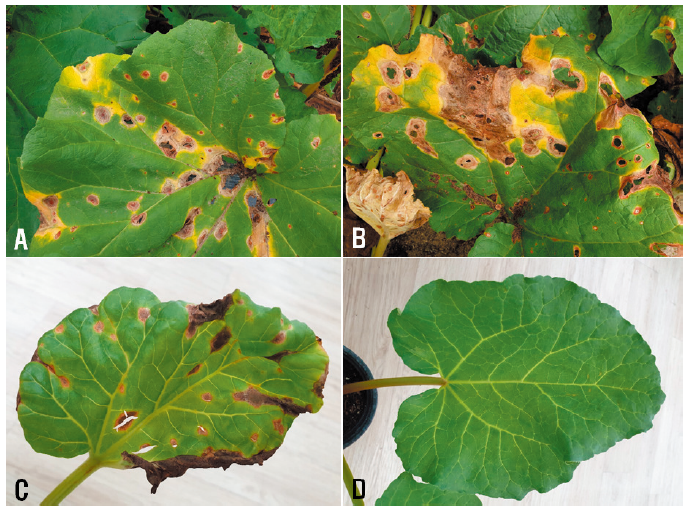
Rhubarb (Rheum rhabarbarum L., synonym: R. undulatum L.) belongs to the family Polygonaceae, and has been used as food and medicine [1]. The native range of the plant is southern Siberia to north and central China; it was introduced into several countries including Canada, USA, Czechoslovakia, and Korea [2]. The plant has been grown as a medicinal plant in Korea. Leaf spot symptoms in rhubarb were frequently observed in the plants growing in fields located in Cheolwon, Taebaek, and Inje in Gangwon Province, Korea, during disease surveys from 2019 to 2021. The symptoms initially appeared as circular red spots with a diameter of 1–5 mm on the leaves, and the peripheries of the spots displayed yellow halos (Fig. 1A). With the progression of the disease, the lesions expanded to more than 10 mm in diameter, and some lesions were perforated and torn (Fig. 1B). Later, the diseased leaves turned yellowish-brown and blighted. The spot lesions sometimes occurred on stems of the diseased plants. Leaves of 10 plants in each field were investigated in triplicate. The incidence of the diseased leaves in the plants ranged from 2% to 80% (Table 1).
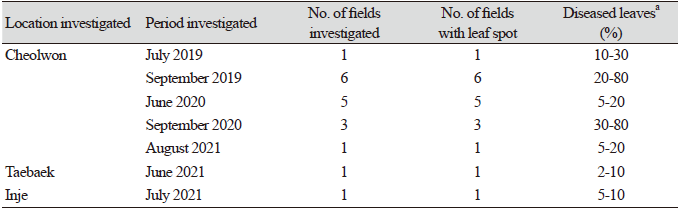
Table 1. Occurrence of leaf spot on rhubarb plants in fields located in Cheolwon, Inje, and Taebaek in Korea from 2019 to 2021.
|
|
aLeaves of 10 plants in each field were investigated in triplicae |
The diseased leaves of rhubarb were collected from the fields, investigated for the presence of the pathogens, and a fungal pathogen was isolated from them. Three to 5 mm-long lesion pieces were cut from the diseased leaves and were surface-sterilized using 1% sodium hypochlorite solution for one min. Then, the lesion pieces were plated on 2% water agar (WA; FUJIFILM Wako Chemicals, Chuo-Ku, Osaka, Japan). The fungal mycelia growing from the lesion pieces were transferred to potato dextrose agar (PDA; Difco, Sparks, MD, USA) slants after incubating the plates at 25℃ for 2-3 days. Morphological characteristics of the isolates cultured in PDA slants for 3‒4 weeks were examined using a compound microscope (Nikon Eclipse Ci-L, Japan). Most of the isolates were identified as Phoma sp. based on the morphological characteristics, as per descriptions given in a previous study [3]. The Phoma sp. isolates were transferred to oatmeal agar (OA; Sigma-Aldrich, St. Louis, MO, USA) plates and incubated at 22℃ for 2 weeks. Conidial suspension of each isolate was prepared from the OA plate cultures and streaked on WA plate using a sterile loop. After incubation of the WA plate for 24 hr at 22℃, germinated conidia were picked up under a dissecting microscope (Nikon SMZ 1780, Japan) and transferred to new WA plates. Nine single-spore isolates of Phoma sp. were obtained from the WA plate cultures and cultured in PDA slants. The isolates were used for identification and pathogenicity tests.
