469 | |
| NIBRFGC000509938 | Plant (Prunus padus) | Yangpyeong, GG | OQ326727 | OQ318477 | |
| NIBRFGC000509939 | Plant (Lindera obtusiloba) | Yangpyeong, GG | OQ326728 | OQ318478 | |
| NIBRFGC000509940 | Plant (Sorbus alnifolia) | Yangpyeong, GG | OQ326729 | OQ318479 | |
| NIBRFGC000509941 | Plant (Rubus crataegifolius) | Yangpyeong, GG | OQ326730 | OQ318480 | |
| NIBRFGC000509942 | Plant (Tilia mandshurica) | Yangpyeong, GG | OQ326731 | OQ318481 | |
| NIBRFGC000509943 | Plant (Zanthoxylum schinifolium) | Yangpyeong, GG | OQ326732 | OQ318482 | |
| NIBRFGC000509944 | Plant (Corylus heterophylla) | Yangpyeong, GG | OQ326733 | OQ318483 | |
| NIBRFGC000509945 | Air & insect | Incheon | OQ326734 | OQ318484 | |
| NIBRFGC000509946 | Plant (rotten wood) | Gwangju | OQ326735 | OQ318485 | |
| NIBRFGC000511147 | Plant (Arisaema amurense f. serratum) | Jeongsun, GW | PP589019 | PP589507 | |
| NIBRFGC000511148 | Plant (Rhamnus davurica) | Jeongsun, GW | PP589021 | PP589548 | |
| NIBRFGC000511164 | Plant (Sorbus commixta) | Yangpyeong, GG | PP589017 | ||
| NIBRFGC000511191 | Air & insect | Incheon | PP589020 | PP589547 | |
| NIBRFGC000511192 | Air & insect | Incheon | PP589018 | PP589505 | |
| NIBRFGC000511206 | Plant (Dendropanax trifidus) | Jeju | PP589023 | PP589610 | |
| NIBRFGC000511207 | Plant (Aralia elata) | Jeju | PP589025 | PP589613 | |
| NIBRFGC000511217 | Plant (Hydrangea serrata var. acuminata) | Jeju | PP589026 | PP589614 | |
| NIBRFGC000511218 | Plant (Callicarpa mollis) | Jeju | PP589024 | PP589611 | |
| NIBRFGC000512349 | Plant (Nandina domestica) | Gochang, JB | PP589014 | PP589609 | |
| NIBRFGC000512350 | Plant (Persicaria thunbergii) | Gochang, JB | PP589012 | PP589506 | |
| NIBRFGC000512351 | Plant (Dendranthema boreale) | Gochang, JB | PP589015 | PP589365 | |
| NIBRFGC000512352 | Plant (Lactuca indica) | Gochang, JB | PP589022 | PP589305 | |
| NIBRFGC000512353 | Plant (Rhamnus yoshinoi) | Gochang, JB | PP589016 | PP589579 | |
| NRRL Y-8284T | Food (mirabelle-plum conserve) | Russia | NR111334 | NG042626 | |
| L. waltii | NIBRFGC000136094 | Plant (Chaenomeles sinensis) | Daejeon | OQ326708 | OQ318458 |
| NIBRFGC000512379 | Plant (Aralia cordata var. continentalis) | Ulleung, GB | PP589011 | PP589442 | |
| NRRL Y-8285T | Plant (Ilex integra) | Japan | KY104011 | NG055070 | |
| Kluyveromyces dobzhanskii | CBS 2104T | Insect (Drosophila pseudoobscura) | USA | NR138156 | KY107999 |
| K. marxianus | NRRL Y-8281T | Unknown | Unknown | NR111251 | NG042627 |
The strains in bold were phenotypical characteristics examined. Type strains are denoted with the superscript ‘T’.
CN, Chungcheongnam-do; GB, Gyeongsangbuk-do; GG, Gyeonggi-do; GW, Gangwon-do; JB, Jeollabuk-do; JN, Jeollanam-do.
