Tae-ho Kim, Jae-Eui Cha, Yun-Jeong Kim, Ji-Won Kim, and Ahn-Heum Eom*
Department of Biology Education, Korea National University of Education, Cheongju 28173, Korea
*Correspondence to eomah@knue.ac.kr
Korean Journal of Mycology (Kor J Mycol) 2025 June, Volume 53, Issue 2, pages 127-135.
https://doi.org/10.4489/kjm.2025.53.2.8
Received on May 02, 2025, Revised on June 12, 2025, Accepted on June 12, 2025, Published on June 30, 2025.
Copyright © The Korean Society of Mycology.
This is an Open Access article which is freely available under the Creative Commons Attribution-NonCommercial 4.0 International License (CC BY-NC) (https://creativecommons.org/licenses/by-nc/4.0/).
We report two previously unrecorded endophytic fungal species belonging to the order Xylariales—Hypoxylon vinosopulvinatum and Apiospora sasae—collected from Torreya nucifera and Phyllostachys nigra var. henonis, respectively, in Korea. Morphological characteristics and phylogenetic analyses based on internal transcribed spacer region, large subunit ribosomal RNA gene, β-tubulin gene, and translation elongation factor 1-α region confirmed the isolates as H. vinosopulvinatum and A. sasae, respectively. These findings expand our knowledge of the fungal biodiversity in Korean forest ecosystems and highlight the ecological significance of endophytic Xylariales.
Apiospora sasae, Biodiversity, Endophytic fungi, Hypoxylon vinosopulvinatum
The order Xylariales, belonging to the class Sordariomycetes, plays a key ecological role as wood decomposers, endophytes, and pathogens [1]. Early taxonomic studies on this order were primarily based on morphological characteristics. In the 19th century, the Tulasne brothers provided foundational systematic records of asci and ascospore structures [2]. However, morphological characteristics alone were insufficient to resolve interspecific variations within the order. The advent of molecular techniques has significantly reshaped Xylariales classification. Initial phylogenetic analyses have revealed fluctuating numbers of families within this order [2]. More comprehensive, recent multilocus studies have confirmed the monophyly of the order as well as led to major systematic revisions [2,3]. Continued phylogenetic investigations have resulted in the reclassification of several genera and new genera and species continue to be described [4,5].
Xylariales are generally characterized by darkly pigmented stromata and embedding perithecia [6]. For instance, the closely related families, Xylariaceae and Hypoxylaceae, typically produce brown, unicellular ascospores, often bearing a mucilaginous sheath or a conspicuous germ slit [7]. In contrast, species belonging to the families Diatrypaceae and Apiosporaceae frequently exhibit hyaline ascospores and poorly developed stromata [2]. These morphological characteristics are key diagnostic features for species identification. Recent phylogenetic studies using genetic markers such as internal transcribed spacer (ITS), large subunit (LSU), β-tubulin (TUB), and elongation factor 1-α (EF1-α) regions have been actively conducted [8].
Endophytic fungi colonize the internal plant tissues without causing apparent disease symptoms and are found in a wide range of host plants worldwide [9,10]. These fungi engage in mutualistic interactions with their hosts, contributing to plant growth, resistance to pathogens, and environmental stress tolerance [4,11]. Several members of Xylariales play important roles in these relationships [10].
Despite the diverse forest ecosystems and favorable climatic conditions for Xylariales in Korea, studies on endophytic fungi within this order remain limited and are often limited to a few genera or species, thereby highlighting a considerable gap in regional mycological research [12]. In this study, we report the morphological and molecular phylogenetic characteristics of two previously unrecorded endophytic fungi in Korea, namely Hypoxylon vinosopulvinatum and Apiospora sasae.
The samples were collected between December 2022 and July 2023. Leaves and twigs of Torreya nucifera (L.) Siebold & Zucc. were collected in December 2022 from Seoqwipo-si, Jeju-do (33°18′51.484″N, 126°27′ 51.962″E). The leaves and twigs of Phyllostachys nigra var. henonis (Mitford) Stapfex Rendle were collected in July 2023 from Hwasun-gun, Jeollanam-do (35°9′17.899″N, 127°10′8.353″E). Healthy leaves and twigs without disease symptoms or damage were collected and transported to the laboratory within 24 h.
