Introduction
Luttrell [1] proposed the family Microascaceae to accommodate the genus Microascus Zukal, and its emended description was published by Malloch in 1970 [2]. Currently, the family consists of a morphologically heterogeneous group of saprobic, plant pathogenic, and opportunistic human pathogenic fungi [3-6]. The genus Acaulium was established as a sexual morph within the family Microascaceae, and its type species is A. albonigrescens Sopp [7]. Previously considered a synonym of Scopulariopsis, the genus Acaulium was recently re-instated by Sandoval-Denis et al. [8] based on the descriptions of three species, i.e., A. acremonium (Delacr.) Sandoval-Denis, Guarro & Gene, A. albonigrescens Sopp, Skr. Vidensk.-Selsk, and A. caviariforme (Malloch & Hubart) Sandoval-Denis, Guarro & Gene. At the time of this writing, the genus Acaulium consists of at least seven species. Acaulium album was formerly known as Doratomyces putredinis, but it has been transferred to the genus Acaulium and redescribed by Woudenberg based on morphological, physiological, and molecular phylogenetic analyses [9]. Acaulium pannemaniae became a member of the genus in 2018 [10] and, recently, two new members, namely, A. peruvianum and A. retardatum, were proposed by Su et al. [11]. Morphologically, members of the genus Acaulium undergo annellidic conidiogenesis, have guttulate conidia, and their mycelia form abundant hyphal fascicles.
The aim of this study is to investigate the cultural and morphological diversity of native fungal species in a special environment in Korea and analyze their molecular phylogenetic relationships. Herein, we describe a novel, insect-associated fungal species belonging to the genus Acaulium, which was isolated from the bean bug Riptortus clavatus.
Materials and methods
Sample collection and fungal isolation
A specimen of the insect (Riptortus clavatus) was collected from Gyeongbuk, Gunwi, Korea (36°06'50.1''N, 128°38'24.4''E) and then transferred to the laboratory and stored at 4℃ until use. The ground insect body was mixed with double-distilled water, and the suspension was serially diluted. We spread 200 μL of each dilution onto potato dextrose agar (PDA; Difco, Detroit, MI, USA) plates and incubated the plates for 2–3 days at 25℃. Single isolated colonies were transferred to fresh PDA plates and incubated for another 4–5 days at 25℃. One colony, designated KNUF-20-INY03, was selected for further molecular analyses based on its distinct cultural characteristics. Stocks of strain KNUF-20-INY03 were maintained in 20% glycerol at -80℃ for further study.
Morphological characterization
Cultural and morphological characteristics of strain KNUF-20-INY03 were recorded after incubating cultures on PDA and oatmeal agar (OA; Difco, Detroit, Ml, USA) for 14 days at 25℃. During this time, fungal growth was measured and colony characteristics such as color, shape, and size were recorded. Morphological characteristics of the strain were observed using light microscopy (BX-50, Olympus, Tokyo, Japan).
Genomic DNA extraction, PCR amplification, and sequencing
Fungal mycelia were grown on PDA plates for 4–5 days at 25℃ and then scraped off with a sterile blade. Genomic DNA was extracted from the mycelia using a HiGene Genomic DNA Prep Kit (Biofact, Daejeon, Korea) following the manufacturer's instructions. PCR targeted the internal transcribed spacer regions (ITS) and partial sequences of the large subunit of the nuclear ribosomal RNA (LSU), translation elongation factor 1-alpha (TEF1-α), and β-tubulin (β-TUB) genes. The ITS regions were amplified using the primers ITS1F/ITS4 [12,13]; the partial LSU gene was amplified using the primer pair LROR/LR5 [14,15]; and the partial β-TUB and TEF1-α genes were amplified using the primer pairs Bt2a/Bt2b [16,17] and EF1-983F/EF1-2218R [18], respectively. The PCR yields were verified on 1% agarose gels and stained with ethidium bromide. The amplified PCR products were purified using the EXOSAP-IT kit (Thermo Fisher Scientific, Waltham, MA, USA) following the manufacturer’s instructions, and the purified DNA was sequenced by Macrogen Co., Ltd. (Daejeon, Korea). Sequence data were analyzed using SeqMan Lasergene software (DNAStar Inc., Madison, Wisconsin, USA). The ITS regions, β-TUB, TEF1-α, and LSU gene sequences of KNUF-20-INY03 were deposited in GenBank under accession numbers LC645069, LC645071, LC645072, and LC645070, respectively.