Species identification was performed by phylogenetic analysis of internal transcribed spacer (ITS) region and domains 1 and 2 (D1/D2) of the large subunit (LSU) rRNA gene. DNA extraction, PCR amplification, and cycle sequencing were performed as previously described [17]. The sequences were edited and aligned using the Geneious® 9 program (Biomatters, New Zealand). A phylogenetic tree was constructed using maximum-likelihood analysis performed with RAxML [18], implemented on the CIPRES web portal [19]. The analysis employed the general time reversible (GTR) model with 1,000 bootstrap replicates.
Five representative strains were examined for physiological (carbon source assimilation and oxidation) and biochemical properties following standard protocols [20]. Utilization of major carbon sources was determined using YT plates (Biolog Inc., Hayward, CA, USA). All experiments were conducted in triplicates.
Yeast strains were grown on yeast mold agar media (YMA) separately for 3 and 7 days each at 25℃; cell and colony morphology were observed using Nikon Eclipse 80i microscope (Nikon, Tokyo, Japan). Formation of hyphae and pseudohyphae were observed using Dalmau plates, after incubation for 2 weeks on cornmeal agar at 25℃. To induce sporulation, strains were incubated at 25℃ on four types of media- acetate agar (AA), cornmeal agar (CMA), 5% malt extract agar (MEA), and YMA.
We conducted a taxonomic examination of 58 Lachancea strains isolated from samples in Korea and identified four distinct species: L. fermentati, L. kluyveri, L. thermotolerans, and L. waltii (Table 1). A phylogenetic tree was constructed using concatenated sequences of ITS region and D1/D2 regions of the LSU rRNA gene (Fig. 1). The analysis revealed that the four species formed well-defined phylogenetic clades, each supported by bootstrap values close to 100%, thereby strongly confirming their classification as distinct taxa. Additionally, all strains exhibited high sequence similarity to the type strains in both the ITS and D1/D2 regions, with similarities exceeding 99.5%.
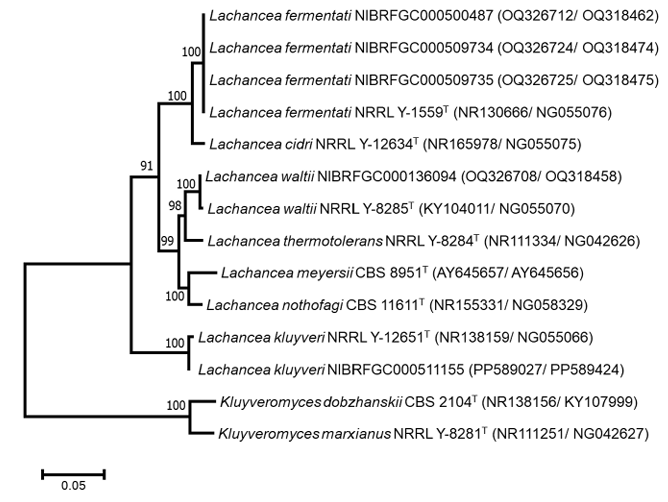
Fig. 1. Phylogenetic tree based on a maximum-likelihood analysis of combined internal transcribed spacer (ITS) and D1/D2 of the large subunit (LSU) rRNA gene sequences to determine the phylogenetic placement of Lachancea strains isolated in Korea and related species. Bootstrap values greater than 70% are indicated at the respective nodes. Kluyveromyces dobzhanskii and K. marxianus were used as outgroups. Strain numbers from culture collections follow the scientific names, and type strains are denoted with the superscript ‘T’. The scale bar represents 0.05 substitutions per site.
The strains also displayed morphological and physiological traits consistent with the type strains of the identified species. Among them, L. fermentati was unique in its ability to grow at 40℃, while L. kluyveri stood out for its capacity to utilize D-melibiose, a feature absent in the other three species. L. thermotolerans was the most frequently identified species, accounting for 23 of the 34 isolates, and was recovered from diverse substrates, including enrichment broths under trees, as well as plants (fruits, flowers, and tree stumps) and mushrooms. In contrast, L. waltii was distinguished by its inability to utilize turanose, maltose, or D-galactose (Table 2) and is considered a relatively rare species, with only a few isolates reported globally. Notably, even the CBS culture collection-one of the largest global repositories-contains only four preserved strains of L. waltii.