The plant samples were rinsed with distilled water and surface sterilization was performed by immersion in 35% hydrogen peroxide for 1 min and 70% ethanol for 2 min, followed by rinsing with sterile distilled water [13]. Sterilized plant tissues were cut into 1.5 cm segments and placed on potato dextrose agar (PDA; Difco Lab., Detroit, USA). The samples were incubated at 25℃ in the dark to promote fungal growth.
Mycelia emerging from the plant tissues were subcultured to obtain pure isolates. These isolates were grown on PDA and malt extract agar (MEA; Kisan Bio, Seoul, Korea) at 25℃ in the dark to observe colony morphology. The slide culture method was used for microscopic observations. To induce spore formation, sterilized needles of Pinus densiflora Siebold & Zucc. were placed on 2% water agar and synthetic nutrient-poor agar (SNA) along with the fungal strains [14,15]. Colony morphology was recorded visually, and conidiomata and other microscopic structures were examined using a light microscope (Axio Imager A2; Carl Zeiss, Oberkochen, Germany).
Genomic DNA was extracted from fungal mycelial cultures using the HiGene Genomic DNA Prep Kit (BioFACT, Daejeon, Korea). Four DNA regions were amplified using polymerase chain reaction (PCR) for molecular phylogenetic analysis. The following primers sets were used for PCR amplification of fungal DNA regions: ITS1F/ITS4 for ITS [16,17], LR0R/LR5 for LSU [18], T1/T22 for TUB [19], and EF1983F/EF1-1567R for EF1-α [20]. The EF1-α region was only amplified for H. vinosopulvinatum, whereas the EF1-α region was used instead for A. sasae
DNA sequencing was performed by Solgent Co., Ltd. (Daejeon, Korea), using the Sanger method. The obtained sequences were compared to those in the National center for Biotechnology Information (NCBI) database using the Basic Local Alignment Search Tool (BLAST). Phylogenetic trees were constructed using the maximum-likelihood (ML) method implemented in MEGA11 software with bootstrap support from 1,000 replicates [21].
Morphological characteristics of strain KNUE23N245: After 7 d of growth at 25℃ in the dark, colonies grown on MEA reached 48.4–57.3 mm in diameter and exhibited a circular shape with concentric rings. The hyphae radiated outward from the center in a sequence of strong greenish-yellow, moderate olive, and white cottony hyphae. Similar concentric color patterns were observed on the reverse side, following the same sequence from the center outward (Fig. 1A). On PDA, colonies measured 32.6–40.1 mm in diameter and also exhibited a circular shape. Colonies developed outward from the center, showing strong greenish-yellow and white zones. Grayish olive aerial mycelia formed in the central region, and the margin was surrounded by white cottony hyphae. The reverse side of the colony appeared dark olive in the center, which gradually faded toward the edge, with the margin showing a cottony-white texture (Fig. 1B). The conidiophores were branched and brown (Fig. 1D). Conidia were obovate to ellipsoid, hyaline, and smooth surface measuring (4.21–) 5.67 (–7.67) × (1.93–) 2.58 (–3.34) µm (n = 20) (Figs. 1E and F).
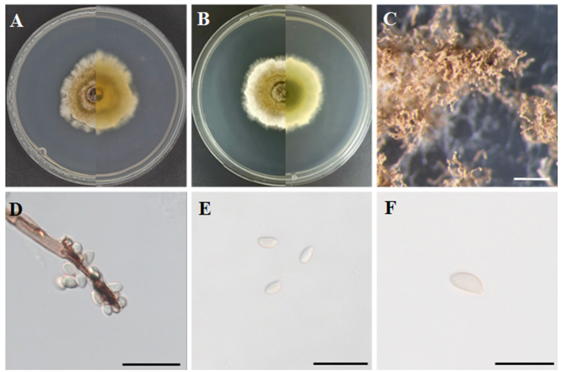
Fig. 1. Morphology of Hypoxylon vinosopulvinatum KNUE 23N245. (A, B) Colonies after 7 d of growth at 25℃ on malt extract agar (MEA; A) and potato dextrose agar (PDA; B). The left side of each image shows the front view, and the right side shows the reverse view. (C) Conidiomata (scale bar = 1 mm), (D) conidiophore (scale bar = 20 µm), (E) conidia (scale bar = 20 µm), and (F) conidia (scale bar = 10 µm).