Molecular phylogenetic analysis
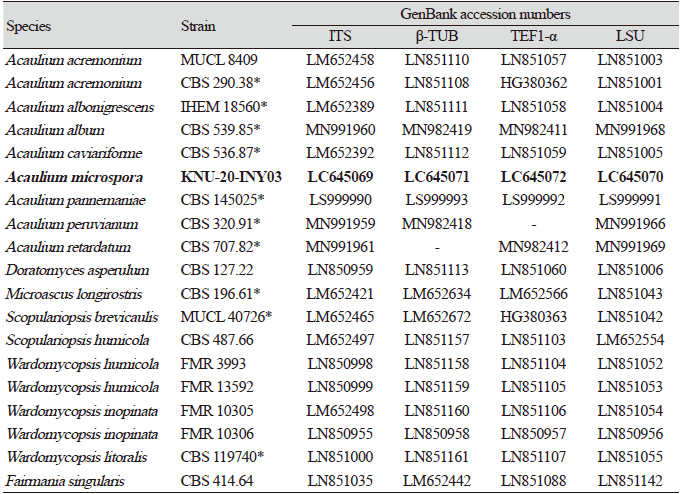
Sequences of species related to genus Acaulium were retrieved from the National Center for Biotechnology Information (NCBI) (Table 1). The multiple sequences were aligned using Clustal X 2.0 [19]. Concatenated nucleotide sequences consisting of the ITS regions and the partial sequences of β-TUB, TEF1-α, and LSU genes were phylogenetically analyzed using neighbor-joining (NJ) [20], maximum-likelihood (ML) [21], and maximum-parsimony (MP) [22] methods, as implemented in MEGA7 [23]. The NJ analysis used Kimura two-parameter distances [24]; the ML analysis used the nearest neighbor interchange heuristic search method and Kimura’s two-parameter model; and the MP analysis used the subtree pruning and re-grafting heuristic search method with gaps removed from the analysis. The robustness of the NJ, ML, and MP trees are reflected by bootstrap values, which are based on 1,000 replicates.
Results
Taxonomy
Acaulium microspora S.Y. Lee, L.N. Ten & H.Y. Jung sp. nov. (Fig. 1)
MycoBank: MB 841133
Etymology: The name “microspora” refers to the small conidia produced by this species.
Typus: Gyeongbuk, Gunwi, Korea (36°06'50.1''N, 128°38'24.4''E), isolated from insect (Riptortus clavatus). The metabolically inactive stock culture (NIBRFGC000508600) has been deposited in the National Institute of Biological Resources (NIBR).
Habitat: The fungus is associated with the insect (Riptortus clavatus).
Known distribution: Korea
Description: The hyphae are smooth-walled, thin and hyaline. Conidiophores are both branched and unbranched, hyaline, and smooth-walled. The conidiogenous cells are annellidic, smooth-walled, elliptical at the tip and smooth and rounded. The conidia are hyaline, visible, obovoid to oval, smooth-walled, and forming chains measuring 5.4–8.9 × 3.6–5.6 µm (av. 7.2 × 4.6 µm, n=100); their length-to-width (L/W) ratio is 1.6 (Fig. 1). Sexual morph was not observed. Colonies on PDA grew to a diameter of 12.5 mm after culturing at 25℃ for 14 days. Growth is slow and rising, colonies are initially white to ivory, bright on the outside, with the appearance of white powder. Colonies eventually spread wide and form branches, their reverse side is ivory colored. Colonies cultured on OA reach 16 mm after 14 days of growth at 25℃. They are round, white to ivory, with a flat powder appearance; their outer round forms a band, and their reverse round is brown (Fig. 1).