Table 2. Oxidation and assimilation of different carbon sources by three unregistered Lachancea yeast species in National Species List of Korea
| Carbon sources | L. fermentatia | L. kluyverib | L. waltiic | |
|---|---|---|---|---|
| Oxidation | ||||
| water | – | – | – | |
| acetic acid | – | – | – | |
| formic acid | – | – | – | |
| propionic acid | – | – | – | |
| succinic acid | – | – | – | |
| succinic acid mono-methyl ester | – | – | – | |
| L-aspartic acid | – | – | – | |
| L-glutamic acid | – | – | – | |
| L-proline | – | – | – | |
| D-gluconic acid | – | – | – | |
| dextrin | – | – | – | |
| inulin | – | – | – | |
| D-cellobiose | – | – | – | |
| gentiobiose | – | – | – | |
| maltose | + | + | – | |
| maltotriose | – | – | – | |
| D-melezitose | v | – | – | |
| D-melibiose | – | + | – | |
| palatinose | + | – | – | |
| D-raffinose | – | + | + | |
| stachyose | – | + | + | |
| sucrose | + | + | + | |
| D-trehalose | + | – | – | |
| turanose | + | + | – | |
| N-acetyl-D-glucosamine | – | – | – | |
| α-D-glucose | + | + | + | |
| D-galactose | + | + | – | |
| D-psicose | – | – | – | |
| L-sorbose | – | – | – | |
| salicin | – | – | – | |
| D-mannitol | – | – | – | |
| D-sorbitol | – | – | – | |
| D-arabitol | – | – | – | |
| xylitol | – | – | – | |
| glycerol | – | – | – | |
| tween 80 | – | – | – | |
| Assimilation | ||||
| water | – | – | – | |
| fumaric acid | – | – | – | |
| L-malic acid | – | – | – | |
| succinic acid mono-methyl ester | – | – | – | |
| bromo-succinic acid | – | – | – | |
| L-glutamic acid | – | – | – | |
| γ-amino-butyric acid | – | – | – | |
| α-keto-glutaric acid | – | – | – | |
| 2- keto-D-gluconic acid | – | – | – | |
| D-gluconic acid | – | – | – | |
| dextrin | – | – | – | |
| inulin | – | – | – | |
| D-cellobiose | – | – | – | |
| gentiobiose | – | – | – | |
| maltose | + | + | – | |
| maltotriose | – | – | – | |
| D-melezitose | – | – | – | |
| D-melibiose | – | + | – | |
| palatinose | + | – | – | |
| D-raffinose | v | + | + | |
| stachyose | – | + | + | |
| sucrose | + | + | + | |
| D-trehalose | + | – | – | |
| turanose | + | + | – | |
| N-acetyl-D-glucosamine | – | – | – | |
| D-glucosamine | – | – | – | |
| α-D-glucose | + | + | + | |
| D-galactose | + | + | – | |
| D-psicose | – | – | – | |
| L-rhamnose | – | – | – | |
| L-sorbose | – | – | – | |
| α-methyl-D-glucoside | + | – | – | |
| β-methyl-D-glucoside | – | – | – | |
| amygdalin | – | – | – | |
| arbutin | – | – | – | |
| salicin | – | – | – | |
| maltitol | + | – | – | |
| D-mannitol | – | – | – | |
| D-sorbitol | – | – | – | |
| adonitol | – | – | – | |
| D-arabitol | – | – | – | |
| xylitol | – | – | – | |
| i-erythritol | – | – | – | |
| glycerol | – | – | – | |
| tween 80 | – | – | – | |
| L-arabinose | – | – | – | |
| D-arabinose | – | – | – | |
| D-ribose | – | – | – | |
| D-xylose | – | – | – | |
| succinic acid mono-methyl ester plus D-xylose | – | – | – | |
| N-acetyl-L-glutamic acid plus D-xylose | – | – | – | |
| quinic acid plus D-xylose | – | – | – | |
| D-glucuronic acid plus D-xylose | – | – | – | |
| dextrin plus D-xylose | – | – | – | |
| α-D-lactose plus D-xylose | – | – | – | |
| D-melibiose plus D-xylose | – | + | – | |
| D-galactose plus D-xylose | + | + | – | |
| m-inositol plus D-xylose | – | – | – | |
| 1,2-propanediol plus D-xylose | – | – | – | |
| acetoin plus D-xylose | – | – | – | |
Examined strains: a, NIBRFGC000500487, NIBRFGC000509734, and NIBRFGC000509735; b, NIBRFGC000511155, c, NIBRFGC000136094. Growth reactions: +, strong growth; w, weak growth; v, variable; -, no growth.