Specimen examined: Seoqwipo-si, Jeju, Korea, 33°18’51.484″N 126°27’51.962″E, December 1, 2022, H. vinosopulvinatum, isolated from the leaf of T. nucifera, strain KNUE23N245 (NIBRFGC000510696), GenBank No. PV384440 (ITS), PV384442 (LSU), and PV454694 (TUB).
Phylogenetic analysis: The ITS sequence of KNUE23N245 showed 98.5% similarity with H. vinosopulvinatum FACTAS 4374 (OQ316446), the LSU sequence showed 99.9% similarity with FACTAS 4374 (OQ348552), and the TUB sequence showed 99.85% similarity with FACTAS4374 (OQ303966). ML phylogenetic trees were constructed from the combined ITS and LSU sequences, as well as a tree based on the TUB sequence. KNUE23N245 clustered with H. vinosopulvinatum voucher FCATAS4374 (Figs. 2 and 3).
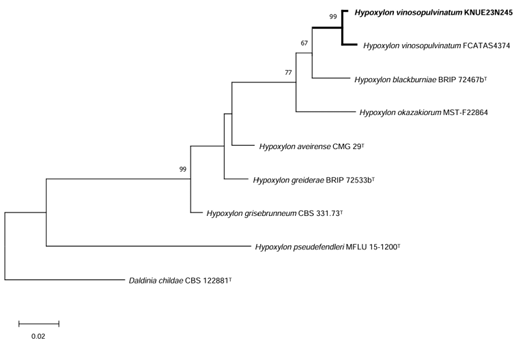
Fig. 2. Maximum-likelihood (ML) phylogenetic tree of Hypoxylon vinosopulvinatum KNUE 23N245. The tree was constructed based on concatenated sequences of internal transcribed spacer (ITS) and large subunit rDNA (LSU) regions. Daldinia childae was used as an outgroup. Numbers on the nodes indicate bootstrap values of >50% (1,000 replicates).
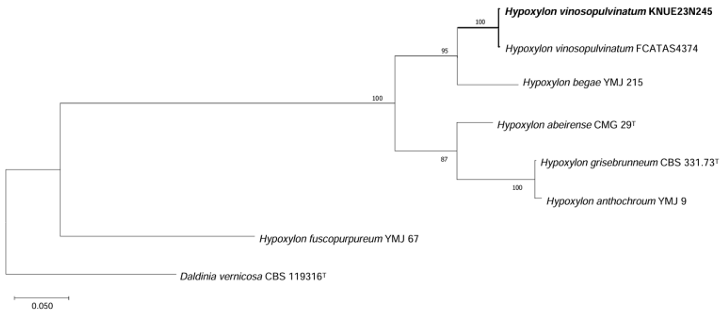
Fig. 3. Maximum-likelihood (ML) phylogenetic tree of Hypoxylon vinosopulvinatum KNUE 23N245. The tree was constructed based on β-tubulin (TUB) gene sequences. Hypoxylon fuscopurpureum was used as an outgroup. Numbers on the nodes indicate bootstrap values of >50% (1,000 replicates).
Notes: H. vinosopulvinatum was first collected from dead wood in 1988, as described by Ju. and Rogers in 1996 and 1999 [22]. However, the holotype was not designated because of the limited quantity and quality of the original specimens. In 2001, Ju and Hsieh recorded this species from decorticated wood in Taiwan, which was redescribed in 2004 [22]. H. vinosopulvinatum is currently classified within the family Hypoxylaceae, which is characterized by pigmented stromata containing perithecia, brown to hyaline ascospores typically with germ slits, and diverse anamorphic states [3, 6]. Although most species of genus Hypoxyloncea possess virgariella-like conidiogenous structures, H. vinosopulvinatum exhibits a nodulisporium-like structure, which was also observed in KNUE23N245 [22]. Although the conidial size was smaller than that described by Ju et al. [22], other morphological characteristics of strain KNUE23N245 adhered to the original description. Notably, H. vinosopulvinatum was recently reported to produce purine compounds with antifungal activity [23].