Three single-spore isolates of Phoma sp. were cultured on malt extract agar (MEA; Sigma-Aldrich, St. Louis, MO, USA), OA, and PDA at 22℃ for 14 days for investigation of their cultural and morphological characteristics using the methods described in previous studies [3,4]. Five culture plates of each isolate were used for each medium. Average diameters of 7-day-old cultures of the isolates grown on MEA, OA, and PDA at 22°C under darkness were 5.9 cm, 5.5 cm, and 5.7 cm, respectively. NaOH spot tests [4] of the isolates on MEA cultures presented negative reactions. Colony morphology of the isolates on the three media was investigated after an additional incubation of the cultures in alternating cycles of 13 hr NUV light and 11 hr darkness for 7 days. The colony on MEA showed white to light-brown concentric rings (Fig. 2A). The colony on OA showed olivaceous buff with gray olivaceous rings (Fig. 2B). The colony on PDA showed similar characteristics to that on MEA (Fig. 2C). Morphological features of 10 pycnidia and 30 conidia per isolate produced in 2-week-old OA cultures were examined using the compound microscope. Pycnidia were globose to subglobose, solitary or confluent, with non-papillate to 1‒2 papillate ostioles, olivaceous to olivaceous black (Fig. 2D), and measured 137‒340 μm in diameter. Conidia were ellipsoidal to cylindrical, with bipolar small guttules, mainly single-celled, rarely 1-septate (Fig. 2E), and measured 3.9‒11.4 μm × 1.6‒3.2 μm (av. 5.5 μm × 2.4 μm). All the isolates were identified as Phoma rhei (Ellis & Everh.) Aa & Boerema based on the cultural and morphological characteristics described in a previous study [4].
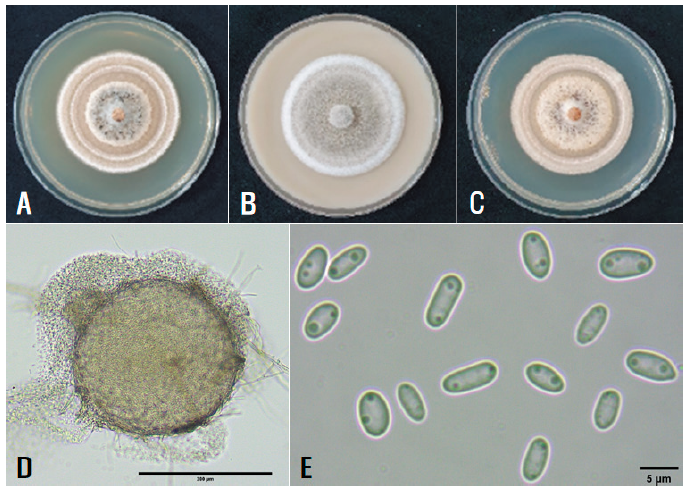
Fig. 2. Cultural and morphological features of Didymella rhei isolate from rhubarb. Colonies of the isolate grown on malt extract agar (A), oatmeal agar (B), and potato dextrose agar (C) at 22°C for 14 days. A pycnidium of the isolate produced in oatmeal agar (D) and conidia produced in the pycnidium (E).
P. rhei was reclassified as Didymella rhei (Ellis & Everh.) Q. Chen & L. Cai, the teleomorph of the fungus, based on the phylogenetic analyses [5]. To confirm the identification of P. rhei isolates based on the cultural and morphological characteristics, three target genes, partial large subunit nrDNA (28S, LSU), internal transcribed spacer regions 1 & 2 and intervening 5.8S nrDNA (ITS), and partial gene regions of β-tubulin (TUB2) regions were investigated. Genomic DNA of the isolates was extracted using molecular grade Chelex 100 resin (Bio-rad, USA) according to the protocol described by Simon et al. [6], with slight modifications. Following primers for PCR amplification of the genomic DNA were used with LR0R (5’-ACCCGCTGAACTTAAGC-3’) [7] and LR7 (5’-TACTACCACCAAGATCT-3’) [8] for LSU, V9G (5’-TTACGTCCCTGCCCTTTGTA-3’) [9] and ITS4 (5’-TCCTCCGCTTATTGATATGC-3’) for ITS, and TUB2Fd (5’-GTBCACCTYCARACCGGYCARTG-3’) and TUB4Rd (5’-CCRGAYTGRCCRAARACRAAGTTGTC-3’) [10] for TUB2. Conditions of PCR amplification for all the genes were followed as in a previous study [11]. All alignment works were conducted using Bioedit version 7.2.5 [12]. All PCR products were checked with 100 bp plus DNA ladder (Bioneer, Korea). Sequencing of genomic DNA was carried out based on the same primer sets at Bionics (Seoul, Korea). To construct a phylogenetic tree of the concatenated sequences with the three target genes, maximum likelihood (ML) method was utilized. This method included 1,000 bootstrap replicates and was conducted by MEGA version 7 [13] with a general time reversible model. The final concatenated alignment included 11 ingroup taxa with a total of 2,126 characters containing gaps (1,306 for LSU, 488 for ITS, and 332 for TUB2). Coniothyrium palmarum Corda (CBS 400.71) was used as the outgroup taxon. The reference sequence data were obtained from the NCBI GenBank database. The phylogenetic tree based on loci LSU, ITS, and TUB2 combined sequence data showed that all the isolates were clustered in a group with D. rhei CBS 109177 (Fig. 3). The sequences of the three loci genes from the isolates were 99‒100% identical to those of the reference strain D. rhei CBS 109177 of the GenBank database. The nucleotide sequences of LSU, ITS, and TUB2 genes obtained from the nine isolates were deposited in NCBI GenBank with accession numbers of OL721954-OL721962, OL744200-OL744208, and OL792697-OL792705, respectively.