Of the four Lachancea species, three (L. fermentati, L. kluyveri, and L. waltii) have not been previously recorded in Korea. Therefore, here we describe the taxonomic characteristics of these three Lachancea species, newly added to the National Species List of Korea (NSLK).
Lachancea fermentati (H. Nagan.) Kurtzman, FEMS Yeast Res 4(3):240 (2003)
The cells are broadly ellipsoidal after 3 days on YM agar at 25℃, 2.6–5.3 × 3.6–7.3 µm, and usually occur singly or in pairs (Fig. 2). Budding is by multilateral on a narrow base. After 1 week on YM agar at 25℃, colonies are smooth, butyrous, entire to weakly filamentous margin, and tannish-white colored. After 2 weeks of culture on Dalmau plates at 25℃, branched pseudohyphae but not true hyphae are formed. Asci are persistent and each ascus forms one to four spherical ascospores. Ascospores were observed on YM, CMA and MEA after 2 weeks at 25℃.
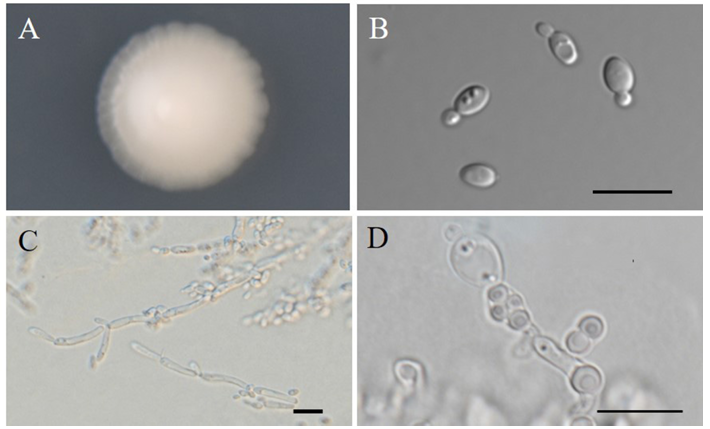
Fig. 2. Morphology of Lachancea fermentati NIBRFGC000500487. A: Colony on yeast mold (YM) agar 7 days at 25℃. B: Budding cells on YM agar 3 days at 25℃. C: Pseudohyphae on Dalmau plate with cornmeal agar for 2 weeks at 25℃. D: Ascospores on cornmeal agar 2 weeks at 25℃. Bars, 10 µm.
On the Biolog YT plate, the strains NIBRFGC000500487, NIBRFGC000509734 and NIBRFGC000509735 are positive for the oxidation of maltose, palatinose, sucrose, D-trehalose, turanose, α-D-glucose, and D-galactose. Assimilation of carbon compounds: maltose, palatinose, D-raffinose (variable), sucrose, D-trehalose, turanose, α-D-glucose, D-galactose, α-methyl-D-glucoside, maltitol, and D-galactose plus D-xylose (Table 2). Grow in 50% glucose, but not in 10% NaCl. All strains grow at 40℃, but not at 42℃. Growth occurs in the presence of 0.1% cycloheximide.
Examined strains: NIBRFGC000500487, Jeonju-si, Jeonbuk, Korea, 19 Oct. 2017, isolated from fruits of Rosa multiflora; NIBRFGC000509734, Gwangju, Korea, 09 Aug. 2022, isolated from tree stump; NIBRFGC000509735, Gapyeong-gun, Korea, 28 Jul. 2022, isolated from fruits of Crataegus pinnatifida.
Remarks: This species appears cosmopolitan and was isolated from diverse habitats such as plants, insects, and even beverages (tequila and kombucha). L. fermentati has been reported in the literature as Saccharomyces montanus and Zygosaccharomyces fermentati in Korea without a taxonomic description [21–23]. Saccharomyces montanus was isolated from Quercus varialilis [21] and fluid yogurt [22], and Z. fermentati was isolated from rotten potato [23]. In addition, one strain (CBS6772) isolated from a spoiled strawberry drink in Korea is being stored in CBS without reference.