Morphological characteristics of strain KNUE23P495: After 7 d of incubation at 25℃ in the dark, the colonies grown on MEA measured 44.8–48.5 mm in diameter and exhibited a circular shape. The front side of the colony appeared light gray, flat, and entirely covered with coarse cottony mycelia. The reverse side of the colony was pale yellow at the center and became light gray toward the edges. The colony margins had a cottony appearance (Fig. 4A). On PDA, the colonies reached diameters of 67.7–71.6 mm and also exhibited a circular shape. The front side of the colony was light gray, flat, and densely covered with coarse cottony mycelia. The reverse side showed a color gradient from vivid yellow at the center to light yellow toward the periphery with yellowish-white cottony margins (Fig. 4B). The conidia were globose, polygonal, or ampulliform with thick smooth surfaces. Numerous gutules are observed within the conidia. Initially, the hyaline phase turned dark brown upon maturation. The size of the conidia was (17.03–) 21.13 (–26.53) × (8.13–) 16.13 (–20.11) µm (n = 20) (Figs. 4C and D).
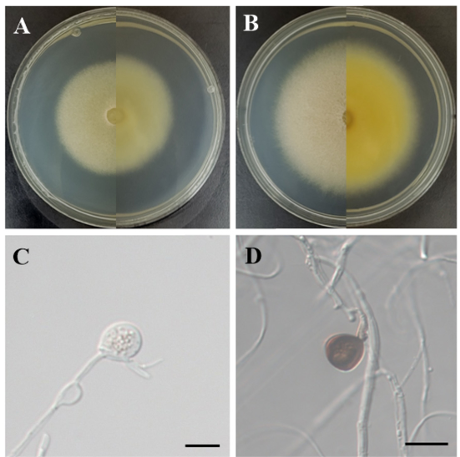
Fig. 4. Morphology of Apiospora sasae KNUE 23P495. (A, B) Colonies after 7 d of growth at 25℃ on malt extract agar (MEA; A) and potato dextrose agar (PDA; B). The left side of each image shows the front view, and the right side shows the reverse view. (C) Immature conidia (scale bar = 20 µm), (D) mature conidia (scale bar = 20 µm).
Specimen examined: Hwasun-gun, Jeollanam-do, Korea, 35°9’17.899″N 127°10’8.353″E, July 14, 2023, A. sasae, isolated from twigs of P. nigra var. henonis strain KNUE23P495 (NIBRFGC000510707), GenBank No. PV384448 (ITS), PV384449 (LSU), PV460718 (TUB), and PV460719 (TEF).
Phylogenetic analysis: The ITS sequence of KNUE showed 99.5% similarity to A. sasae CPC38165 (NR173004). The LSU sequence showed 99.9% similarity with CPC38465 (MW883797), the TUB sequence showed 98.1% similarity with CPC38165 (MW890120), and the TEF sequence exhibited 97.1% similarity with CPC38165(MW890104). In the ML phylogenetic tree constructed using the combined ITS, LSU, TUB, and TEF sequences, KNUE23P495 formed a clade with A. sasae strain CPC38165 (Fig. 5).
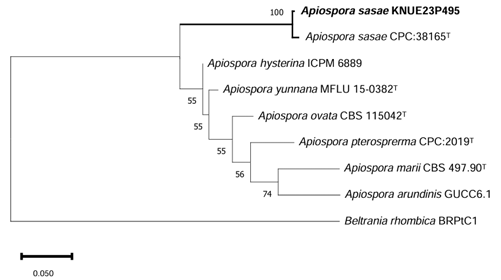
Fig. 5. Maximum-likelihood (ML) phylogenetic tree of Apiospora sasae KNUE 23P495. The tree was constructed based on concatenated sequences of internal transcribed spacer (ITS), large subunit rDNA (LSU), β-tubulin (TUB), and translation elongation factor 1-α (TEF1-α). Apiospora marii was used as an outgroup. The numbers on the nodes indicate bootstrap values of >50% (1,000 replicates).
