Genus Lambertella Höhn., the largest genus in the family Rutstroemiaceae, has 63 species reported around the world [1]. In Korea, the Lambertella sp. was discovered in the leaves of Pinus koraiensis Siebold et Zucc. but has not been described at the species level [2]. The Genus Lambertella was first reported by Höhnel in 1918; Dumont's specimen studies established a clear definition of this genus [3,4]. Lambertella has similar morphological characteristics to other genera Lanzia or Polculum of the same family; it can be distinguished by its substratal stroma structure and spore pigmentation [5].
Abies koreana E. H. Wilson has been selected by the Ministry of Environment (National Institute of Ecology) as a vulnerable species to climate change; it inhabits alpine regions of 1,000 m or higher in Korea but its population is rapidly declining due to rising temperatures and decreasing soil moisture [6,7]. In this study, Lambertella corni-maris was discovered while studying the biodiversity of the endophyte fungi in alpine conifers, and it was confirmed as an unrecorded endophytic fungus in Korea based on its morphological and molecular characteristics and so is reported here.
The needle leaves of Korean fir used in the study were collected from Mt. Halla in the Jeju Special Self-governing Province in 2018. The collected samples were placed in a sealed zipper bag and stored at a low temperature until fungi were separated within 24 h. All needle leaves were washed in running water and cut into approximately 1 cm pieces. The surface sterilization conditions were 96% ethyl alcohol, sodium hypochlorite, and 96% ethyl alcohol in that order. Each treatment was performed for 1 min, 3 min, and 30 s, and finally, samples were washed twice using sterile water. The needles were placed on potato dextrose agar (PDA; MBcell, Seoul, Korea) and then cultured at 25℃ in the dark to isolate endophytic fungi. The isolated mycelium was cultured using PDA, and maltose extract agar (MEA; MBcell, Seoul, Korea) media were used for the pure culture of endophytic fungi to observe the colony’s morphological characteristics [8]. Microstructures in the colony, including spores, were observed after staining with lactophenol cotton blue under a light microscope (DM2500, Leica Microsystems, Wetzlar, Germany). The unrecorded endophytic fungus used for observation was deposited with the Korean Collection for Type Cultures (KCTC).
Genomic DNA was extracted from the fungus using a plant tissue genomic DNA extraction kit (Xi'an Tianlong Science & Technology, Shaanxi, Taiwan) following the manufacturer’s instructions. Polymerase chain reaction (PCR) was performed using primers ITS1F and ITS4, which can selectively amplify from the internal transcribed spacer region (ITS) 1, 2, including 5.8S ribosomal DNA and 28S ribosomal DNA [9,10]. The conditions during PCR were as follows: Pre-denaturing for 5 min at 94℃ with one cycle, denaturing for 30 s at 94℃, annealing for 30 s at 50℃, extending for 1 min at 72℃ across 30 cycles, and then finally stabilizing for 10 min at 72℃ for one cycle. The PCR product was confirmed by electrophoresis using 1.5% agarose gel. DNA sequencing was supplied to Cosmogenetech Inc. (Daejeon, Korea), and the analyzed DNA sequence was then identified based on similarity with the National Center for Biotechnology Information (NCBI, http://www.ncbi.nlm.nih.gov/) using the basic local alignment search tool (BLAST). A neighbor-joining tree was generated by MEGA10.0.5 based on the Kimura-2 parameter distance model with the 1,000-times bootstrap method [11].
Lambertella corni-maris Höhn., Sber. Akad. Wiss. Wien, Math.-naturw. Kl., Abt. 1 127 (4-5): 375 (1918) [MB#166748]
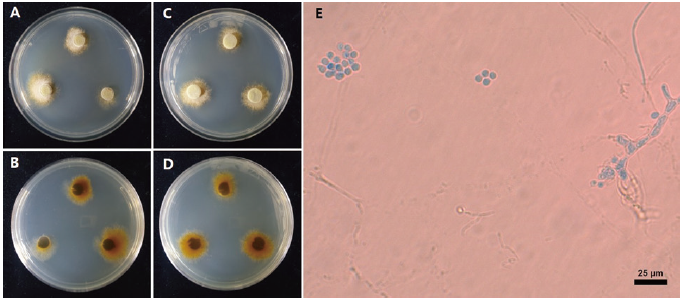
The colony diameter after 14 d was 14.03-19.39 mm on MEA and 10.35-19.24 mm on PDA. The mycelium was rather dense. The surface color was light gambogeish gray (Munsell color notation: 10Y 9/2) in MEA and grayish gamboge (5Y 7/4) in PDA, it has a texture similar to downy or cotton with an irregular margin and no exudate. The reverse was moderate gamboge (Munsell color notation: 10YR 4/8) in MEA and moderate gamboge (7.5 YR 5/10) in PDA [12]. Conidia measured about (5.20-)5.63(-6.41) µm (n=20) were circular with a hyaline well with lactophenol cotton blue and showed no septa inside (Fig. 1; Table 1).