Lachancea kluyveri (Phaff, M.W. Mill. & Shifrine) Kurtzman, FEMS Yeast Res 4(3):240 (2003)
The cells are globose to subglobose, 5.2–8.0 × 5.6–8.2 µm, occur singly or in pairs (Fig. 3). Cell division is by budding. Colonies are glossy or dull, cream colored to tan after 1 week on YM agar at 25℃. After 2 weeks of culture on Dalmau plates at 25℃, pseudohyphae may be formed, but true hyphae are not produced. Abundant ascosporulation was observed on acetate, cornmeal, 5% malt extract, YM agars after 3–5 days at 25℃. One to four spherical ascospores were formed in each ascus.
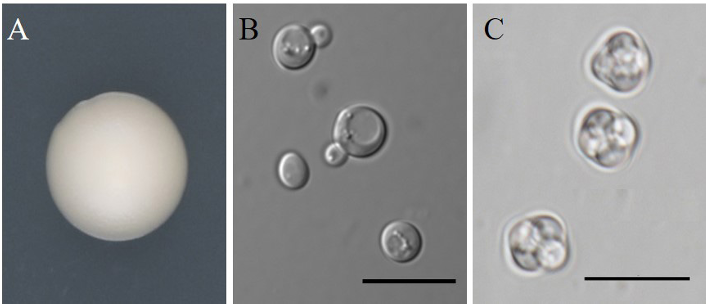
Fig. 3. Morphology of Lachancea kluyveri NIBRFGC000511155. A: Colony on yeast mold (YM) agar 7 days at 25℃. B: Budding cells on YM agar 3 days at 25℃. C: Ascospores on cornmeal agar 2 weeks at 25℃. Bars, 10 µm.
On the Biolog YT plate, the strain NIBRFGC000511155 is positive for the oxidation of maltose, maltotriose, D-melezitose, D-melibiose, palatinose, D-raffinose, stachyose, sucrose, D-trehalose, turanose, N-acetyl-D-glucosamine, α-D-glucose, and D-galactose. Assimilation of carbon compounds: maltose, D-melibiose, D-raffinose, stachyose, sucrose, turanose, α-D-glucose, D-galactose, D-melibiose plus D-xylose, and D-galactose plus D-xylose (Table 2). Grow in 50% glucose, but not in 10% NaCl. The maximum temperature for growth is 37℃. Sensitive to 0.01% cycloheximide.
Examined strain: NIBRFGC000511155, Jeongseon-gun, Korea, 14 Sep. 2022, isolated from the cone of Pinus koraiensis.
Remarks: The type strain of L. kluyveri was isolated from Drosophila sp. in the USA [24]. Strains of this species were mainly isolated from environmental samples such as soils, insects, and plants, and it exhibits a wide distribution, considering it was isolated from the USA, Sweden, Russia, and Spain. This species was reported in Korea as Saccharomyces kluyveri isolated from Diospyros kaki [25] and Meju [26], a raw material used to make Korean soy sauce.
Lachancea waltii (K. Kodama) Kurtzman, FEMS Yeast Res 4(3):240 (2003)
The cells are spherical, subglobose or broadly ellipsoidal after 3 days on YM agar at 25℃, 3.3–5.9 × 3.8–6.8 µm, occur singly or in pairs (Fig. 4). Multilateral budding on a narrow base. Colonies are butyrous, smooth, sometimes umbonate, glossy and cream colored with entire margin after 1 week on YM agar at 25℃. After 2 weeks of culture on Dalmau plates at 25℃, rudimentary pseudohyphae are occasionally formed while septated hyphae are absent. Ascospores are observed on acetate, cornmeal, 5% malt extract, YM agars after 2–4 weeks at 25℃. One to four spherical ascospores are formed from each ascus.