Notes: A. sasae was first collected in 2019 from Sasa veitchii (Poaceae) in the Netherlands by van der Linde and reported in 2021 [15]. A. sasae is known to be related to, but morphologically distinct from Apiospora yunnana and Apiospora esporlensis [15]. Although phylogenetically close to one another, strain KNUE23P495 is morphologically distinguishable from A. yunnana, which has smaller conidia (10–16 µm) [24], by its significantly greater conidial width up to 6 µm wider-consistent with the original description of A. sasae [15]. A. esporlensis has smaller conidia (8–13µm) [25] than KNUE23P495, which exhibits a conidial width up to 13 µm greater consistent with the description provided by Crous et al. [15].
In the present study, we identified two endophytic fungal species, H. vinosopulvinatum and A. sasae, which have not been previously recorded in Korea. These species were isolated from T. nucifera and P. nigra var. henonis, respectively. Their identification was based on their morphological characteristics and molecular phylogenetic analysis using ITS, LSU, TUB, and TEF sequences.. Endophytic fungi exhibit a wide range of biological activities through interactions with their host plants, and further investigation is required to understand their ecological roles and potential functional applications. The results of this study provide fundamental data on fungal diversity in temperate forest ecosystems in Korea and support future studies on the ecological and biotechnological potential of endophytic fungi.
The authors declare no competing interests.
This work was supported by a grant from the National Institute of Biological Resources (NIBR NIBR202502103), funded by the Ministry of Environment (MOE) of the Republic of Korea.
1. Ferrari R, Gautier V, Silar P. Lignin degradation by ascomycetes. In: Morel-Rouhier M, Sormani R, editors. Advances in botanical research. Cambridge, MA: Academic Press; 2021. p. 77-113. [DOI]
2. Smith GJD, Liew ECY, Hyde KD. The Xylariales: A monophyletic order containing 7 families. Fungal Divers 2003;13:185-218.
3. Wendt L, Sir EB, Kuhnert E, Heitkämper S, Lambert C, Hladki AI, Romero AI, Luangsaard JJ, Srikitikulchai P, Peršoh D, et al. Resurrection and emendation of the Hypoxylaceae, recognised from a multigene phylogeny of the Xylariales. Mycol Progress 2018;17:115-54. [DOI]
4. Li QR, Habib K, Long SH, Wu YP, Zhang X, Hu HM, Wu QZ, Liu LL, Zhou SX, Shen XC, et al. Unveiling fungal diversity in China: New species and records within the Xylariaceae. Mycosphere 2024;15:275-364. [DOI]
5. Maharachchikumbura SSN, Hyde KD, Jones EBG, McKenzie EHC, Huang SK, Abdel-Wahab MA, Daranagama DA, Dayarathne M, D’souza MJ, Goonasekara ID, et al. Towards a natural classification and backbone tree for Sordariomycetes. Fungal Divers 2015;72:199-301. [DOI]