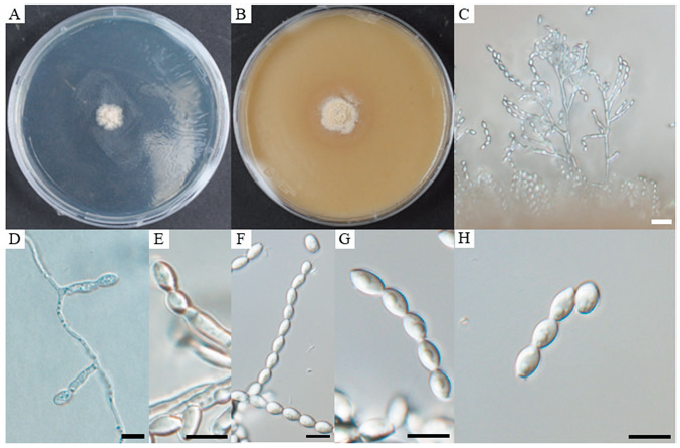
Fig. 1. Cultural and morphological characteristics of Acaulium microspora. A, colony on potato dextrose agar (PDA) after 14 days of growth at 25℃; B, colony on oatmeal agar (OA) after 14 days of growth at 25℃; C, D, and E, conidiophores and conidiogenous cells; F, G, chain of conidia; H, conidia. Scale bars: C = 20 μm; D–H = 10 μm.
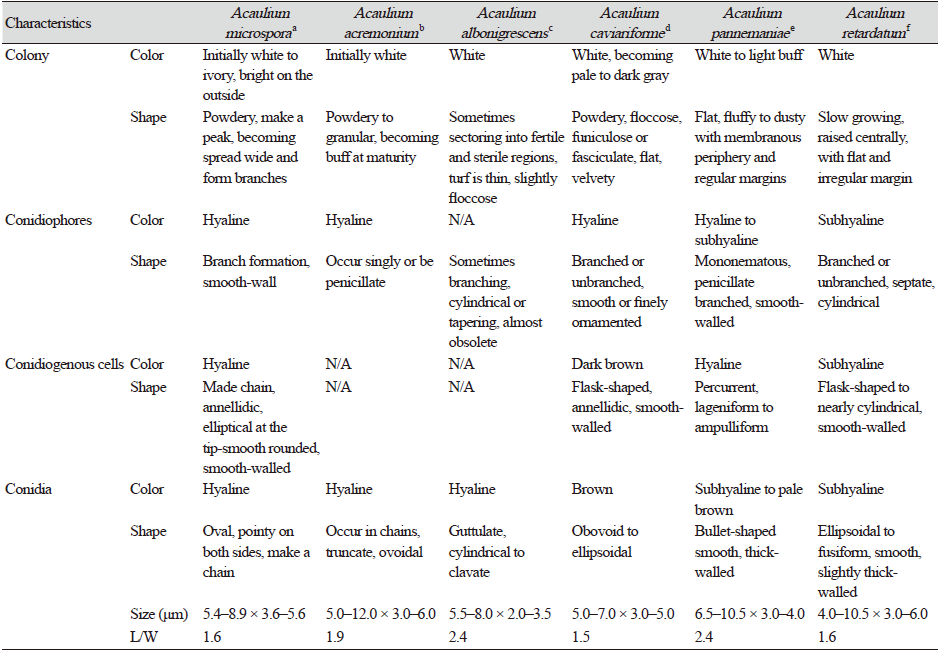
Note: Morphologically, strain KNUF-20-INY03 resembles the phylogenetically closely related Acaulium acremonium, A. albonigrescens, A. caviariformis, A. pannemaniae, and A. retardatum; however, the strain is readily distinguishable from these species based on the shape, size, and color of conidia and conidiogenous cells (Table 2). The average length of conidia of strain KNUF-20-INY03 (5.4–8.9 × 3.6–5.6 μm; av. 7.2 × 4.6 μm, L/W=1.6) is much shorter than that of A. acremonium (5–12 × 3–6 μm, av. 8.5 × 4.5 μm, L/W=1.9) and A. pannemaniae (6.5–10.5 × 3–4 μm, av. 8.5 × 3.5 μm, L/W=2.4). The average width of the conidia of the isolate (4.6 μm) is significantly larger than that of A. albonigrescens (5.5–8 × 2–3.5 μm, av. 6.8 × 2.8 μm, L/W=2.4) and A. pannemaniae (3.5 μm). The conidial L/W values clearly differentiate strain KNUF-20-INY03 (1.6) from A. acremonium (1.9), A. albonigrescens (2.4), and A. pannemaniae (2.4). The conidia of the isolate are slightly larger than that of A. caviariformis (5–7 × 3–5 μm, av. 6 × 4 μm, L/W=1.5), and its conidia are hyaline while that of A. caviariformis are brown. The conidial length range of A. microspora (5.4–8.9 μm) is less than that of A. retardatum (4–10.5 × 3–6 μm, av. 7.2 × 4.5 μm, L/W=1.6) and the novel isolate produces elliptical conidiogenous cells while A. retardatum forms flask-shaped conidiogenous cells. Furthermore, unlike KNUF-20-INY03, A. albonigrescens, A. caviariformis, and A. retardatum have a sexual morph [8].
Phylogenetic analysis