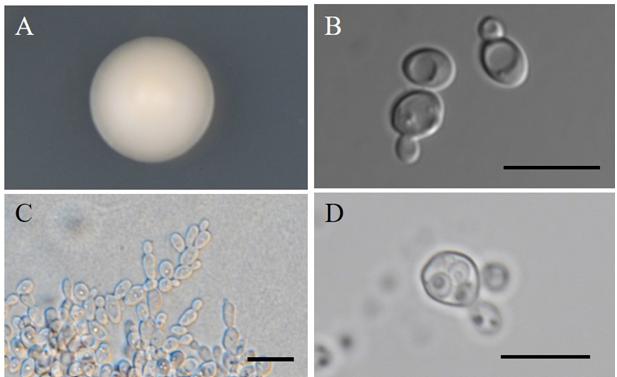
Fig. 4. Morphology of Lachancea waltii NIBRFGC000136094. A: Colony on yeast mold (YM) agar 7 days at 25℃. B: Budding cells on YM agar 3 days at 25℃. C: Pseudohyphae on Dalmau plate with cornmeal agar for 2 weeks at 25℃. D: Ascospores on cornmeal agar 2 weeks at 25℃. Bars, 10 µm.
On the Biolog YT plate, the strain NIBRFGC000136094 is positive for the oxidation of D-raffinose, stachyose, sucrose, and α-D-glucose (Table 2). Assimilation of carbon compounds: D-raffinose, stachyose, sucrose, and α-D-glucose. Grow in 50% glucose, but not in 10% NaCl. The strain grows 37℃, but not in 40℃. The strain is sensitive to 0.01% cycloheximide.
Examined strain: NIBRFGC000136094, Daejeon, Korea, 01 Sep. 2013, isolated from unripe fruits of Chaenomeles sinensis.
Remarks: Lachancea waltii has been isolated from Japan and Canada [27]. Through this study, it has been revealed that this species inhabits Korea. This species was mainly isolated from fruit, oak gall, and black knot of plants (Ilex integra and Prunus virginiana).
The authors declare no competing interests.
This research was supported by the National Institute of Biological Resources (NIBR202402104, NIBR202502103) under the Ministry of Environment, Republic of Korea.
1. Kurtzman CP. Phylogenetic circumscription of Saccharomyces, Kluyveromyces and other members of the Saccharomycetaceae, and the proposal of the new genera Lachancea, Nakaseomyces, Naumovia, Vanderwaltozyma and Zygotorulaspora. FEMS Yeast Res 2003;4:233-45. [DOI]
2. Fell JW, Statzell-Tallman A, Kurtzman CP. Lachancea meyersii sp. nov., an ascosporogenous yeast from mangrove regions in the Bahama Islands. Stud Mycol 2004;50:359-63.
3. Lee CF, Yao CH, Liu YR, Hsieh CW, Young SS. Lachancea dasiensis sp. nov., an ascosporogenous yeast isolated from soil and leaves in Taiwan. Int J Syst Evol Microbiol 2009;59:1818-22. [DOI]
4. Mestre MC, Ulloa JR, Rosa CA, Lachance MA, Fontenla S. Lachancea nothofagi sp. nov., a yeast associated with Nothofagus species in Patagonia, Argentina. Int J Syst Evol Microbiol 2010;60:2247-50. [DOI]
5. Pereira LF, Costa Jr CRL, Brasileiro BTRV, de Morais Jr MA. Lachancea mirantina sp. nov., an ascomycetous yeast isolated from the cachaça fermentation process. Int J Syst Evol Microbiol 2011;61:989-92. [DOI]
6. González SS, Alcoba-Flórez J, Laich F. Lachancea lanzarotensis sp. nov., an ascomycetous yeast isolated from grapes and wine fermentation in Lanzarote, Canary Islands. Int J Syst Evol Microbiol 2013;63:358-63. [DOI]
7. Freel KC, Charron G, Leducq JB, Landry CR, Schacherer J. Lachancea quebecensis sp. nov., a yeast species consistently isolated from tree bark in the Canadian province of Québec. Int J Syst Evol Microbiol 2015;65:3392-9. [DOI]
8. Bellut K, Krogerus K, Arendt EK. Lachancea fermentati strains isolated from kombucha: Fundamental insights, and practical application in low alcohol beer brewing. Front Microbiol 2020;11:764. [DOI]
9. Vicente J, Navascués E, Calderón F, Santos A, Marquina D, Benito S. An integrative view of the role of Lachancea thermotolerans in wine technology. Foods 2021;10:2878. [DOI]
10. Jolly NP, Varela C, Pretorius IS. Not your ordinary yeast: Non-Saccharomyces yeasts in wine production uncovered. FEMS Yeast Res 2014;14:215-37. [DOI]
11. Ministry of Food and Drug Safety. Standards and specifications for each food product [Internet]. Cheongju: MFDS; 2022 [cited 2025 Apr 15]. Available from: https://various.foodsafetykorea. go.kr/fsd/#/ext/Document/FC?searchNm=thermotol&itemCode=FC0A074001002A166.