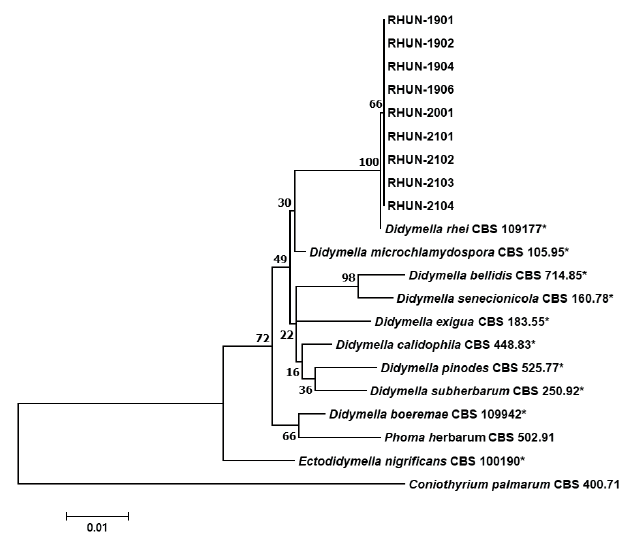
Fig. 3. Phylogenetic tree based on the concatenated sequences of partial large subunit nrDNA (28S, LSU), internal transcribed spacer regions 1 & 2 and 5.8S nrDNA (ITS), and β-tubulin (TUB2) of nine isolates from rhubarb and reference species. The reference sequence data were obtained from the NCBI GenBank database. The tree was generated using maximum likelihood method based on general time reversible model by MEGA version 7. The bootstrap support values are given at the nodes. The bar represents the number of nucleotide substitutions per site. Ex-type strains are marked by an asterisk (*).
Three isolates of D. rhei from rhubarb were used to confirm their pathogenicity to the host plant by artificial inoculation. A conidial suspension (1‒2 × 106 conidia/mL) of each isolate was harvested from 2-week-old cultures on OA and used for the inoculation test on 78-day-old rhubarb plants grown in plastic pots (height: 15 cm; upper diameter: 17 cm; lower diameter: 10 cm) in a vinyl greenhouse. Twenty milliliters of conidial suspension of each isolate was sprayed onto each rhubarb plant. Control plants were treated with sterile distilled water. Inoculated plants were placed in plastic boxes (71.0 cm × 53.5 cm × 40.5 cm) under 100% relative humidity at room temperature (24‒26°C) for 5 days. Thereafter, the inoculated plants were taken out of the plastic boxes and placed in a vinyl greenhouse. Ten days after inoculation, pathogenicity of the tested isolates was rated based on the degree of leaf spot symptoms. The pathogenicity test was conducted in triplicate. All the tested isolates caused leaf spot symptoms in the inoculated plants (Fig. 1C), but no symptoms occurred in the control plants (Fig. 1D). The symptoms induced by the artificial inoculation of plants were similar to those observed in plants from the fields investigated. The inoculated isolates were re-isolated from the lesions.
In this study, the P. rhei isolates causing leaf spot in rhubarb in Korea were identified as D. rhei by phylogenetic analyses. It has been reported that D. rhei (anamorph: P. rhei) causes leaf spot in rhubarb [4,14]. The fungus also causes the disease in several Rheum spp. [3,15]. Leaf spot of rhubarb caused by Phoma sp. has been recorded in Korea [16]. However, there has been no report on the species identification and pathogenicity of the fungus. This is the first report of D. rhei causing leaf spot in rhubarb in Korea.