Amplicons of the ITS, β-TUB, TEF1-α, and LSU loci of strain KNU-20-INY03 were 623, 544, 907, and 782 bp long, respectively. A BLAST search of the NCBI database reveals that the LSU gene sequence of KNU-20-INY03 is closely related to those of Acaulium acremonium DTO 401-F9 (100% similarity), A. albonigrescens CBS 109.69 (99.2%), A. album CBS 539.85 (98.9%), A. retardatum CBS 707.82 (98.9%), and A. caviariformis CBS 536.87 (formerly Microascus caviariformis) (98.9%). The ITS sequence of strain KNU-20-INY03 is 100%, 98.6%, and 96.4% identical with those of A. acremonium DTO 401-F9, A. album CBS 539.85, and A. peruviarum CBS 320.91, respectively. The partial TEF1-α gene sequence of the isolate is 95.9%, 94.9%, and 93.6% similar to those of A. acremonium MUCL9028 (former Scopulariopsis acremonium), A. album CBS 539.85, and A. caviariformis CBS 536.87, respectively. The β-TUB gene sequence of strain KNU-20-INY03 is 99.3% similar to that of A. acremonium DTO 401-F9, while it is only 90.0% similar to those of A. albonigrescens IHEM 18560 (former M, albonigrescens) and A. caviariformis RB002.1 (former Microascus caviariformis). Overall, these results clearly indicate that none of the gene sequences allowed for a precise identification of the novel fungal strain at the species level. Therefore, we performed a multilocus sequence analysis using concatenated sequences of the ITS regions, β-TUB, TEF1-α, and LSU genes of KNU-20-INY03 (Table 1). Combining these four molecular markers has been highly effective in resolving the species within the genus Acaulium [11]. The ML phylogenetic tree based on the concatenated sequences clearly shows that KNU-20-INY03 is distinct from other Acaulium species (Fig. 2). Moreover, the ML tree shares corresponding nodes with the NJ and MP trees, as indicated by the filled circles in Fig. 2. These results indicate that the strain is a single, novel, phylogenetically distinct Acaulium species.