|
Table 1. Morphological characteristics of Lambertella corni-maris NIE7191 isolated from the needle leaves of Abies korenana. 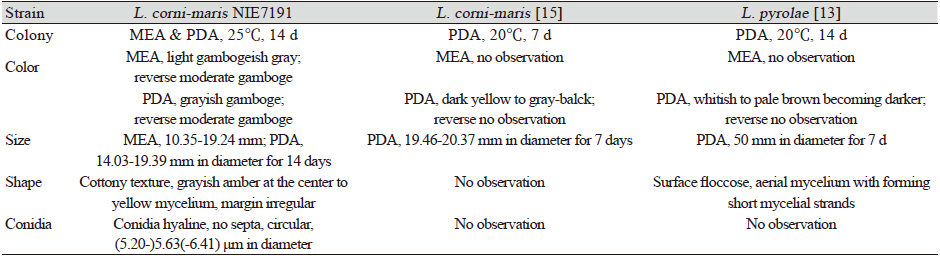
|
|
MEA: Maltose extract agar; PDA: Potato dextrose agar |
Specimen examined: Mt. Halla, Jeju Special Self-governing Province, Korea, 33°22'49.87"N, 126°35'37.86"E, 2018.9.12., isolated from the leaves of Abies koreana, strain NIE7191, KCTC no. 56756, GenBank no. MW563952 (ITS) and MW786688 (LSU).
Note: In the genus Lambertella, a total of 63 species have been reported so far [1,3,4]. Among reports on Lambertella spp., there have been no reports on the specificity of the host plant. L. corni-maris was first isolated and reported to be on Cornus mas L. living in Europe [4]. In this study, it is believed that the distribution range of L. corni-maris is wide, from broad-leafed trees to coniferous trees such as A. koreana [2,13]. The nucleotide sequence of the ITS region, including 5.8S ribosomal DNA and 28S ribosomal DNA, were analyzed and used for the molecular identification of L. corni-maris. Each sequences were shown 100% and 99.9% homology with ITS region of L. corni-maris (AB705239.1) and 28S ribosomal DNA of L. corni-maris (KC964858.1) (Fig. 2). Lambertella spp. are found in Australia, China, Europe, India, Japan, New Zealand, and North America, and Andromeda sp., Aucuba sp., Camellia sp., Cornus sp., Fatsia sp., Larix sp., Mallotus sp., Pinus sp., Pyrola sp. and Torreya sp. have been reported as host plants to date [2,13,14]. Through this study, L. corni-maris was first isolated from A. koreana, and there have been no previous studies on Lambertella sp. in Korea. Most previous studies on the morphology of Lambertella spp. are based on the apothecium or ascospore directly observed in the fruit or leaf of a host plant, so morphological observations cultured in medium are insufficient. In particular, the coloration of spores occurring in ascus has not been confirmed in conidia. Thus, in the future, further taxonomic studies of Lambertella spp. to describe the morphological characteristics and microstructure on a culture medium should be performed.
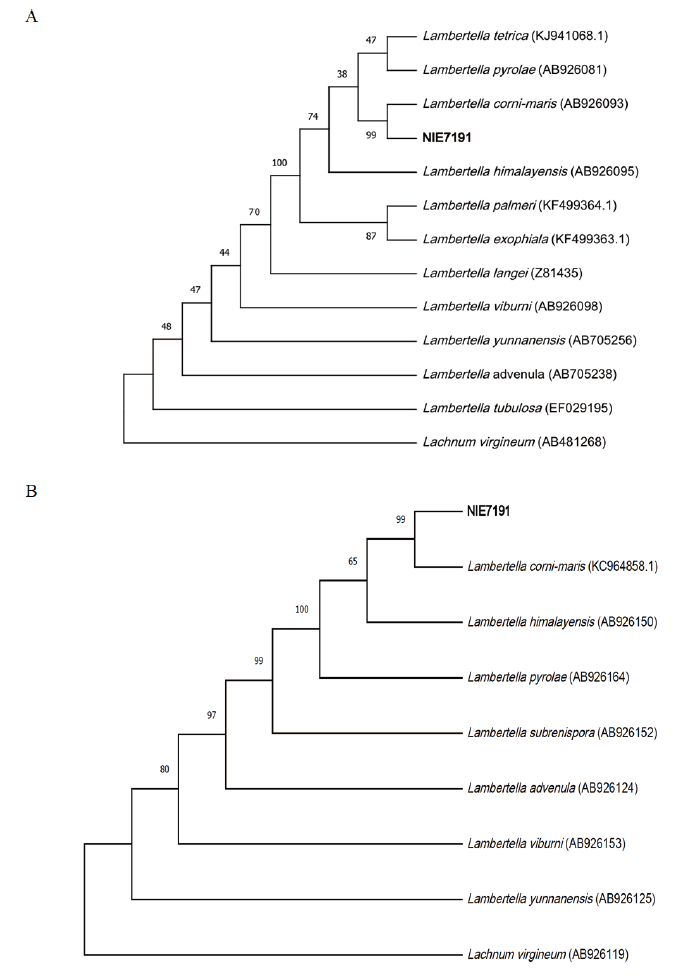
Fig. 2. Phylogenic tree of Lambertella corni-maris NIE7191 isolated from Abies koreana. (A) Internal transcribed spacer region (ITS) region, including the 5.8S ribosomal DNA region, and (B) 28S ribosomal DNA were used for sequence analysis to confirm the topological appropriation of the fungal isolates. Lachnum virgineum was used as an out-group, and bootstrap values are shown at the branches (1,000 replicates).