12. Vaquero C, Izquierdo-Cañas PM, Mena-Morales A, Marchante-Cuevas L, Heras JM, Morata A. Use of Lachancea thermotolerans for biological vs. chemical acidification at pilot-scale in white wines from warm areas. Fermentation 2021;7:193. [DOI]
13. Porter TJ, Divol B, Setati ME. Investigating the biochemical and fermentation attributes of Lachancea species and strains: Deciphering the potential contribution to wine chemical composition. Int J Food Microbiol 2019;290:273-87. [DOI]
14. Hyun SH, Mun HY, Lee HB, Kim HK, Lee JS. Isolation of yeasts from wild flowers in Gyonggi-do province and Jeju island in Korea and the production of anti-gout xanthine oxidase inhibitor. Microbiol Biotechnol Lett 2013;41:383-90. [DOI]
15. Ahn C, Kim M, Kim C. Zygotorulaspora cornina sp. nov. and Zygotorulaspora smilacis sp. nov., two novel ascomycetous yeast species isolated from plant flowers and fruits. Mycobiology 2021;49:521-6. [DOI]
16. Han SM, Lee SY, Lee JS. Isolation of wild yeasts from humus-rich soil in city park of Daejeon metropolitan city, Korea, and characterization of the unrecorded wild yeasts. Kor J Mycol 2018;46:75-82.
17. Ahn C, Kim S, Kim C. Taxonomic study on six yeast species unlisted in the national species list of Korea. Kor J Mycol 2023;51:7-24.
18. Stamatakis A. RAxML-VI-HPC: Maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 2006;22:2688-90. [DOI]
19. Miller MA, Pfeiffer W, Schwartz T. Creating the CIPRES Science Gateway for inference of large phylogenetic trees. In: 2010 Gateway Computing Environments Workshop (GCE); New Orleans, LA, USA. New York: IEEE; 2010. p. 1-8. [DOI]
20. Kurtzman CP, Fell JW, Boekhout T, Robert V. Methods for isolation, phenotypic characterization and maintenance of yeasts. In: Kurtzman CP, Fell JW, Boekhout T, editors. The yeasts: a taxonomic study, 5th ed. Amsterdam: Elsevier; 2011. p. 87-110. [DOI]
21. Park MS. Studies on the wild yeasts in Korea(III): Investigation of wild yeast distribution. Korean J Microbiol 1972;10:51-68.
22. Suh D, Hwang I. Isolation and identification of yeasts occurred in inflated fluid yogurts. Korean J Microbiol Biotechnol 1987;15:15-20.
23. Jeong YJ, Kim OM, Seo JH, Lee MH, Jung SH, Kim TH. Characteristics of alcohol fermentation yeast isolated from potatoes. Korean J Postharvest Sci Technol 2000;7:228-32.
24. Phaff HJ, Miller MW, Shifrine M. The taxonomy of yeasts isolated from Drosophila in the Yosemite region of California. Antonie van Leeuwenhoek 1956;22:145-61. [DOI]
25. Rhee C, Park H. Isolation and characterization of alcohol fermentation yeasts from persimmon. Korean J Microbiol Biotechnol 1997;25:266-70.
26. Lee JS, Yi SH, Kwon SJ, Ahn C, Yoo JY. Isolation, identification and cultural conditions of yeasts from traditional meju. Korean J Microbiol Biotechnol 1997;25:435-41.
27. Lachance MA, Kurtzman CP. Lachancea Kurtzman (2003). In: Kurtzman CP, Fell JW, Boekhout T, editors. The yeasts: a taxonomic study, 5th ed. Amsterdam: Elsevier; 2011. p. 511-9. [DOI]