Discussion
Malloch emended the description of the family Microascaceae [2] based on the characteristics of five genera, although according to the Mycobank database (), this family consists of more than forty genera. Molecular studies performed by Sandoval-Denis et al. [8] demonstrated that several genera within the Microascaceae are closely related and are difficult to differentiate based only on morphological characteristics. Over the last decade, several studies have sought to determine the best molecular markers for resolving the phylogenetic relationships between members of Microascaceae. A study of clinical isolates of Scopulariopsis showed the inadequacy of the LSU sequence at differentiating between species [25], although the combination of LSU, β-TUB, and TEF1-α gene sequences successfully identified several Scopulariopsis species isolated from cheese [26]. Recently, the branches of the Microascus, Scopulariopsis, and Pithoascus genera within the family Microascaceae were revised based on an updated phylogenetic analysis using the concatenated sequences of the ITS regions and the LSU, TEF1-α, and β-TUB genes [5]. The analysis excluded several taxa that were previously associated with these genera, which formed a new lineage within the Microascaceae. In particular, the analysis proposed that the genus Acaulium, previously considered synonymous with Scopulariopsis, constitutes a distinct genus, and thus the old genus Acaulium was revalidated to include three species, namely, A. acremonium, A. albonigrescens, and A. caviariformis. Meanwhile, Woudenberg et al. [9] used a combination of LSU and ITS sequences to show that Cephalotrichum album does not belong to Cephalotrichum and should be reclassified as a synnematous species of the genus Acaulium. The same multi-gene approach has been applied to revise the taxonomy of the genera Kernia and Acaulium [11]; these genera were clearly separated by phylogenetic analysis based on four combined loci (LSU, ITS, TEF1-α, and β-TUB). This analysis resulted in the addition of A. peruvianum and A. retardatum within the genus Acaulium. Besides that, based on morphological features and phylogenetic analyses of LSU, A. pannemaniae was introduced in this genus by Crous et al. [10]. Thus, this genus now comprises seven species.
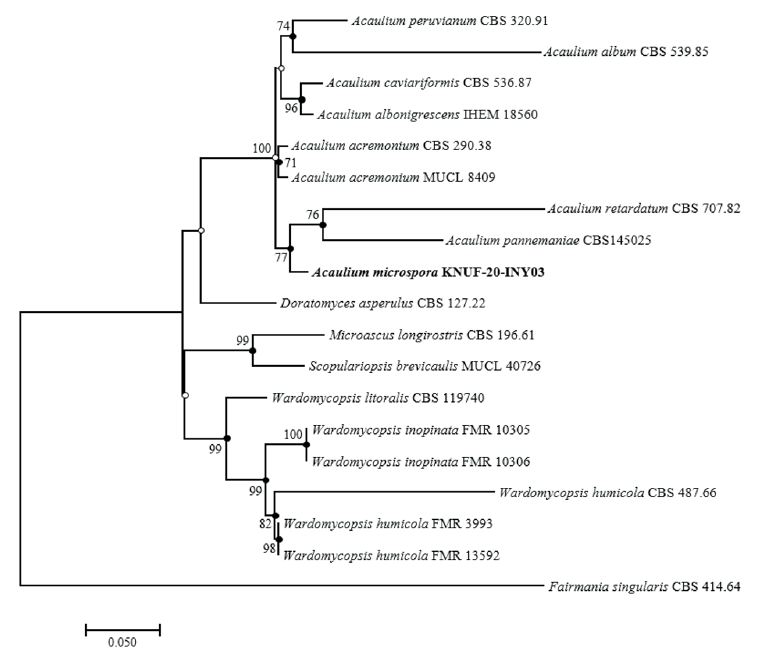
Fig. 2. Maximum-likelihood phylogenetic tree of KNUF-20-INY03 based on the combined sequences internal transcribed spacer (ITS) regions, β-tubulin (β-TUB), translation elongation factor 1-alpha (TEF1-α) and large subunit of the nuclear ribosomal RNA (LSU), showing the phylogenetic position of novel strain KNUF-20-INY03 among Acaulium spp. and other closely related fungal species. Bootstrap values (based on 1,000 replications) greater than 70% are shown at branch points. Filled circles indicate that the corresponding nodes were also recovered in trees generated using the neighborjoining and maximum-parsimony algorithms. Open circles indicate that the corresponding nodes were also recovered in the tree generated using the neighbor-joining or maximum-parsimony algorithm. The isolated strain is shown in bold. Fairmania singularis CBS 414.64 was used as an outgroup. Bar, 0.05 substitutions per nucleotide position.