6. Ju YM, Rogers JD. A revision of the genus Hypoxylon. St. Paul, MN: American Phytopathological Society; 1996.
7. Stadler M, Kuhnert E, Peršoh D, Fournier J. The Xylariales as model example for a unified nomenclature following the “One Fungus-One Name” (1F1N) concept. Mycology 2013;4:521. [DOI]
8. Daranagama DA, Hyde KD, Sir EB, Thambugala KM, Tian Q, Samarakoon MC, McKenzie EHC, Jayasiri SC, Tibpromma S, Bhat JD, et al. Towards a natural classification and backbone tree for Xylariales based on multigene phylogenetic analyses. Fungal Divers 2018;88:1-165. [DOI]
9. Rodriguez RJ, White JFJ, Arnold AE, Redman RS. Fungal endophytes: Diversity and functional roles. New Phytol 2009;182:314-30. [DOI]
10. Maharachchikumbura SSN, Hyde KD, Jones EBG, McKenzie EHC, Bhat JD, Dayarathne MC, Huang SK, Norphanphoun C, Senanayake IC, Perera RH, et al. Families of Sordariomycetes. Fungal Divers 2016;79:1-317. [DOI]
11. Pintos Á, Alvarado P. Phylogenetic delimitation of Apiospora and Arthrinium. Fungal Syst Evol 2021;7:197-221. [DOI]
12. Eo JK, Choi JW, Eom AH. Diversity, distribution, and host plant of endophytic fungi: A focus on Korea. Mycobiology 2022;50:399-407. [DOI]
13. Choi JW, Lee JM, Park SY, Eom AH. Acrodontium burrowsianum and Pestalotiopsis humicola: Two previously unrecorded fungal species isolated from conifer leaves in Korea. Kor J Mycol 2022;50:311-8.
14. Dos Santos GD, Gomes RR, Gonçalves R, Fornari G, Maia BHLNS, Schmidt-Dannert C, Gaascht F, Glienke C, Schneider GX, Colombo IR, et al. Molecular identification and antimicrobial activity of foliar endophytic fungi on the Brazilian pepper tree (Schinus terebinthifolius) reveal new species of Diaporthe. Curr Microbiol 2021;78:3218-29. [DOI]
15. Crous PW, Hernández-Restrepo M, Schumacher RK, Cowan DA, Maggs-Kölling G, Marais E, Wingfield MJ, Yilmaz N, Adan OCG, Akulov A, et al. New and interesting fungi. 4. Fungal Syst Evol 2021;7:255-343. [DOI]
16. Gardes M, Bruns TD. ITS primers with enhanced specificity for basidiomycetes-application to the identification of mycorrhizae and rusts. Mol Ecol 1993;2:113-8. [DOI]
17. White TJ, Bruns TD, Lee SB, Taylor JW. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, editors. PCR protocols: A guide to methods and applications. New York: Academic Press; 1990. p. 315-22. [DOI]
18. Vilgalys R, Hester M. Rapid genetic identification and mapping of enzymatically amplified ribosomal DNA from several Cryptococcus species. J Bacteriol 1990;172:4238-46. [DOI]
19. O’Donnell K, Cigelnik E. Two divergent intragenomic rDNA ITS2 types within a monophyletic lineage of the fungus Fusarium are nonorthologous. Mol Phylogenet Evol 1997;7:103-16. [DOI]
20. Rehner S. Primers for elongation factor 1-a (EF1-a) [Internet]. Beltsville, MD: Insect Biocontrol Laboratory; 2001 [cited 2025 Apr 30]. Available from: https://web.archive.org/ web/20060518132929/http://ocid.NACSE.ORG/research/deephyphae/EF1primer.pdf.
21. Tamura K, Stecher G, Kumar S. MEGA11: Molecular evolutionary genetics analysis version 11. Mol Biol Evol 2021;38:3022-7. [DOI]
22. Ju YM, Rogers JD, Hsieh HM. New Hypoxylon species and notes on some names associated with or related to Hypoxylon. Mycologia 2004;96:154-61. [DOI]
23. Lv J, Zhou H, Dong L, Wang H, Yang L, Yu H, Wu P, Zhou L, Yang Q, Liang Y, et al. Three new furanones from endophytic fungus Hypoxylon vinosopulvinatum DYR-1-7 from Cinnamomum cassia with their antifungal activity. Nat Prod Res 2023;38:3165-72. [DOI]
24. Dai DQ, Phookamsak R, Wijayawardene NN, Li WJ, Bhat DJ, Xu JC, Taylor JE, Hyde KD, Chukeatirote E. Bambusicolous fungi. Fungal Divers 2017;82:1-105. [DOI]
25. Pintos Á, Alvarado P, Planas J, Jarling R. Six new species of Arthrinium from Europe and notes about A. caricicola and other species found in Carex spp. hosts. MycoKeys 2019;49:1548. [DOI]