In this study, we characterized strain KNU-20-INY03, which was isolated from the body of Riptortus clavatus collected in Korea. BLAST searches of sequences of the strain’s ITS regions and its LSU, TEF1-α, and β-TUB genes revealed its affiliation to the genus Acaulium. Depending on the molecular marker used, similarity analysis indicates that strain KNU-20-INY03 is most closely related to A. acremonium, A. albonigrescens, A. caviariformis, A. retardatum, or A. album. Clearly, multi-gene phylogenetic analysis is required to properly identify the novel strain at the species level. Using previously successful approaches [8,11] that achieved a fine level of phylogenetic resolution, we carried out multilocus sequence analysis of strain KNU-20-INY03 using concatenated sequences of its ITS regions, along with partial sequences of the TEF1-α, β-TUB, and LSU genes. The topologies of the resulting ML, NJ, and MP trees show that the isolate is distinct from the seven valid Acaulium species (Fig. 2). The closest neighbors of Acaulium microspora (KNU-20-INY03) are A. acremonium, A. albonigrescens, A. caviariformis, A. pannemaniae, and A. retardatum and these Acaulium species share certain typical morphological features of the genus [8]. A. microspora differs from the five above-mentioned Acaulium species by the shape, size, and color of its conidia and conidiogenous cells. The conidia of strain KNU-20-INY03 are shorter (5.4–8.9 × 3.6–5.6 μm) than those of A. acremonium (5.0–12.0 × 3.0–6.0 μm). The average length of the KNU-20-INY03 conidia is approximately the same as that of A. albonigrescens (5.5–8.0 μm); however, the conidial width of KNU-20-INY03 is significantly greater (3.6–5.6 μm vs 2.0–3.5 μm). In addition to the conidial dimensions, the L/W ratios can differentiate between Acaulium species, which is reflected by the L/W-based differentiation of strain KNU-20-INY03 (L/W=1.6) from A. acremonium (L/W=1.9), A. albonigrescens (L/W=2.4), and A. pannemaniae (L/W=2.4). Moreover, the hyaline conidia and elliptical conidiogenous cells of KNU-20-INY03 are readily distinguished from the brown conidia of A. caviariformis and flask-shaped conidiogenous cells of A. retardatum. Finally, in contrast to A. albonigrescens, A. caviariformis, and A. retardatum, KNU-20-INY03 does not have a sexual morph. Therefore, both morphological and phylogenetic analyses indicate that strain KNU-20-INY03 is a novel species, and we have designated Acaulium microspora in this study.
Acaulium species have been isolated from a variety of sources on different continents. The type species is A. albonigrescens, a well-known fungus that was originally isolated from soil, dung, and wood in Scandinavia, northern North America, and Japan [8]. A. album is broadly distributed in Europe and North America on heavily decayed organic material and various kinds of dung. A. pannemaniae and A. retardatum are soil-borne fungi with strains that have been isolated in Peru and Japan [11]. Among the Acaulium species, A. caviariforme appears to occupy a unique niche, having been isolated from meat in caves in Europe and North America [8], while only A. acremonium has been reported to cause skin and nail infections in humans [3] (although its identification in cases of clinical infection has not been confirmed by molecular methods [8]).
A. microspora was isolated from the bean bug (Riptortus clavatus), thus extending the distribution of members of this genus; it is the first insect-associated species of the genus Acaulium. Our results raise awareness regarding the distribution of Acaulium and highlight the need for further studies on the ecological and biological roles of this genus.
In conclusion, phylogenetic and morphological analyses show that strain KNU-20-INY03 is distinct from previously identified Acaulium species and should be considered a novel species within the genus. We propose the name Acaulium microspora sp. nov.